Biology:YcaO
| YcaO-like family | |||||||||
|---|---|---|---|---|---|---|---|---|---|
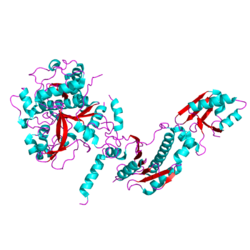 Structure of the heterocyclase TruD. A YcaO-like protein from Prochloron sp. 06037A PDB entry 4bs9[1] | |||||||||
| Identifiers | |||||||||
| Symbol | YcaO | ||||||||
| Pfam | PF02624 | ||||||||
| InterPro | IPR003776 | ||||||||
| |||||||||
YcaO is a protein found in bacteria which is involved in the synthesis of thiazole/oxazole modified microcin antibiotics, such as bottromycin. YcaO performs ATP dependent cyclodehydration to form the oxazole and thiazole moieties of the microcin.[2][3][4]
The YcaO name origin is from a gene naming rubric that was established from the bacterium Escherichia coli. If a gene has an unknown function, it was given a four-letter name starting with the letter Y and the next three letters are given based on the genomic location. [5]
Methyl coenzyme M reductase (MCR) or Coenzyme-B sulfoethylthiotransferase is a protein known in thioamidation (a posttranslational modification). A Ycao enzyme dependent on ATP is needed for MCR thioamidation as well as a sulfide source. YcaO enzymes are needed to catalyze the ATP-dependent backbone cyclodehydration of polar amino acids such as Cysteine, Serine, and Threonine to the correct thiazoline and (methyl) oxazoline Heterocycle.[6] The side chains of these amino acids can act as Nucleophiles. The Thiol group in cysteine and the hydroxyl group of serine and threonine are strong nucleophiles.
References
- ↑ "The cyanobactin heterocyclase enzyme: a processive adenylase that operates with a defined order of reaction". Angewandte Chemie 52 (52): 13991–13996. December 2013. doi:10.1002/anie.201306302. PMID 24214017.
- ↑ "YcaO domains use ATP to activate amide backbones during peptide cyclodehydrations". Nature Chemical Biology 8 (6): 569–575. April 2012. doi:10.1038/nchembio.944. PMID 22522320.
- ↑ "Discovery of a new ATP-binding motif involved in peptidic azoline biosynthesis". Nature Chemical Biology 10 (10): 823–829. October 2014. doi:10.1038/nchembio.1608. PMID 25129028.
- ↑ "Structural analysis of leader peptide binding enables leader-free cyanobactin processing". Nature Chemical Biology 11 (8): 558–563. August 2015. doi:10.1038/nchembio.1841. PMID 26098679.
- ↑ "YcaO-Dependent Posttranslational Amide Activation: Biosynthesis, Structure, and Function". Chemical Reviews 117 (8): 5389–5456. April 2017. doi:10.1021/acs.chemrev.6b00623. PMID 28256131.
- ↑ "Enzymatic reconstitution of ribosomal peptide backbone thioamidation". Proceedings of the National Academy of Sciences of the United States of America 115 (12): 3030–3035. March 2018. doi:10.1073/pnas.1722324115. PMID 29507203.
 |

