Biology:CsrC RNA family
| CsrC RNA family | |
|---|---|
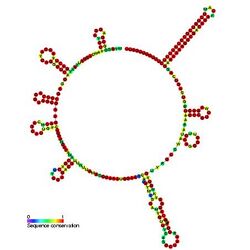 Predicted secondary structure and sequence conservation of CsrC | |
| Identifiers | |
| Symbol | CsrC |
| Alt. Symbols | SraK |
| Rfam | RF00084 |
| Other data | |
| RNA type | Gene; sRNA |
| Domain(s) | Bacteria |
| SO | 0000655 |
| PDB structures | PDBe |
The 245 nucleotide sRNA of Escherichia coli, CsrC, was discovered using a genetic screen for factors that regulate glycogen biosynthesis. CsrC RNA binds multiple copies of CsrA, a protein that post-transcriptionally regulates central carbon flux, biofilm formation and motility in E. coli. CsrC antagonises the regulatory effects of CsrA, presumably by sequestering this protein. The discovery of CsrC is intriguing, in that a similar sRNA, CsrB, performs essentially the same function. Both sRNAs possess similar imperfect repeat sequences (18 in CsrB, nine in CsrC), primarily localised in the loops of predicted hairpins, which may serve as CsrA binding elements. Transcription of csrC increases as the culture approaches the stationary phase of growth and is indirectly activated by CsrA via the response regulator UvrY [1]. This RNA was also discovered in E. coli during a large scale screen [2]. The gene called SraK, was highly abundant in stationary phase, but low levels could be detected in exponentially growing cells as well [2].
See also
- CsrB/RsmB RNA family
- PrrB/RsmZ RNA family
- RsmY RNA family
- RsmX
- CsrA protein
References
Further reading
- "A novel sRNA component of the carbon storage regulatory system of Escherichia coli". Molecular Microbiology 48 (3): 657–670. May 2003. doi:10.1046/j.1365-2958.2003.03459.x. PMID 12694612.
- "Novel small RNA-encoding genes in the intergenic regions of Escherichia coli". Current Biology 11 (12): 941–950. June 2001. doi:10.1016/S0960-9822(01)00270-6. PMID 11448770.
- "A direct link between the global regulator PhoP and the Csr regulon in Y. pseudotuberculosis through the small regulatory RNA CsrC". RNA Biology 11 (5): 580–593. Apr 2, 2014. doi:10.4161/rna.28676. PMID 24786463.
External links
 |

