Biology:Fusarium oxysporum
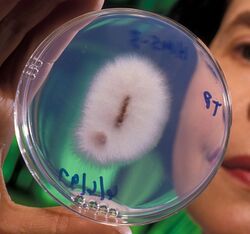
| Fusarium oxysporum | |
|---|---|
| |
| Scientific classification | |
| Domain: | Eukaryota |
| Kingdom: | Fungi |
| Division: | Ascomycota |
| Class: | Sordariomycetes |
| Order: | Hypocreales |
| Family: | Nectriaceae |
| Genus: | Fusarium |
| Species: | F. oxysporum
|
| Binomial name | |
| Fusarium oxysporum Schlecht. emend. Snyder & Hansen
| |
Fusarium oxysporum (Schlecht as emended by Snyder and Hansen),[1] an ascomycete fungus, comprises all the species, varieties and forms recognized by Wollenweber and Reinking[2] within an infrageneric grouping called section Elegans. It is part of the family Nectriaceae.
Although their predominant role in native soils may be as harmless or even beneficial plant endophytes or soil saprophytes, many strains within the F. oxysporum complex are soil borne pathogens of plants, especially in agricultural settings.
Taxonomy
While the species, as defined by Snyder and Hansen, has been widely accepted for more than 50 years,[3][4] more recent work indicates this taxon is actually a genetically heterogeneous polytypic morphospecies,[5][6] whose strains represent some of the most abundant and widespread microbes of the global soil microflora.[7]
Genome
The Fot1 family of transposable elements was first discovered by Daboussi et al., 1992 in several formae speciales[8][9] and Davière et al., 2001 and Langin et al., 2003 have since found them in most strains at copy numbers as high as 100.[8]
Habitat
These diverse and adaptable fungi have been found in soils ranging from the Sonoran Desert, to tropical and temperate forest, grasslands and soils of the tundra.[10] F. oxysporum strains are ubiquitous soil inhabitants that have the ability to exist as saprophytes, and degrade lignin[11][12] and complex carbohydrates[13][14][1] associated with soil debris. They are pervasive plant endophytes that can colonize plant roots[15][16] and may even protect plants or form the basis of disease suppression.[17][18]
Because the hosts of a given forma specialis usually are closely related, many have assumed that members of a forma specialis are also closely related and descended from a common ancestor.[19] However, results from research conducted on Fusarium oxysporum f. sp. cubense forced scientists to question these assumptions. Researchers used anonymous, single-copy restriction fragment length polymorphsims (RFLPs) to identify 10 clonal lineages from a collection of F. oxysporum f.sp. cubense from across the world. These results showed that pathogens of banana causing Panama disease could be as closely related to other host's pathogens, such as melon or tomato, as they are to each other. Exceptional amounts of genetic diversity within F. oxysporum f.sp. cubense were deduced from the high level of chromosomal polymorphisms found among strains, random amplified polymorphic DNA fingerprints and from the number and geographic distribution of vegetative compatibility groups.[20]
Pathogen
Presented with the wide-ranging occurrence of F. oxysporum strains that are nonpathogenic, it is reasonable to conclude that certain pathogenic forms were descended from originally nonpathogenic ancestors. Given the association of these fungi with plant roots, a form that is able to grow beyond the cortex and into the xylem could exploit this ability and hopefully gain an advantage over fungi that are restricted to the cortex.[citation needed]
The progression of a fungus into vascular tissue may elicit an immediate host response, successfully restricting the invader; or an otherwise ineffective or delayed response, reducing the vital water-conducting capacity and induce wilting.[21] On the other hand, the plant might be able to tolerate limited growth of the fungus within xylem vessels, preceded by an endophytic association.[22] In this case, any further changes in the host or parasite could disturb the relationship, in a way that fungal activities or a host response would result in the generation of disease symptoms.
Pathogenic strains of F. oxysporum have been studied for more than 100 years. The host range of these fungi is broad and includes animals, ranging from arthropods[23] to humans,[24] as well as plants, including a range of both gymnosperms and angiosperms. While collectively, plant pathogenic F. oxysporum strains have a broad host range, individual isolates usually cause disease only in a narrow range of plant species. This observation has led to the idea of "special form" or forma specialis in F. oxysporum. Formae speciales have been defined as "…an informal rank in Classification… used for parasitic fungi characterized from a physiological standpoint (e.g. by the ability to cause disease in particular hosts) but scarcely or not at all from a morphological standpoint." Exhaustive host range studies have been conducted for relatively few formae speciales of F. oxysporum.[25] For more information on Fusarium oxysporum as a plant pathogen, see Fusarium wilt and Koa wilt.
Different strains of F. oxysporum have been used for the purpose of producing nanomaterials (especially Silver nanoparticles).
"Agent Green" in Colombia
In 2000, the government of Colombia proposed dispersing strains of Crivellia and Fusarium oxysporum, also known as Agent Green, as a biological weapon to forcibly eradicate coca and other illegal crops.[26][self-published source?] The weaponized strains were developed by the US government, who originally conditioned their approval of Plan Colombia on the use of this weapon, but ultimately withdrew the condition.[27] In February 2001, the EU Parliament issued a declaration specifically against the use of these biological agents in warfare.[27]
Gold interactions
The fungus has the ability to dissolve gold, then precipitate it onto its surface, encrusting itself with gold. This phenomenon was first observed in Boddington, West Australia.[28] As a result of this discovery, F. oxysporum is currently being evaluated as a possible way to help detect hidden underground gold reserves.[29] It also is used to manufacture gold nanoparticles.[30]
Formae speciales
- Fusarium oxysporum f.sp. albedinis
- Fusarium oxysporum f.sp. asparagi
- Fusarium oxysporum f.sp. batatas
- Fusarium oxysporum f.sp. betae
- Fusarium oxysporum f.sp. cattleyae
- Fusarium oxysporum f.sp. cannabis
- Fusarium oxysporum f.sp. cepae
- Fusarium oxysporum f.sp. ciceris
- Fusarium oxysporum f.sp. citri
- Fusarium oxysporum f.sp. coffea
- Fusarium oxysporum f.sp. cubense
- Fusarium oxysporum f.sp. cyclaminis
- Fusarium oxysporum f.sp. herbemontis
- Fusarium oxysporum f.sp. dianthi
- Fusarium oxysporum f. sp. fragariae
- Fusarium oxysporum f.sp. gladioli
- Fusarium oxysporum f.sp. koae
- Fusarium oxysporum f.sp. lactucae
- Fusarium oxysporum f.sp. lentis
- Fusarium oxysporum f.sp. lilli[31]
- Fusarium oxysporum f.sp. lini
- Fusarium oxysporum f.sp. lycopersici
- Fusarium oxysporum f.sp. medicaginis
- Fusarium oxysporum f.sp. melonis
- Fusarium oxysporum f.sp. momordicae
- Fusarium oxysporum f.sp. narcissi [32][33]
- Fusarium oxysporum f.sp. nicotianae
- Fusarium oxysporum f.sp. niveum
- Fusarium oxysporum f.sp. palmarum
- Fusarium oxysporum f.sp. passiflorae
- Fusarium oxysporum f.sp. perniciosum
- Fusarium oxysporum f.sp. phaseoli
- Fusarium oxysporum f.sp. pisi
- Fusarium oxysporum f.sp. radicis-lycopersici
- Fusarium oxysporum f.sp. ricini
- Fusarium oxysporum f.sp. strigae
- Fusarium oxysporum f.sp. tuberosi
- Fusarium oxysporum f.sp. tulipae
- Fusarium oxysporum f.sp. vasinfectum
See also
References
- ↑ 1.0 1.1 "The Species Concept in Fusarium". American Journal of Botany 27 (2): 64–67. 1940. doi:10.1002/j.1537-2197.1940.tb14217.x.
- ↑ (in German) Die Fusarien, ihre Beschreibung, Schadwirkung und Bekampfung. Berlin: P. Parey. 1935. OCLC 1123368362.[page needed]
- ↑ The genus Fusarium. Commonwealth Agricultural Bureaux [for the] Commonwealth Mycological Institute. 1971. ISBN 978-0-85198-046-1. OCLC 281402.[page needed]
- ↑ Fusarium species: an illustrated manual for identification. The Pennsylvania State University Press. 1983. ISBN 978-0-271-00349-8. OCLC 802515637.[page needed]
- ↑ "Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous". Molecular Phylogenetics and Evolution 7 (1): 103–16. February 1997. doi:10.1006/mpev.1996.0376. PMID 9007025.
- ↑ "Discordant Groupings of Fusarium spp. from Sections Elegans, Liseola and Dlaminia Based on Ribosomal ITS1 and ITS2 Sequences". Mycologia 88 (3): 361–368. 1996. doi:10.1080/00275514.1996.12026663.
- ↑ "The evolutionary biology of Fusarium oxysporum". Annual Review of Phytopathology 35: 111–28. 1997. doi:10.1146/annurev.phyto.35.1.111. PMID 15012517.
- ↑ 8.0 8.1 Daboussi, Marie-Josée; Capy, Pierre (2003). "Transposable Elements in Filamentous Fungi". Annual Review of Microbiology (Annual Reviews) 57 (1): 275–299. doi:10.1146/annurev.micro.57.030502.091029. ISSN 0066-4227. PMID 14527280.
- ↑ Daboussi, Marie-Josée; Langin, Thierry; Brygoo, Yves (1992). "Fot1, a new family of fungal transposable elements". Molecular & General Genetics (Springer) 232 (1): 12–16. doi:10.1007/bf00299131. ISSN 0026-8925. PMID 1313143.
- ↑ "Ecology of Fusarium in noncultivated soils". Fusarium: diseases, biology, and taxonomy. Pennsylvania State University Press. 1981. pp. 276–286. ISBN 978-0-271-00293-4.
- ↑ "Degradation of natural lignins and lignocellulosic substrates by soil-inhabiting fungi imperfecti". FEMS Microbiology Ecology 21 (3): 213–219. November 1996. doi:10.1111/j.1574-6941.1996.tb00348.x. Bibcode: 1996FEMME..21..213R.
- ↑ "Lignocellulose degradation by Fusarium species". Canadian Journal of Botany 61 (4): 1194–1198. 1 April 1983. doi:10.1139/b83-126.
- ↑ "Purification and mode of action of a low molecular mass endo-1,4-β-d-glucanase from Fusarium oxysporum". Journal of Biotechnology 39 (1): 85–93. February 1995. doi:10.1016/0168-1656(94)00147-5.
- ↑ "Purification and characterization of two low molecular mass alkaline xylanases from Fusarium oxysporum F3". Journal of Biotechnology 51 (2): 181–9. November 1996. doi:10.1016/0168-1656(96)01619-7. PMID 8987884.
- ↑ "Colonization of muskmelon and nonsusceptible crops by Fusarium oxysporum f. sp. melonis and other species of Fusarium". Phytopathology 79 (10): 1095–1100. 1989. doi:10.1094/Phyto-79-1095. https://www.apsnet.org/publications/phytopathology/backissues/Documents/1989Abstracts/Phyto79_1095.htm.
- ↑ "Symptomless carriers of the tomato Fusarium wilt pathogen". Phytopathology 61 (10): 1213–1217. 1971. doi:10.1094/Phyto-61-1213. https://www.apsnet.org/publications/phytopathology/backissues/Documents/1971Abstracts/Phyto61_1213.htm.
- ↑ "Effect of successive watermelon plantings on Fusarium oxysporum and other microorganisms in soils suppressive and conducive to fusarium wilt of watermelon". Phytopathology 83 (10): 1097–1105. 1993. doi:10.1094/Phyto-83-1097. https://www.apsnet.org/publications/phytopathology/backissues/Documents/1993Abstracts/Phyto_83_1097.htm.
- ↑ "Antagonistic Effect of Nonpathogenic Fusarium oxysporum Fo47 and Pseudobactin 358 upon Pathogenic Fusarium oxysporum f. sp. dianthi". Applied and Environmental Microbiology 59 (1): 74–82. January 1993. doi:10.1128/AEM.59.1.74-82.1993. PMID 16348860. Bibcode: 1993ApEnM..59...74L.
- ↑ "Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies". Proceedings of the National Academy of Sciences of the United States of America 95 (5): 2044–9. March 1998. doi:10.1073/pnas.95.5.2044. PMID 9482835. Bibcode: 1998PNAS...95.2044O.
- ↑ "Evolutionary relationships among the Fusarium oxysporum f. sp. cubense vegetative compatibility groups". Applied and Environmental Microbiology 75 (14): 4770–81. July 2009. doi:10.1128/AEM.00370-09. PMID 19482953. Bibcode: 2009ApEnM..75.4770F.
- ↑ "Fusarium Wilt of Banana Is Caused by Several Pathogens Referred to as Fusarium oxysporum f. sp. cubense". Phytopathology 96 (6): 653–6. June 2006. doi:10.1094/PHYTO-96-0653. PMID 18943184.
- ↑ "Local and regional variation in populations of Fusarium oxysporum from agricultural field soils". Phytopathology 84 (8): 786–791. 1994. doi:10.1094/Phyto-84-786. https://www.apsnet.org/publications/phytopathology/backissues/Documents/1994Abstracts/Phyto_84_786.htm.
- ↑ "Entomogenous Fusarium species". Mycopathologia 84 (1): 3–16. December 1983. doi:10.1007/BF00436991. PMID 6369143.
- ↑ "Taxonomy, biology, and clinical aspects of Fusarium species". Clinical Microbiology Reviews 7 (4): 479–504. October 1994. doi:10.1128/cmr.7.4.479. PMID 7834602.
- ↑ "Evolution of host specificity in Fusarium oxysporum". Fusarium: Paul E. Nelson Memorial Symposium. APS Press. 2001. pp. 70–82. ISBN 978-0-89054-268-2. OCLC 46786813.
- ↑ "Grain — Sprouting Up: Battle Lines Drawn over Agent Green". https://www.grain.org/article/entries/306-sprouting-up-battle-lines-drawn-over-agent-green.
- ↑ 27.0 27.1 "EU Parliament Rejects Agent Green for Colombia" (Press release). The Sunshine Project. 1 February 2001.
- ↑ CSIRO. "Gold-coated fungi" (in en). https://www.csiro.au/en/work-with-us/industries/mining-resources/exploration/sampling-methods/gold-fungi.
- ↑ "Evidence for fungi and gold redox interaction under Earth surface conditions". Nature Communications 10 (1): 2290. May 2019. doi:10.1038/s41467-019-10006-5. PMID 31123249. Bibcode: 2019NatCo..10.2290B.
- Mindy Weisberger (May 24, 2019). "This Fungus Mines For Gold, Then Wears It". https://www.livescience.com/65562-gold-studded-fungus-australia.html.
- ↑ Sayadi, Khali; Akbarzadeh, Fatemeh; Pourmardan, Vahid; Saravani-Aval, Mehdi; Sayadi, Jalis; Chauhan, Narendra Pal Singh; Sargazi, Ghasem (2021). "Methods of green synthesis of Au NCs with emphasis on their morphology: A mini-review". Heliyon (Cell Press) 7 (6): e07250. doi:10.1016/j.heliyon.2021.e07250. ISSN 2405-8440. PMID 34189304. Bibcode: 2021Heliy...707250S.
- ↑ "Detection, Diagnosis and Control of Lily Diseases". Acta Horticulturae (900): 313–324. July 2011. doi:10.17660/ActaHortic.2011.900.40.
- ↑ "Control of Fusarium oxysporum f.sp. narcissi, the cause of narcissus basal rot, with thiabendazole and other fungicides". Crop Protection 15 (6 September): 549–558. 1996. doi:10.1016/0261-2194(96)00023-3.
- ↑ Hanks, Gordon; Carder, John (2003). "Management of basal rot - the narcissus disease". Pesticide Outlook 14 (6): 260. doi:10.1039/B314848N.
Wikidata ☰ Q139958 entry
 |

