Biology:Sul1 RNA motif
| sul1 | |
|---|---|
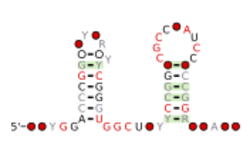 Consensus secondary structure and sequence conservation of sul1 RNA | |
| Identifiers | |
| Symbol | sul1 |
| Rfam | RF03058 |
| Other data | |
| RNA type | Cis-reg; Riboswitch |
| SO | 0000035 |
| PDB structures | PDBe |
The sul1 RNA motif is a conserved RNA structure that was discovered by bioinformatics.[1] Energetically stable tetraloops often occur in this motif. sul1 motif RNAs are found in Alphaproteobacteria.
sul1 motif RNAs likely function as cis-regulatory elements, in view of their positions upstream of protein-coding genes. Indeed, the RNAs are upstream of multiple genes that encode non-homologous proteins. If all examples of the RNA were upstream of homologous genes, there is the possibility that the RNAs were conserved in that position simply by inheritance. The non-homology of the genes downstream of sul1 RNAs makes this scenario less likely.
Among the proteins encoded by genes that are apparently regulated by sul1 RNAs, the most common protein domains (whose gene is named sul1) is believed to function as a sulfate transporter. Other common protein domains function as Serine O-acetyltransferase, Cyclopropane-fatty-acyl-phospholipid synthase, S-adenosylmethionine-dependent methyltransferase or glycosyltransferase. It was observed[1] that many of these genes are related to sulfur metabolism or to methionine metabolism, and therefore sul1 RNAs' function might relate to these pathways. If sul1 RNAs function by sensing ions such as sulfate or metabolites involved in these pathways, they would qualify as riboswitches.
References
- ↑ 1.0 1.1 "Detection of 224 candidate structured RNAs by comparative analysis of specific subsets of intergenic regions". Nucleic Acids Res. 45 (18): 10811–10823. October 2017. doi:10.1093/nar/gkx699. PMID 28977401.
 |

