Biology:Haplogroup M-P256
| Haplogroup M-P256 | |
|---|---|
 | |
| Possible time of origin | 32,000-47,000 years BP (Scheinfeldt 2006) |
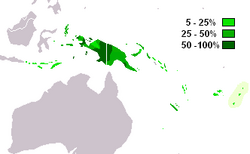
| Possible place of origin | Wallacea (eastern Indonesia) or New Guinea [1] |
| Ancestor | K2b1 |
| Defining mutations | P256 |
Haplogroup M, also known as M-P256 and Haplogroup K2b1b (previously K2b1d) is a Y-chromosome DNA haplogroup. M-P256 is a descendant haplogroup of Haplogroup K2b1, and is believed to have first appeared between 32,000 and 47,000 years ago (Scheinfeldt 2006).
M-P256 is the most frequently occurring Y-chromosome haplogroup in West Papua and western Papua New Guinea (Kayser 2003). It is also found in neighbouring parts of Melanesia, Indonesia and among indigenous Australians.
It and Haplogroup S (B254) are the only primary subclades of K2b1, also known as haplogroup MS.
Phylogenetic structure
This phylogenetic tree of haplogroup subclades is based primarily on the trees published by ISOGG in 2016 (ISOGG 2016) and YCC in 2008.(Karafet 2008)
- M* (P256)
- M1 (M4, M5/P73, M106, M186, M189, M296, P35)
- M1a(P34_1, P34_2, P34_3, P34_4, P34_5)
- M1a1 (P51)
- M1a2 (P94)
- M1b (P87)
- M1b1 (M104_1/P22_1, M104_2/P22_2)
- M1b1a (M16)
- M1b1b (M83)
- M1b1 (M104_1/P22_1, M104_2/P22_2)
- M1a(P34_1, P34_2, P34_3, P34_4, P34_5)
- M2 (M353, M387)
- M2a (M177/SRY9138)
- M3 (P117, P118)
- M1 (M4, M5/P73, M106, M186, M189, M296, P35)
Distribution
M* (M-P256*)
The paragroup M-P256* is found at low incidences[spelling?] in New Guinea (6.3%) and Flores (2.5%).[1]
M1 (M-M4)
| Haplogroup M-M4 | |
|---|---|
| Possible time of origin | 8,200 [3,800–20,600] years BP (Kayser 2003) |
| Possible place of origin | Southeast Asia - Melanesia[citation needed] |
| Ancestor | M-P256 |
| Defining mutations | M4, M5/P73, M106, M186, M189, M296, P35[citation needed] |
Found frequently in New Guinea and Melanesia, with a moderate distribution in neighboring parts of Indonesia, Micronesia, and Polynesia.
- Una 100% (Kayser 2003)
- Ketengban 100% (Kayser 2003)
- Awyu 100% (Kayser 2003)
- Citak 86% (Kayser 2003)
- Asmat 75% (Kayser 2003)
- West Papua
- lowlands/coast 77.5% (Kayser 2003)
- highlands 74.5% (Kayser 2003)
- Kombai/Korowai 46% (Kayser 2003)
- Papua New Guinea
- coast 29% (Kayser 2003)
- highlands 35.5% (Kayser 2003)
- Tolai (New Britain) 31% (Kayser 2003)
- Trobriand Islands 30% (Kayser 2003)
- Maluku (Moluccas) 21% (Kayser 2003)
- Torres Strait Islanders (Australia): up to 2.0% – i.e. 0.9% of samples, when 45% of the total were deemed to be "non-indigenous".(Nagle 2015)
An extreme geographical outlier was apparently identified in a 2012 study, which reported a Hazara individual from Mazar-e Sharif, Afghanistan, with M1 among a sample of 60 males from Mazar-e Sharif.(Haber 2012). The Hazara individual carried the SNP M186 (which is believed to be equivalent to M4).
| Old names (YCC 2002/2008) | M-M4 |
| Jobling and Tyler-Smith 2000 | 24 |
| Underhill 2000 | VIII |
| Hammer 2001 | 1U |
| Karafet 2001 | 37 |
| Semino 2000 | Eu16 |
| Su 1999 | H17 |
| Capelli 2001 | E |
| YCC 2002 (Longhand) | M* |
| YCC 2005 (Longhand) | M |
| YCC 2008 (Longhand) | M1 |
| YCC 2010r (Longhand) | M1 |
M1a (M-P34)
M1a (M-P34) is the most frequently occurring Y-chromosome DNA haplogroup in Western New Guinea. It is also found with moderate frequency in neighboring parts of Indonesia (Maluku, Nusa Tenggara) and throughout Papua New Guinea, including offshore islands (Karafet 2005 and Kayser 2008).
| Old names (YCC 2002/2008) | M-P34 |
| Jobling and Tyler-Smith 2000 | 24 |
| Underhill 2000 | VIII |
| Hammer 2001 | 1U |
| Karafet 2001 | 37 |
| Semino 2000 | Eu16 |
| Su 1999 | H17 |
| Capelli 2001 | E |
| YCC 2002 (Longhand) | M1 |
| YCC 2005 (Longhand) | M1 |
| YCC 2008 (Longhand) | M1a |
| YCC 2010r (Longhand) | M1a |
M1b (M-P87)
M1b M-P87(xM104/P22) has been found in approximately 18% (20/109) of a pool of samples from New Ireland, approximately 12% (5/43) of a sample of Lavongai from New Hanover, approximately 5% (19/395) of a pool of samples from New Britain (and, in particular, in about 24% (15/63) of Baining from East New Britain), in one Saposa individual from northern Bougainville, and in another individual from the north coast of Papua New Guinea (Scheinfeldt 2006).
The subclade M1b1 (M104_1/P22_1, M104_2/P22_2) is found frequently in populations of the Bismarck Archipelago and Bougainville Island, with a moderate distribution in New Guinea, Fiji, Tonga, East Futuna, and Samoa. (Kayser 2008 and Scheinfeldt 2006).
| Old names (YCC 2002/2008) | M-P22 |
| Jobling and Tyler-Smith 2000 | 24 |
| Underhill 2000 | VIII |
| Hammer 2001 | 1U |
| Karafet 2001 | 38 |
| Semino 2000 | Eu16 |
| Su 1999 | H17 |
| Capelli 2001 | E |
| YCC 2002 (Longhand) | M2* |
| YCC 2005 (Longhand) | M2a |
| YCC 2008 (Longhand) | M1b1 |
| YCC 2010r (Longhand) | M1b1 |
M2 (M-M353)
Found at a low frequency in Fiji and East Futuna (Kayser 2006).
The subclade M2a (M-M177 a.k.a. M-SRY9138) is found in one Nasioi individual from the eastern coast of Bougainville and in one individual from Malaita Province of the Solomon Islands (Cox 2006).
Historic names for M-SRY9138 (a.k.a. M-M177) from peer reviewed literature.
| Old names (YCC 2002/2008) | K-SRY9138/M-SRY9138 AKA M-M177 |
| Jobling and Tyler-Smith 2000 | 23 |
| Underhill 2000 | VIII |
| Hammer 2001 | 1E |
| Karafet 2001 | 25 |
| Semino 2000 | Eu16 |
| Su 1999 | H5 |
| Capelli 2001 | F |
| YCC 2002 (Longhand) | K1 |
| YCC 2005 (Longhand) | K1 |
| YCC 2008 (Longhand) | M2a |
| YCC 2010r (Longhand) | M2a |
M3 (M-P117)
M3 (P117, P118) is found frequently in populations of New Britain, and also observed occasionally in northern Bougainville, Fiji, and East Futuna (Kayser 2008 and Scheinfeldt 2006).
Previous phylogenetic history
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M4 | 24 | VIII | 1U | 37 | Eu16 | H17 | E | M* | M | M1 | M1 | - | - | - | - | - | - | - |
| M-P34 | 24 | VIII | 1U | 37 | Eu16 | H17 | E | M1 | M1 | M1a | M1a | - | - | - | - | - | - | - |
| M-P22/M-M104 | 24 | VIII | 1U | 38 | Eu16 | H17 | E | M2* | M2a | M1b1 | M1b1 | - | - | - | - | - | - | - |
| M-M16 | 24 | VIII | 1U | 39 | Eu16 | H17 | E | M2a | M2a1 | M1b1a | M1b1a | - | - | - | - | - | - | - |
| M-M83 | 24 | VIII | 1U | 38 | Eu16 | H17 | E | M2b | M2a2 | M1b1b | M1b1b | - | - | - | - | - | - | - |
| K-SRY9138/M-SRY9138 | 23 | VIII | 1E | 25 | Eu16 | H5 | F | K1 | K1 | M2a | M2a | - | - | - | - | - | - | - |
- Sources
The following research teams per their publications were represented in the creation of the YCC Tree.
Karafet's 2008 paper introduced a number of changes, compared to the previous 2006 ISOGG tree. Before the discovery of the P256 marker, the current subgroup M-M4 (defined by the M4 marker) previously represented the whole of Haplogroup M-P256; and subgroups M2 and M3 were formerly classed as subgroups K1 and K7 of the parent Haplogroup K.[citation needed]
References
Footnotes
Works cited
- Cox, Murray P.; Mirazón Lahr, Marta (2006). "Y-chromosome diversity is inversely associated with language affiliation in paired Austronesian- and Papuan-speaking communities from Solomon Islands". American Journal of Human Biology 18 (1): 35–50. doi:10.1002/ajhb.20459. PMID 16378340.
- Haber, Marc; Platt, Daniel E.; Ashrafian Bonab, Maziar; Youhanna, Sonia C.; Soria-Hernanz, David F.; Martínez-Cruz, Begoña; Douaihy, Bouchra; Ghassibe-Sabbagh, Michella et al. (2012). Kayser, Manfred. ed. "Afghanistan's Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events". PLOS ONE 7 (3): e34288. doi:10.1371/journal.pone.0034288. PMID 22470552. Bibcode: 2012PLoSO...734288H.
- Karafet, Tatiana M.; Lansing, J. S.; Redd, Alan J.; Watkins, Joseph C.; Surata, S. P. K.; Arthawiguna, W. A.; Mayer, Laura; Bamshad, Michael et al. (2005). "Balinese Y-Chromosome Perspective on the Peopling of Indonesia: Genetic Contributions from Pre-Neolithic Hunter-Gatherers, Austronesian Farmers, and Indian Traders". Human Biology 77 (1): 93–114. doi:10.1353/hub.2005.0030. PMID 16114819. https://kuscholarworks.ku.edu/bitstream/1808/13586/1/Karafet_et_al_2005.pdf.
- Karafet, T. M.; Mendez, F. L.; Meilerman, M. B.; Underhill, P. A.; Zegura, S. L.; Hammer, M. F. (2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research 18 (5): 830–8. doi:10.1101/gr.7172008. PMID 18385274.
- Kayser, Manfred; Brauer, Silke; Weiss, Gunter; Schiefenhövel, Wulf; Underhill, Peter; Shen, Peidong; Oefner, Peter; Tommaseo-Ponzetta, Mila et al. (2003). "Reduced Y-Chromosome, but Not Mitochondrial DNA, Diversity in Human Populations from West New Guinea". The American Journal of Human Genetics 72 (2): 281–302. doi:10.1086/346065. PMID 12532283.
- Kayser, M.; Brauer, S; Cordaux, R; Casto, A; Lao, O; Zhivotovsky, LA; Moyse-Faurie, C; Rutledge, RB et al. (2006). "Melanesian and Asian Origins of Polynesians: MtDNA and Y Chromosome Gradients Across the Pacific". Molecular Biology and Evolution 23 (11): 2234–44. doi:10.1093/molbev/msl093. PMID 16923821. https://hal.archives-ouvertes.fr/hal-00117338/document.
- Kayser, M.; Choi, Y.; Van Oven, M.; Mona, S.; Brauer, S.; Trent, R. J.; Suarkia, D.; Schiefenhovel, W. et al. (2008). "The Impact of the Austronesian Expansion: Evidence from mtDNA and Y Chromosome Diversity in the Admiralty Islands of Melanesia". Molecular Biology and Evolution 25 (7): 1362–74. doi:10.1093/molbev/msn078. PMID 18390477.
- Scheinfeldt, L.; Friedlaender, F; Friedlaender, J; Latham, K; Koki, G; Karafet, T; Hammer, M; Lorenz, J (2006). "Unexpected NRY Chromosome Variation in Northern Island Melanesia". Molecular Biology and Evolution 23 (8): 1628–41. doi:10.1093/molbev/msl028. PMID 16754639.
- Nagle, N. (2015). "Antiquity and diversity of aboriginal Australian Y-chromosomes". American Journal of Physical Anthropology 159 (3): 367–81. doi:10.1002/ajpa.22886. PMID 26515539.
- "ISOGG 2016 Y-DNA Haplogroup M". http://www.isogg.org/tree/ISOGG_HapgrpM.html.
Sources for conversion tables
- Capelli, Cristian; Wilson, James F.; Richards, Martin; Stumpf, Michael P.H. et al. (February 2001). "A Predominantly Indigenous Paternal Heritage for the Austronesian-Speaking Peoples of Insular Southeast Asia and Oceania". The American Journal of Human Genetics 68 (2): 432–443. doi:10.1086/318205.
- Hammer, Michael F.; Karafet, Tatiana M.; Redd, Alan J.; Jarjanazi, Hamdi et al. (1 July 2001). "Hierarchical Patterns of Global Human Y-Chromosome Diversity". Molecular Biology and Evolution 18 (7): 1189–1203. doi:10.1093/oxfordjournals.molbev.a003906.
- Jobling, Mark A.; Tyler-Smith, Chris (2000), "New uses for new haplotypes", Trends in Genetics 16 (8): 356–62, doi:10.1016/S0168-9525(00)02057-6, PMID 10904265
- Kaladjieva, Luba; Calafell, Francesc; Jobling, Mark A; Angelicheva, Dora et al. (February 2001). "Patterns of inter- and intra-group genetic diversity in the Vlax Roma as revealed by Y chromosome and mitochondrial DNA lineages". European Journal of Human Genetics 9 (2): 97–104. doi:10.1038/sj.ejhg.5200597.
- Karafet, Tatiana; Xu, Liping; Du, Ruofu; Wang, William et al. (September 2001). "Paternal Population History of East Asia: Sources, Patterns, and Microevolutionary Processes". The American Journal of Human Genetics 69 (3): 615–628. doi:10.1086/323299.
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA et al. (2000), "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective", Science 290 (5494): 1155–9, doi:10.1126/science.290.5494.1155, PMID 11073453, Bibcode: 2000Sci...290.1155S
- Su, Bing; Xiao, Junhua; Underhill, Peter; Deka, Ranjan et al. (December 1999). "Y-Chromosome Evidence for a Northward Migration of Modern Humans into Eastern Asia during the Last Ice Age". The American Journal of Human Genetics 65 (6): 1718–1724. doi:10.1086/302680.
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics 26 (3): 358–361. doi:10.1038/81685.
External links
- Spread of Haplogroup M, from National Geographic
See also
- Genetic genealogy
- Haplogroup
- Haplotype
- Human Y-chromosome DNA haplogroup
- Molecular phylogeny
- Paragroup
- Subclade
- Y-chromosome haplogroups in populations of the world
- Y-DNA haplogroups by ethnic group
- Y-DNA haplogroups in populations of East and Southeast Asia
 |

