Biology:PubMed
 | |
|---|---|
| Contact | |
| Research centre | United States National Library of Medicine (NLM) |
| Release date | January 1996 |
| Access | |
| Website | pubmed |
PubMed is a free search engine accessing primarily the MEDLINE database of references and abstracts on life sciences and biomedical topics. The United States National Library of Medicine (NLM) at the National Institutes of Health maintain the database as part of the Entrez system of information retrieval.[1]
From 1971 to 1997, online access to the MEDLINE database had been primarily through institutional facilities, such as university libraries.[2] PubMed, first released in January 1996, ushered in the era of private, free, home- and office-based MEDLINE searching.[3] The PubMed system was offered free to the public starting in June 1997.[2]
Content
In addition to MEDLINE, PubMed provides access to:
- older references from the print version of Index Medicus, back to 1951 and earlier
- references to some journals before they were indexed in Index Medicus and MEDLINE, for instance Science, BMJ, and Annals of Surgery
- very recent entries to records for an article before it is indexed with Medical Subject Headings (MeSH) and added to MEDLINE
- a collection of books available full-text and other subsets of NLM records[4]
- PMC citations
- NCBI Bookshelf
Many PubMed records contain links to full text articles, some of which are freely available, often in PubMed Central[5] and local mirrors, such as Europe PubMed Central.[6]
Information about the journals indexed in MEDLINE, and available through PubMed, is found in the NLM Catalog.[7]
As of 23 May 2023[update], PubMed has more than 35 million citations and abstracts dating back to 1966, selectively to the year 1865, and very selectively to 1809. As of the same date[update], 24.6 million of PubMed's records are listed with their abstracts, and 26.8 million records have links to full-text versions (of which 10.9 million articles are available, full-text for free).[8] Over the last 10 years (ending 31 December 2019), an average of nearly one million new records were added each year.
In 2016, NLM changed the indexing system so that publishers are able to directly correct typos and errors in PubMed indexed articles.[9]
PubMed has been reported to include some articles published in predatory journals. MEDLINE and PubMed policies for the selection of journals for database inclusion are slightly different. Weaknesses in the criteria and procedures for indexing journals in PubMed Central may allow publications from predatory journals to leak into PubMed.[10] The National Library of Medicine had respond that individual journal articles can be included in PMC to support the public access policies of research funders and that rigorous policies about journals and publishers ensure integrity of NLM literature databases.[11]
Characteristics
Website design
A new PubMed interface was launched in October 2009 and encouraged the use of such quick, Google-like search formulations; they have also been described as 'telegram' searches.[12] By default the results are sorted by Most Recent, but this can be changed to Best Match, Publication Date, First Author, Last Author, Journal, or Title.[13]
The PubMed website design and domain was updated in January 2020 and became default on 15 May 2020, with the updated and new features.[14] There was a critical reaction from many researchers who frequently use the site.[15]
PubMed for handhelds/mobiles
PubMed/MEDLINE can be accessed via handheld devices, using for instance the "PICO" option (for focused clinical questions) created by the NLM.[16] A "PubMed Mobile" option, providing access to a mobile friendly, simplified PubMed version, is also available.[17]
Search
Standard search
Simple searches on PubMed can be carried out by entering key aspects of a subject into PubMed's search window.
PubMed translates this initial search formulation and automatically adds field names, relevant MeSH (Medical Subject Headings) terms, synonyms, Boolean operators, and 'nests' the resulting terms appropriately, enhancing the search formulation significantly, in particular by routinely combining (using the OR operator) textwords and MeSH terms.[citation needed]
The examples given in a PubMed tutorial[18] demonstrate how this automatic process works:
Causes Sleep Walking is translated as ("etiology"[Subheading] OR "etiology"[All Fields] OR "causes"[All Fields] OR "causality"[MeSH Terms] OR "causality"[All Fields]) AND ("somnambulism"[MeSH Terms] OR "somnambulism"[All Fields] OR ("sleep"[All Fields] AND "walking"[All Fields]) OR "sleep walking"[All Fields])
Likewise,
soft Attack Aspirin Prevention is translated as ("myocardial infarction"[MeSH Terms] OR ("myocardial"[All Fields] AND "infarction"[All Fields]) OR "myocardial infarction"[All Fields] OR ("heart"[All Fields] AND "attack"[All Fields]) OR "heart attack"[All Fields]) AND ("aspirin"[MeSH Terms] OR "aspirin"[All Fields]) AND ("prevention and control"[Subheading] OR ("prevention"[All Fields] AND "control"[All Fields]) OR "prevention and control"[All Fields] OR "prevention"[All Fields])
Comprehensive search
For optimal searches in PubMed, it is necessary to understand its core component, MEDLINE, and especially of the MeSH (Medical Subject Headings) controlled vocabulary used to index MEDLINE articles. They may also require complex search strategies, use of field names (tags), proper use of limits and other features; reference librarians and search specialists offer search services.[19][20]
The search into PubMed's search window is only recommended for the search of unequivocal topics or new interventions that do not yet have a MeSH heading created, as well as for the search for commercial brands of medicines and proper nouns. It is also useful when there is no suitable heading or the descriptor represents a partial aspect. The search using the thesaurus MeSH is more accurate and will give fewer irrelevant results. In addition, it saves the disadvantage of the free text search in which the spelling, singular/plural or abbreviated differences have to be taken into consideration. On the other side, articles more recently incorporated into the database to which descriptors have not yet been assigned will not be found. Therefore, to guarantee an exhaustive search, a combination of controlled language headings and free text terms must be used.[21]
Journal article parameters
When a journal article is indexed, numerous article parameters are extracted and stored as structured information. Such parameters are: Article Type (MeSH terms, e.g., "Clinical Trial"), Secondary identifiers, (MeSH terms), Language, Country of the Journal or publication history (e-publication date, print journal publication date).
Publication Type: Clinical queries/systematic reviews
Publication type parameter allows searching by the type of publication, including reports of various kinds of clinical research.[22]
Secondary ID
Since July 2005, the MEDLINE article indexing process extracts identifiers from the article abstract and puts those in a field called Secondary Identifier (SI). The secondary identifier field is to store accession numbers to various databases of molecular sequence data, gene expression or chemical compounds and clinical trial IDs. For clinical trials, PubMed extracts trial IDs for the two largest trial registries: ClinicalTrials.gov (NCT identifier) and the International Standard Randomized Controlled Trial Number Register (IRCTN identifier).[23]
See also
A reference which is judged particularly relevant can be marked and "related articles" can be identified. If relevant, several studies can be selected and related articles to all of them can be generated (on PubMed or any of the other NCBI Entrez databases) using the 'Find related data' option. The related articles are then listed in order of "relatedness". To create these lists of related articles, PubMed compares words from the title and abstract of each citation, as well as the MeSH headings assigned, using a powerful word-weighted algorithm.[24] The 'related articles' function has been judged to be so precise that the authors of a paper suggested it can be used instead of a full search.[25]
Mapping to MeSH
PubMed automatically links to MeSH terms and subheadings. Examples would be: "bad breath" links to (and includes in the search) "halitosis", "heart attack" to "myocardial infarction", "breast cancer" to "breast neoplasms". Where appropriate, these MeSH terms are automatically "expanded", that is, include more specific terms. Terms like "nursing" are automatically linked to "Nursing [MeSH]" or "Nursing [Subheading]". This feature is called Auto Term Mapping and is enacted, by default, in free text searching but not exact phrase searching (i.e. enclosing the search query with double quotes).[26] This feature makes PubMed searches more sensitive and avoids false-negative (missed) hits by compensating for the diversity of medical terminology.[26]
PubMed does not apply automatic mapping of the term in the following circumstances: by writing the quoted phrase (e.g., "kidney allograft"), when truncated on the asterisk (e.g., kidney allograft*), and when looking with field labels (e.g., Cancer [ti]).[21]
My NCBI
The PubMed optional facility "My NCBI" (with free registration) provides tools for
- saving searches
- filtering search results
- setting up automatic updates sent by e-mail
- saving sets of references retrieved as part of a PubMed search
- configuring display formats or highlighting search terms
and a wide range of other options.[27] The "My NCBI" area can be accessed from any computer with web-access. An earlier version of "My NCBI" was called "PubMed Cubby".[28]
LinkOut
LinkOut is an NLM facility to link and make available full-text local journal holdings.[29] Some 3,200 sites (mainly academic institutions) participate in this NLM facility (as of March 2010[update]), from Aalborg University in Denmark to ZymoGenetics in Seattle.[30] Users at these institutions see their institution's logo within the PubMed search result (if the journal is held at that institution) and can access the full-text. Link out is being consolidated with Outside Tool as of the major platform update coming in the Summer of 2019.[31]
PubMed Commons
In 2016, PubMed allows authors of articles to comment on articles indexed by PubMed. This feature was initially tested in a pilot mode (since 2013) and was made permanent in 2016.[32] In February 2018, PubMed Commons was discontinued due to the fact that "usage has remained minimal".[33][34]
askMEDLINE
askMEDLINE, a free-text, natural language query tool for MEDLINE/PubMed, developed by the NLM, also suitable for handhelds.[35]
PubMed identifier
A PMID (PubMed identifier or PubMed unique identifier)[36] is a unique integer value, starting at 1, assigned to each PubMed record. A PMID is not the same as a PMCID (PubMed Central identifier) which is the identifier for all works published in the free-to-access PubMed Central.[37]
The assignment of a PMID or PMCID to a publication tells the reader nothing about the type or quality of the content. PMIDs are assigned to letters to the editor, editorial opinions, op-ed columns, and any other piece that the editor chooses to include in the journal, as well as peer-reviewed papers. The existence of the identification number is also not proof that the papers have not been retracted for fraud, incompetence, or misconduct. The announcement about any corrections to original papers may be assigned a PMID.
Each number that is entered in the PubMed search window is treated by default as if it were a PMID. Therefore, any reference in PubMed can be located using the PMID.
Alternative interfaces
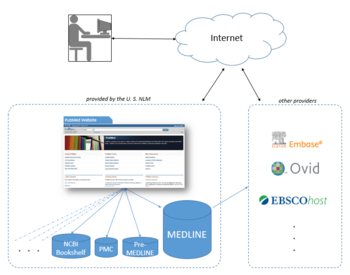
The National Library of Medicine leases the MEDLINE information to a number of private vendors such as Embase, Ovid, Dialog, EBSCO, Knowledge Finder and many other commercial, non-commercial, and academic providers.[38] As of October 2008[update], more than 500 licenses had been issued, more than 200 of them to providers outside the United States. As licenses to use MEDLINE data are available for free, the NLM in effect provides a free testing ground for a wide range[39] of alternative interfaces and 3rd party additions to PubMed, one of a very few large, professionally curated databases which offers this option.
Lu identifies a sample of 28 current and free Web-based PubMed versions, requiring no installation or registration, which are grouped into four categories:[39]
- Ranking search results, for instance: eTBLAST; MedlineRanker;[40] MiSearch;[41]
- Clustering results by topics, authors, journals etc., for instance: Anne O'Tate;[42] ClusterMed;[43]
- Enhancing semantics and visualization, for instance: EBIMed;[44] MedEvi.[45]
- Improved search interface and retrieval experience, for instance, askMEDLINE[46][47] BabelMeSH;[48] and PubCrawler.[49]
As most of these and other alternatives rely essentially on PubMed/MEDLINE data leased under license from the NLM/PubMed, the term "PubMed derivatives" has been suggested.[39] Without the need to store about 90 GB of original PubMed Datasets, anybody can write PubMed applications using the eutils-application program interface as described in "The E-utilities In-Depth: Parameters, Syntax and More", by Eric Sayers, PhD.[50] Various citation format generators, taking PMID numbers as input, are examples of web applications making use of the eutils-application program interface. Sample web pages include Citation Generator – Mick Schroeder, Pubmed Citation Generator – Ultrasound of the Week, PMID2cite, and Cite this for me.
Data mining of PubMed
Alternative methods to mine the data in PubMed use programming environments such as Matlab, Python or R. In these cases, queries of PubMed are written as lines of code and passed to PubMed and the response is then processed directly in the programming environment. Code can be automated to systematically query with different keywords such as disease, year, organs, etc.
In addition to its traditional role as a biomedical database, PubMed has become common resource for training biomedical language models.[51] Recent advancements in this field include the development of models like PubMedGPT, a 2.7B parameter model trained on PubMed data by Stanford CRFM, and Microsoft's BiomedCLIP-PubMedBERT, which utilizes figure-caption pairs from PubMed Central for vision-language processing. These models demonstrate the significant potential of PubMed data in enhancing the capabilities of AI in medical research and healthcare applications. Such advancements underline the growing intersection between large-scale data mining and AI development in the biomedical field.
The data accessible by PubMed can be mirrored locally using an unofficial tool such as MEDOC.[52]
Millions of PubMed records augment various open data datasets about open access, like Unpaywall. Data analysis tools like Unpaywall Journals are used by libraries to assist with big deal cancellations: libraries can avoid subscriptions for materials already served by instant open access via open archives like PubMed Central.[53]
See also
- Europe PubMed Central
- JournalReview.org
- List of academic databases and search engines
- PubMed Central
- PubMed Central Canada
References
- ↑ "PubMed". https://www.nlm.nih.gov/bsd/pubmed.html.
- ↑ 2.0 2.1 "Internet access to the National Library of Medicine". Effective Clinical Practice 3 (5): 256–60. 2000. PMID 11185333. http://www.acponline.org/clinical_information/journals_publications/ecp/sepoct00/nlm.pdf.
- ↑ "PubMed Celebrates its 10th Anniversary". United States National Library of Medicine. 2006-10-05. https://www.nlm.nih.gov/pubs/techbull/so06/so06_pm_10.html.
- ↑ "PubMed: MEDLINE Retrieval on the World Wide Web". United States National Library of Medicine. 2002-06-07. https://www.nlm.nih.gov/pubs/factsheets/pubmed.html.
- ↑ "PubMed Central: The GenBank of the published literature". Proceedings of the National Academy of Sciences of the United States of America 98 (2): 381–2. January 2001. doi:10.1073/pnas.98.2.381. PMID 11209037. Bibcode: 2001PNAS...98..381R.
- ↑ "UKPMC: a full text article resource for the life sciences". Nucleic Acids Research 39 (Database issue): D58-65. January 2011. doi:10.1093/nar/gkq1063. PMID 21062818.
- ↑ "NLM Catalogue: Journals referenced in the NCBI Databases". NCBI. 2011. https://www.ncbi.nlm.nih.gov/nlmcatalog/journals.
- ↑ "PubMed" (in en). https://pubmed.ncbi.nlm.nih.gov/. The search query "1800:2100[dp]" retrieves all results whose date of publication is between 1800 and 2100 inclusive.
- ↑ "MEDLINE/PubMed Production Improvements Underway". NLM Technical Bulletin (411): e1. July–August 2016. https://www.nlm.nih.gov/pubs/techbull/ja16/ja16_medline_pm_production.html. Retrieved 29 July 2016.
- ↑ "How predatory journals leak into PubMed". CMAJ 190 (35): E1042–E1045. September 2018. doi:10.1503/cmaj.180154. PMID 30181150.
- ↑ Topper, Lauren; Marill, Jennifer; Kelly, Christopher; Funk, Kathryn (2019-03-11). "Rigorous policies ensure integrity of NLM literature databases". CMAJ 191 (10): E289. doi:10.1503/cmaj.71602. ISSN 1488-2329. PMID 30858186.
- ↑ "Pragmatic approach is effective in evidence based health care". BMJ 321 (7260): 566–7. September 2000. doi:10.1136/bmj.321.7260.566/a. PMID 10968827.
- ↑ "How to improve your PubMed/MEDLINE searches: 2. display settings, complex search queries and topic searching". Journal of Telemedicine and Telecare 20 (1): 44–55. January 2014. doi:10.1177/1357633X13517067. PMID 24352897.
- ↑ Trawick, Bart (21 January 2020). "A New and Improved PubMed®". NLM Musings From the Mezzanine. https://nlmdirector.nlm.nih.gov/2020/01/21/a-new-and-improved-pubmed/.
- ↑ Price, Michael (22 May 2020). "They redesigned PubMed, a beloved website. It hasn't gone over well". Science. https://www.science.org/content/article/they-redesigned-pubmed-beloved-website-it-hasn-t-gone-over-well.
- ↑ "PubMed via handhelds (PICO)". United States National Library of Medicine. 2004. https://www.nlm.nih.gov/pubs/techbull/mj04/mj04_pico.html.
- ↑ "PubMed Mobile Beta". United States National Library of Medicine. 2011. https://www.nlm.nih.gov/pubs/techbull/ma11/ma11_pm_mobile_beta.html.
- ↑ "Simple Subject Search with Quiz". NCBI. 2010. https://www.nlm.nih.gov/bsd/viewlet/search/subject/subject.html.
- ↑ "Searching the literature. Be systematic in your searching". BMJ 307 (6895): 66. July 1993. doi:10.1136/bmj.307.6895.66-a. PMID 8343701.
- ↑ "The art and science of searching MEDLINE to answer clinical questions. Finding the right number of articles". International Journal of Technology Assessment in Health Care 15 (2): 281–96. Spring 1999. doi:10.1017/S0266462399015214. PMID 10507188. https://escholarship.umassmed.edu/qhs_pp/130. Retrieved 13 December 2019.
- ↑ 21.0 21.1 "Cómo elaborar una estrategia de búsqueda bibliográfica" (in es). Enfermería Intensiva 29 (4): 182–186. 2018. doi:10.1016/j.enfi.2018.09.001. PMID 30291015.
- ↑ Clinical Queries Filter Terms explained. NCBI. 2010. https://www.ncbi.nlm.nih.gov/entrez/query/static/clinicaltable.html. Retrieved 8 September 2017.
- ↑ "Evaluating adherence to the International Committee of Medical Journal Editors' policy of mandatory, timely clinical trial registration". Journal of the American Medical Informatics Association 20 (e1): e169-74. June 2013. doi:10.1136/amiajnl-2012-001501. PMID 23396544.
- ↑ "Computation of Related Articles explained". NCBI. https://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=helppubmed.section.pubmedhelp.Appendices#pubmedhelp.Computation_of_Relat.
- ↑ "Searching the literature using medical subject headings versus text word with PubMed". The Laryngoscope 116 (2): 336–40. February 2006. doi:10.1097/01.mlg.0000195371.72887.a2. PMID 16467730. http://www.escholarship.org/uc/item/8cn089mk. Retrieved 11 September 2018.
- ↑ 26.0 26.1 "How to improve your PubMed/MEDLINE searches: 3. advanced searching, MeSH and My NCBI". Journal of Telemedicine and Telecare 20 (2): 102–12. March 2014. doi:10.1177/1357633X13519036. PMID 24614997.
- ↑ "My NCBI Help". My NCBI explained. NCBI. 13 December 2010. https://www.ncbi.nlm.nih.gov/books/NBK3842/. Retrieved 8 September 2017.
- ↑ "PubMed Cubby". United States National Library of Medicine. 2000. https://www.nlm.nih.gov/pubs/techbull/so00/so00_hands_on_register.html.
- ↑ "LinkOut Overview". NCBI. 2010. https://www.ncbi.nlm.nih.gov/projects/linkout/.
- ↑ "LinkOut Participants 2011". NCBI. 2011. https://www.ncbi.nlm.nih.gov/projects/linkout/journals/htmllists.cgi?type_id=6.
- ↑ "An Updated PubMed is on its Way". https://www.nlm.nih.gov/pubs/techbull/ma19/ma19_pubmed_update.html.
- ↑ PubMed Commons Team (17 December 2015). "Commenting on PubMed: A Successful Pilot". https://pubmedcommonsblog.ncbi.nlm.nih.gov/2015/12/17/commenting-on-pubmed-a-successful-pilot/.
- ↑ "PubMed Commons to be Discontinued" (in en-US). NCBI Insights. 2018-02-01. https://ncbiinsights.ncbi.nlm.nih.gov/2018/02/01/pubmed-commons-to-be-discontinued/.
- ↑ "PubMed shuts down its comments feature, PubMed Commons". 2018-02-02. http://retractionwatch.com/2018/02/02/pubmed-shuts-comments-feature-pubmed-commons/.
- ↑ "askMedline". NCBI. 2005. http://askmedline.nlm.nih.gov/ask/ask.php.
- ↑ "Search Field Descriptions and Tags". National Center for Biotechnology Information. https://www.ncbi.nlm.nih.gov/books/NBK3827/#pubmedhelp.Search_Field_Descrip.
- ↑ Keener, Molly. "PMID vs. PMCID: What's the difference?". University of Chicago. http://chess.uchicago.edu/docs/PMCIDinPubMed.pdf.
- ↑ "Leasing journal citations from PubMed/Medline". NLM. 2011. https://www.nlm.nih.gov/databases/journal.html.
- ↑ 39.0 39.1 39.2 "PubMed and beyond: a survey of web tools for searching biomedical literature". Database 2011: baq036. 2011. doi:10.1093/database/baq036. PMID 21245076.
- ↑ "MedlineRanker: flexible ranking of biomedical literature". Nucleic Acids Research 37 (Web Server issue): W141-6. July 2009. doi:10.1093/nar/gkp353. PMID 19429696.
- ↑ "MiSearch adaptive pubMed search tool". Bioinformatics 25 (7): 974–6. April 2009. doi:10.1093/bioinformatics/btn033. PMID 18326507.
- ↑ "Anne O'Tate: A tool to support user-driven summarization, drill-down and browsing of PubMed search results". Journal of Biomedical Discovery and Collaboration 3: 2. February 2008. doi:10.1186/1747-5333-3-2. PMID 18279519.
- ↑ "ClusterMed". Vivisimo Clustering Engine. 2011. http://demos.vivisimo.com/clustermed.
- ↑ "EBIMed--text crunching to gather facts for proteins from Medline". Bioinformatics 23 (2): e237-44. January 2007. doi:10.1093/bioinformatics/btl302. PMID 17237098.
- ↑ "MedEvi: retrieving textual evidence of relations between biomedical concepts from Medline". Bioinformatics 24 (11): 1410–2. June 2008. doi:10.1093/bioinformatics/btn117. PMID 18400773.
- ↑ "askMEDLINE: a report on a year-long experience". AMIA ... Annual Symposium Proceedings. AMIA Symposium 2006: 923. 2006. PMID 17238542.
- ↑ "MeSH Speller + askMEDLINE: auto-completes MeSH terms then searches MEDLINE/PubMed via free-text, natural language queries". AMIA ... Annual Symposium Proceedings. AMIA Symposium 2005: 957. 2005. PMID 16779244.
- ↑ "PICO Linguist and BabelMeSH: development and partial evaluation of evidence-based multilanguage search tools for MEDLINE/PubMed". Studies in Health Technology and Informatics 129 (Pt 1): 817–21. 2007. PMID 17911830. http://booksonline.iospress.nl/Extern/EnterMedLine.aspx?ISSN=0926-9630&Volume=129&SPage=817. Retrieved 31 May 2014.
- ↑ "PubCrawler: keeping up comfortably with PubMed and GenBank". Nucleic Acids Research 32 (Web Server issue): W16-9. July 2004. doi:10.1093/nar/gkh453. PMID 15215341.
- ↑ Eric Sayers, PhD (24 October 2018). The E-utilities In-Depth: Parameters, Syntax and More. NCBI. https://www.ncbi.nlm.nih.gov/books/NBK25499/. Retrieved 8 September 2017.
- ↑ Singhal, Karan; Azizi, Shekoofeh; Tu, Tao; Mahdavi, S. Sara; Wei, Jason; Chung, Hyung Won; Scales, Nathan; Tanwani, Ajay et al. (3 August 2023). "Large language models encode clinical knowledge". Nature 620 (7972): 172–180. doi:10.1038/s41586-023-06291-2.
- ↑ on GitHub
- ↑ Denise Wolfe (2020-04-07). "SUNY Negotiates New, Modified Agreement with Elsevier – Libraries News Center University at Buffalo Libraries". University at Buffalo. https://library.buffalo.edu/news/2020/04/07/suny-negotiates-new-modified-agreement-with-elsevier/.
External links
Wikidata has the property:
|
 |
