Biology:LONI Pipeline
This article has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
(Learn how and when to remove this template message)
|
 | |
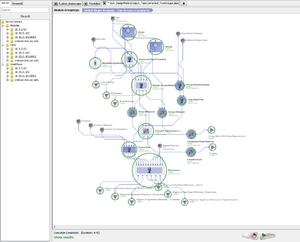 Pipeline Environment | |
| Developer(s) | Samuel Hobel |
|---|---|
| Stable release | 7.0.3
/ March 3, 2020 |
| Written in | Java |
| Operating system | Linux, Mac OS X, Microsoft Windows |
| Type | Scientific workflow system, Workflow processing environments |
| License | LONI License |
| Website | pipeline |
The LONI Pipeline is a free distributed system for designing, executing, monitoring and sharing scientific workflows[1][2] on grid computing architectures. Pipeline allows users to connect and run any number of different software tools, and conveniently visualize and download the results.
Unlike other workflow processing environments, Pipeline does not require new tools and services to include or be built against the core Pipeline libraries. The Pipeline environment references all data, services and tools as external objects. This allows the Pipeline to run as a light-weight middleware, but at the same time, restrict the scope of its applications. For example, the Pipeline does not provide a set of internal core libraries, filters, and processes for rudimentary image processing (e.g., image addition). All tools necessary to complete an analysis protocol must first be built as external stand-alone applications or services, whose interface methods are then described in the Pipeline XML language. Users can connect to the LONI Cranium server to gain quick access to a wide array of pre-built software applications, such as FSL, AFNI, and FreeSurfer already described in XML as modules and workflows. Pipeline allows users to create new workflow descriptions, edit existing ones, and share their work with others.
Typical pipeline server installations include a suite of core resources that are available to all users with access to the specific server, however, different servers will have different suites of default module and module-group (pipeline) definitions. The previous release (version 5) of the LONI Pipeline[3] provided a mechanism for integrating heterogeneous and incongruous data including images, clinical charts and demographic meta-data.
The LONI Pipeline has hundreds of users in a variety of fields (e.g., genomics,[4] neuro-imaging,[5] and Biomedical Informatics[6]) from academic institutions around the world.
Features
Pipeline has cross platform compatibility, and the ability to connect from your local client to a remote server for executing processing and analysis on other operating systems.
Pipeline grants developers the opportunity to create their own plugins to communicate with various grid managers. The default Pipeline package includes the JGDIPlugin and the DRMAAPlugin plugins created for Sun Grid Engine but they may work with Oracle Grid Engine, Univa Grid Engine or Son of Grid Engine. Both plugins are housed under the gridplugins directory which is parented under the dist directory in the installed package of Pipeline. All additional plugins you wish to employ can be downloaded separately.
The Pipeline Library grants users access to hundreds of predefined neuroimaging solutions, including data, modules and workflows that are regularly updated.
Other integral features of the LONI Pipeline are:
- Distributed client-server and platform-agnostic computational infrastructure
- Reliable, asynchronous and secure data processing
- Automated and intelligent data format conversion
- Local and remote file browser
- Imaging and metadata processing modules
- Conditional and iterating modules
- A detailed parameter transformation system
- Launch client from web browser via Pipeline Web Start
Developers
Present:
- Samuel Hobel
Past:
- Denis Trotckii
- David Rex
- Michael Pan
- Celia Cheung
- Kamen Lozev
- Jia-Wei Tam
- Zhizhong Liu
- Wei Yan
- Petros Petrosyan
- Arash Payan
- Ivo Dinov
See also
- Laboratory Of Neuro Imaging
- Kepler scientific workflow system
- Taverna workbench
- Bioinformatics workflow management systems
References
- ↑ Rex, D. E., Ma, J.Q., and Toga, A.W. (2003). "The LONI Pipeline Processing Environment." Neuroimage, 19(3), 1033-48.
- ↑ Rex, D. E., Shattuck, D. W., Woods, R. P., Narr, K. L., Luders, E., Rehm, K., Stolzner, S. E., Rottenberg, D. E., and Toga, A. W. (2004). "A meta-algorithm for brain extraction in MRI." NeuroImage, 23(2), 625–637
- ↑ Dinov ID, Lozev K, Petrosyan P, Liu Z, Eggert P, Pierce, J, Zamanyan, A, Chakrapani, S, Van Horn, JD, Parker, DS, Magsipoc, R, Leung, K, Gutman, B, Woods, RP, Toga, AW. (2010). "Neuroimaging Study Designs, Computational Analyses and Data Provenance Using the LONI Pipeline." PLoS ONE 5(9): e13070. doi:10.1371/journal.pone.0013070.
- ↑ Torri, F., Dinov, ID, Zamanyan, A, Hobel, S, Genco, A, Petrosyan, P, Clark, AP, Liu, Z, Eggert, P, Pierce, J, Knowles, JA, Ames, J, Kesselman, C, Toga, AW, Potkin, SG, Vawter, MP, Macciardi, F. (2012) Next Generation Sequence Analysis and Computational Genomics Using Graphical Pipeline Workflows, Genes, 3(3):545-575; doi:10.3390/genes3030545.
- ↑ Woo MS, Dinov, ID, Hobel, S, Zamanyan, A, Choi, YC, Thompson, PM, Toga, AW and Alzheimer’s Disease Neuroimaging Initiative (ADNI) (2015) Structural Brain Changes in Early-Onset Alzheimer’s Disease Subjects Using the LONI Pipeline Environment. Journal of Neuroimaging., in press. DOI: 10.1111/jon.12252
- ↑ Toga, WA, Dinov, ID. (2015) Sharing big biomedical data. Journal of Big Data., 2(7):1-12. DOI: 10.1186/s40537-015-0016-1.
External links
 |
