Biology:HLA-DQ7
| |
major histocompatibility complex, class II, DQ7
| |
| Haplotypes | DQA1*03:02:DQB1*03:01 DQA1*03:03:DQB1*03:01 DQA1*04:01:DQB1*03:01 DQA1*05:05:DQB1*03:01 DQA1*06:01:DQB1*03:01 |
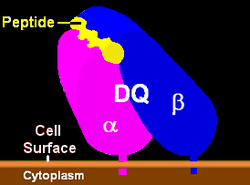
| Structure (See HLA-DQ) | |
| Identifiers | alpha 1 *0302 *0303 *0401 *0505 *0601
|
| Symbol(s) | HLA-DQA1[yes|permanent dead link|dead link}}] |
| EBI-HLA | DQA1*0302 |
| EBI-HLA | DQA1*0303 |
| EBI-HLA | DQA1*0401 |
| EBI-HLA | DQA1*0505 |
| EBI-HLA | DQA1*0601 |
| Identifiers | beta 1 *0301 *0304
|
| Symbol(s) | HLA-DQB1 |
| EBI-HLA | DQB1*0301 |
| EBI-HLA | DQB1*0304 |
| Shared data | |
| Locus | chr.6 6p21.31 |
HLA-DQ7 (DQ7) is an HLA-DQ serotype that recognizes the common HLA DQB1*0301[1] and the less common HLA DQB1*0304 gene products. DQ7 is a form of 'split antigen' of the broad antigen group DQ3 which also contains DQ8 and DQ9.
DQ7 is linked by haplotype to a number of DQA1 (DQ alpha chain) genes, producing in cis-haplotype form, a large number of DQ αβ isoforms. These DQ alpha chains are also known to form transhaplotype isomers with other HLA-DQ.
DQ7 is linked to the following alpha chains genes (DQA1*)
- 03 – *0301, *0302, *0303
- 0401
- 0505
- 0601
Serology
| DQB1* | DQ7 | DQ3 | DQ8 | Sample |
| allele | % | % | % | size (N) |
| 0301 | 85 | 40 | 1 | 12220 |
| 0304 | 40 | 35 | 8 | 111 |
Serotyping efficiency. The serotyping efficiency of DQ7 toward DQB1*0301 is reasonably good, but still results in some false negatives, for *0304 the typing efficiency is poor and cross-reaction with DQ8 is relatively high.
Alleles
DQB1*0301
DQB1*0301 is the major DQ7 allele DQB1*0301 appears to be associated with lupus anticoagulant.[3]
DQB1*0304
DQB1*0304 is the minor DQ7 allele
Haplotypes
| freq | ||
| ref. | Population | (%) |
| [4] | Chukotka Chukchi (Siberia) | 26.7 |
| [4] | Chukotka Eskimos (Siberia) | 25.0 |
| [4] | Koryaks (NE Kamchatka, Siberia) | 19.1 |
| [4] | Polygus Evenks (Siberia) | 11.4 |
| [4] | Khalkh (Ulaanbaatar, Mongolia) | 11.0 |
| [4] | Negidal (Siberia) | 9.6 |
| [4] | Kushun Buryat (Siberia) | 8.0 |
| [4] | Tarialan Khoton (Mongolia) | 7.8 |
| [4] | France Ceph | 6.0 |
| [4] | Russia Tuva (2) | 6.0 |
| [4] | Udegeys Gvaysugi (Siberia) | 4.8 |
| [4] | Irkutsk Tofalar (Siberia ) | 4.7 |
| [4] | Ulchi (Siberia) | 4.1 |
| [4] | Belgian pop2 | 4.1 |
| [4] | England Caucasoid | 4.0 |
| [4] | Italy pop 2 | 2.8 |
| [4] | Russia Tuva Todja | 2.3 |
| [4] | China Ürümqi Kazak | 2.4 |
| [4] | Sulamai Kets (Siberia) | 2.3 |
| [4] | Russia Siberia Nganasan Dudinka | 2.1 |
| [4] | NW Slavic Russia | 2.0 |
| [4] | Japan Fukuoka | 1.2 |
| [4] | Japan (2) | 1.1 |
DQ haplotypes of this serotype are formed between the cis-chromosomal genes of the DQA1 locus. This includes DQA1*0301, *0302, *0303, *0401, *0505, *0601.
There is a rather large degree of disequilibration about DQA1*0301 suggesting that this is one of the older and more established HLA DQB1* alleles in Eurasia. The intron structure of DQB1 suggest that DQB1*0301 DQB1*0302/*0303 split occurred before DQB1*0302/*0303, the distribution of *03 in Africa suggest that recombination DQA1*03:DQB1*0301 are primarily the result of recombination events that have occurred in Africa. A recent study of myasthenia gravis in Houston confirms the presence of A*0505:B*0301 in Nigeria. B1*0301 and A1*03 haplotypes are found at relatively high frequencies in SE Asia and Austronesia, also indicating that it is well established in the exo-African population.
DQ7.3
The DQ7.3 haplotype can be formed by DQA1*0301:DQB1*0301, DQA1*0302:DQB1*0301, DQA1*0303:DQB1*0301. In the west, the DQA1*0303:DQB1*0301 haplotype appears to be more common. The gene products of all 3 function similarly and subunits are interchangeable. In the literature, older DNA tests recognize DQA1*0303 as DQA1*0302, and still oldest DNA tests recognize all three as DQA1*03 or DQA1*0301.
DQA1*0303:DQB1*0301 may be involved in narcolepsy.[5] DQ7.3 appears to be associated with oral ulcerations and gingival disease [6]
DQ7.4
| freq | ||
| ref. | Population | (%) |
| [4] | Chukotka Chukchi (Siberia) | 9.5 |
| [4] | Gvaysugi Udegeys (Siberia) | 9.5 |
| [4] | Chukotka Eskimos (Siberia) | 8.7 |
| [4] | Polygus Evenks (Siberia) | 7.2 |
| [4] | NE Koryaks (Kamchatka) | 6.5 |
| [4] | Cameroon Saa | 4.4 |
| [4] | Sulamai Kets (Siberia) | 2.3 |
| [4] | Gambia | 1.4 |
| [4] | Fukuoka Japan | 1.2 |
| [7] | Caucasian Americans | 0.3 |
DQA1*0401:DQB1*0301 (DQ7.4) This haplotype is found in Siberia, Africa but also at low levels in Western Europe.
DQ7.5
| freq | ||
| ref. | Population | (%) |
| [4] | Lebanon (estimated) | 40.0 |
| [4] | Italy Rome | 29.6 |
| [4] | Netherlands (2) | 15.5 |
| [4] | Tunisia | 14.6 |
| [4] | England (2) | 10.1 |
| [4] | South Korea | 6.8 |
| [4] | Congo Kinshasa Bantu | 4.4 |
DQA1*0505:DQB1*0301 (DQ7.5) was gene-typed as DQA1*0501:DQB1*0301 until it was recognized that there was amino acid sequence variant in the preprocessed DQA1* gene product (proto-α-chain polypeptide encoded DQA1*0505). This proto-alpha, once processed, is identical to the DQA1*0501 encoded α-chain once it is processed. Almost 100% of DQ7.5 haplotypes carry the DQA1*0505 allele.[8] The DR5-DQ7.5 is common in the Southeastern Europe and the Levant, with DQ7.5 reaching a haplotype frequency of 40% in Lebanon. Its high level is probably not by chance, the haplotype appears to protect against juvenile diabetes, which appears to be more common among cereal eating peoples.[9] Cereals were first domesticated in the Near and Middle East more than 10,000 years ago and selection may explain DQ7.5's higher frequencies. (See: Triticeae)
The processed alpha subunit of DQA1*0505 is identical to that of DQA1*0501, but some slight differences in the association with autoimmune disease are observed, possibly as a result of linked DR and DQB1 genes. DQA1*0505 can play into celiac disease under two circumstances. First it can increase risk when DQ2.5 is present, although current studies indicate that it marginally increases risk relative to DQB1*0202 in DQ2.5 cis haplotype. DQA1*0505, without DQ2, is found in a small percentage of coeliac disease (without DQ2 or DQ8).[10]
DQ7.5 is found also high in frequency in the new world, but with DR types less commonly encountered in the old world. DQA1*05 allele is not clear in the new world. DQB1*0301 may be under current positive selection in the human population, at least in areas where DQ2.5 and DQ8 are high, as it confers resistance to type 1 diabetes. For hepatitis type B, DQ7 is associated with persistence but for C, DQ7 is associated with clearance.[11] DQA1*0505, DQB1*0301 appear to increase the risk for melanoma in the Spanish population however this may have a linkage to more recent fair skinned migrants. DQB1*0301 is also associated with allergic fungal sinusitis, human papillomavirus (HPV) induced warts, limited cutaneous systemic sclerosis in Africans, and primary sclerosing cholangitis in Southern Europeans. DQB1*0301 is also predisposing in narcolepsy.[5] DQB1*0301 does not to play a role in any frequently occurring autoimmune disease and its presence in the near east and suppressed frequencies of coeliac disease and Type 1 diabetes in these regions is suggestive that it has a positive selection in Post-Mesolithic cereal based societies in the Western Eurasia.
DQB1*0301 appears to be more associated with early onset myasthenia gravis in Japanese than DQ8, and was also found along with DQB1*0304 to be associated with Chinese MG. DQ7 or associated DR types may play a role in rheumatoid arthritis. In celiac disease the DQ7 (A*0505/1) can mediate celiac disease when HLA DQ2.2 is also present. HLA DQB1*0301 in Turks is associated with Thymoma but the risk may be associated with HLA class I loci.
DQ7.6
| freq | ||
| ref. | Population | (%) |
| [4] | Java Yogyakarta | 48.1 |
| [4] | Kiribati | 37.9 |
| [4] | Nauru | 28.4 |
| [4] | Harbin City (Manchuria, China) | 12.8 |
| [4] | Thailand | 12.7 |
| [4] | South Korean (5) | 4.4 |
| [4] | China Beijing and Xian | 3.5 |
| [4] | Japan | 3.0 |
| [4] | India Bombay | 1.7 |
| [4] | England Caucasoid | 0.6 |
| [4] | Italy Central | 0.6 |
| [4] | Algeria1 | 0.5 |
| [4] | Cameroon | 0.4 |
DQA1*0601:DQB1*0301 (DQ7.6) is a globally rare haplotype, however it is found at high frequencies in the South Pacific and along the West Pacific rim. DQB1*0301 appears to be uniquely linked to DQA1*0601. DQ7.6 is positively associated with asthma,[12] pauciarticular juvenile arthritis without anti-nuclear antibodies,[13] DQ7.6 is negatively associated (Protective against) juvenile diabetes,[14] liver and spleen disease in Schistosoma japonicum infection,[15] pulmonary tuberculosis.[16]
References
- ↑ "Rapid HLA-DQB typing by eight polymerase chain reaction amplifications with sequence-specific primers (PCR-SSP)". Hum. Immunol. 37 (4): 201–6. 1993. doi:10.1016/0198-8859(93)90502-R. PMID 7905469.
- ↑ derived from IMGT/HLA
- ↑ "Molecular analysis of major histocompatibility complex alleles associated with the lupus anticoagulant". J. Clin. Invest. 87 (5): 1490–5. 1991. doi:10.1172/JCI115158. PMID 1673688.
- ↑ 4.00 4.01 4.02 4.03 4.04 4.05 4.06 4.07 4.08 4.09 4.10 4.11 4.12 4.13 4.14 4.15 4.16 4.17 4.18 4.19 4.20 4.21 4.22 4.23 4.24 4.25 4.26 4.27 4.28 4.29 4.30 4.31 4.32 4.33 4.34 4.35 4.36 4.37 4.38 4.39 4.40 4.41 4.42 4.43 4.44 4.45 4.46 4.47 4.48 4.49 4.50 4.51 "New allele frequency database: http://www.allelefrequencies.net". Tissue Antigens 61 (5): 403–7. 2003. doi:10.1034/j.1399-0039.2003.00062.x. PMID 12753660.
- ↑ 5.0 5.1 "DQB1*0301 and DQB1*0601 modulate narcolepsy susceptibility in Koreans". Hum. Immunol. 68 (1): 59–68. 2007. doi:10.1016/j.humimm.2006.10.006. PMID 17207713.
- ↑ "Common major histocompatibility complex class II markers in clinical variants of cicatricial pemphigoid.". Proc Natl Acad Sci U S A 91 (16): 7747–51. Aug 1994. doi:10.1073/pnas.91.16.7747. PMID 8052655. Bibcode: 1994PNAS...91.7747Y.
- ↑ "New HLA haplotype frequency reference standards: high-resolution and large sample typing of HLA DR-DQ haplotypes in a sample of European Americans". Tissue Antigens 62 (4): 296–307. 2003. doi:10.1034/j.1399-0039.2003.00103.x. PMID 12974796.
- ↑ "Novel associations among HLA-DQA1 and -DQB1 alleles, revealed by high-resolution sequence-based typing (SBT)". Tissue Antigens 55 (3): 275–9. 2000. doi:10.1034/j.1399-0039.2000.550313.x. PMID 10777105. https://zenodo.org/record/1231460.
- ↑ "Association of selective HLA class II susceptibility-conferring and protective haplotypes with type 2 diabetes in patients from Bahrain and Lebanon". Clin. Vaccine Immunol. 13 (11): 1296–8. 2006. doi:10.1128/CVI.00206-06. PMID 16988007.
- ↑ "HLA types in celiac disease patients not carrying the DQA1*05-DQB1*02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease". Hum. Immunol. 64 (4): 469–77. 2003. doi:10.1016/S0198-8859(03)00027-2. PMID 12651074.
- ↑ "A comparative review of HLA associations with hepatitis B and C viral infections across global populations". World J. Gastroenterol. 13 (12): 1770–87. 2007. doi:10.3748/wjg.v13.i12.1770. PMID 17465466.
- ↑ "[Association between asthma and the polymorphism of HLA-DQ genes]" (in zh). Zhonghua Jie He He Hu Xi Za Zhi 24 (3): 139–41. 2001. PMID 11802952.
- ↑ "Antinuclear antibodies in early onset pauciarticular juvenile chronic arthritis (JCA) are associated with HLA-DQB1*0603: a possible JCA-associated human leucocyte antigen haplotype". Br. J. Rheumatol. 34 (5): 461–5. 1995. doi:10.1093/rheumatology/34.5.461. PMID 7788177.
- ↑ "HLA DQA1 genotypes and its interaction with HLA DQB1 in Chinese IDDM living in Taiwan". Proc. Natl. Sci. Counc. Repub. China B 19 (2): 73–9. 1995. PMID 7624445.
- ↑ "HLA class II antigens are associated with resistance or susceptibility to hepatosplenic disease in a Chinese population infected with Schistosoma japonicum". Int. J. Parasitol. 28 (4): 537–42. 1998. doi:10.1016/S0020-7519(98)00020-4. PMID 9602373.
- ↑ "Associations of HLA class II alleles with pulmonary tuberculosis in Thais". Eur. J. Immunogenet. 29 (5): 431–4. 2002. doi:10.1046/j.1365-2370.2002.00352.x. PMID 12358854.
 |

