Biology:Ketoacyl synthase
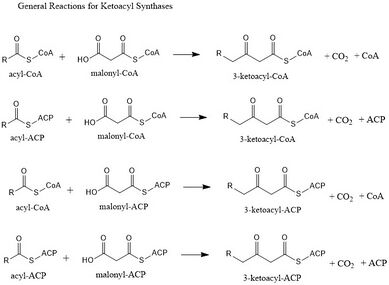
Ketoacyl synthases (KSs) catalyze the condensation reaction of acyl-CoA or acyl-acyl ACP with malonyl-CoA to form 3-ketoacyl-CoA or with malonyl-ACP to form 3-ketoacyl-ACP. This reaction is a key step in the fatty acid synthesis cycle, as the resulting acyl chain is two carbon atoms longer than before. KSs exist as individual enzymes, as they do in type II fatty acid synthesis and type II polyketide synthesis, or as domains in large multidomain enzymes, such as type I fatty acid synthases (FASs) and polyketide synthases (PKSs). KSs are divided into five families: KS1, KS2, KS3, KS4, and KS5.[1]
Multidomain enzyme systems
Fatty acid synthase
Fatty acid synthase (FAS) is the enzyme system involved in de novo fatty acid synthesis. FAS is an iterative multienzyme consisting of several component enzymes, one of which is ketoacyl synthase. There are two types of FASs: type I and type II. Type I FASs are highly integrated multidomain enzymes. They contain discrete functional domains responsible for specific catalytic activities of the reaction sequence, either on a single polypeptide chain or on two different multifunctional proteins. Type II FASs are dissociated systems, meaning the component enzymes are independent proteins encoded by a series of separate genes.[2]
Polyketide synthase
Polyketide synthases (PKS) are structurally and functionally related to FAS’s, both which are enzymes that catalyze the condensation of activated primary metabolites such as acetyl-CoA and malonyl-CoA.
The main reaction they catalyze is:[3]
- CO2-CH2-CO-S-CoA + CH3-CO-S-PKS → CH3-CO-CH2-CO-S-PKS + CoA-H + CO2
Like FASs, PKSs will use a β-ketoacylsynthase (KS), an optional (malonyl) acyl transferase (MAT/AT), and a phosphopantethienylated acyl carrier protein (ACP) or coenzymeA (CoA). They also both used a ketoreductase, dehydratase, and enoyl reductase to create a fully saturated acyl backbone. Unlike FASs, however, PKSs typically use a larger number of biosynthetic building blocks and form a more varied number of tail lengths. The reductive steps that FASs use are also optional for the PKSs. By potentially omitting them, there is potential for a more complex pattern of functionalization.[4]
There are three main types of polyketides: Type I, type II and type III. Type I is very similar to the FAS type I, in that it contains linearly aligned and covalently fused catalytic domains within large multifunctional enzymes. Type II tends to be a more dissociable complex with monofunctional enzyme domains. Another way that PKSs differ is that they have one other type, Type III. Type III PKSs are multifunctional when choosing a starting unit, assembling the chain, and promoting the folding.[4]
Ketoacyl synthase family 1
Nearly all KS1 members are produced by bacteria, with a few formed by eukaryota and only one by an archaeon. There are 12 subfamilies. The dominant enzyme in the KS1 family is 3-ketoacyl-ACP synthase III (KAS III), also known as 3-oxoacyl-ACP synthase III and β-ketoacyl-ACP synthase III, and is defined as EC 2.3.1.180.[5][1]
β-Ketoacyl-ACP synthase III
The characteristic reaction of β-ketoacyl-ACP synthase III is malonyl-ACP + acetyl-CoA => acetoacyl-ACP + CO2 + CoA. Cysteine, histidine, and asparagine form the catalytic triad in KAS III, which uses the ping-pong kinetic mechanism.[1]
In Escherichia coli, one organism KAS III is typically found in, KASIII is weakly inhibited by thiolactomycin.[6] In the same organism, KAS III will have an optimum pH of 7 and an optimum temperature of 30-37 °C.[7] Each organism's inhibitors, optimum pH, and optimum temperatures will vary slightly. However, these numbers are fairly indicative of the enzyme's ideal environment in general.
Ketoacyl synthase family 2
All KS2 enzymes are produced by eukaryota, with nearly all from plants. The most common enzymes in this family are 3-ketoacyl-CoA synthases, fatty acid elongases and very long-chain fatty acid condensing enzymes. The most common general characterization for these enzymes is E.C. 2.3.1.-; however, some are defined as 2.3.1.119. Most enzymes in the KS2 family catalyze reactions to produce very long-chain fatty acids. KS2 can be divided into 10 subfamilies.[1]
3-Ketoacyl-CoA synthase I
3-Ketoacyl-CoA synthase I in Arabidopsis thaliana is involved in very long chain fatty acid synthesis, which plays a role in wax biosynthesis.[8] The enzyme catalyzes the following reaction:
very-long-chain acyl-CoA + malonyl-CoA ⇒ very-long-chain 3-oxoacyl-CoA + CoA + CO2[9]
It is an elongase that appears to be involved in the production of very-long-chain fatty acids that are 26 carbons and longer.[10] Mefluidide and perfluidone are selective inhibitors of this enzyme.[11]
Ketoacyl synthase family 3
The KS3 family is the largest family in the KS system, with 14 subfamilies. KS3 enzymes are primarily produced in bacteria, with a small number of eukaryotes and archaea. KSs in this family contain KS domains present in both Type I FASs and the modular Type I of PKSs. While there are many slightly different enzymes in this family, the two most common 3-ketoacyl-ACP synthase I and synthase II.[1]
3-Ketoacyl-ACP synthase I
3-Ketoacyl-ACP synthase I (E.C. 2.3.1.41) is involved in the process of chain-elongation in type II FAS. A consequence of not having this enzyme will be a deficit in unsaturated fatty acids. It uses fatty acyl thioesters of ACP and CoA as substrates and has a specificity close to that of beta-ketoacyl-ACP synthase II.[12]
Typically, this enzyme is used in condensation reactions, as well as decarboxylation and acyl group transfer.
The reaction proceeds as such:
- acyl-[acyl-carrier protein] + a malonyl-[acyl-carrier protein] → a 3-oxoacyl-[acyl-carrier protein] + CO2 + an [acyl-carrier protein]
In Escherichia coli, for example, this enzyme is used to construct fatty acyl chains through a three step Claisen condensation reaction. The reaction will start with a trans thioesterfication of the acyl primer substrate. The donor substrate is then decarboxylated, forming a carbanion internmediate, which will attack C1 of the primer substrate, and create the elongated acyl chain.[13]
There are a number of molecules known to be inhibitors of synthase I. For example, in certain cases, acyl-CoA itself inhibits the enzyme at high concentrations in Escherichia coli. Cerulenin is known to inhibit synthase I in Carthamus tinctorius, Spinacia oleracea, Brassica napus, Allium ampeloprasu, Streptococcus pneumoniae, Escherichia coli, Mycobacterium tuberculosis, and many more. In Mycobacterium tuberculosis, palmitoyl-CoA is an inhibitor, and thiolactomycin is as well in a number of organisms.[12]
The optimal pH range varies greatly from organism to organism, but overall tends to lie between 5.5-8.5. Optimal temperature is the same, with 20 °C on one end of the spectrum, but 37 °C on the other.
3-Ketoacyl-ACP synthase II
3-Ketoacyl-ACP synthase II[14] is involved in type II FAS that occurs in plants and bacteria. While very similar to beta-ketoacyl-ACP synthase I, there is a slight difference between the two. One main difference is that synthase II is able to easily use palmitoleoyl-ACP as a substrate, while synthase I cannot. This allows for the control of the temperature-dependent regulation of fatty-acid composition.[15]
The reaction proceeds as such:
- (Z)-hexadec-11-enoyl-[acyl-carrier protein] + malonyl-[acyl-carrier protein] → (Z)-3-oxooctadec-13-enoyl-[acyl-carrier protein] + CO2 + [acyl-carrier protein
In Streptococcus pneumoniae, for example, synthase II is used as an elongation condensing enzyme. It contains a catalytic triad of Cys134, His337, and His303, as well as Phe396 and a water molecule bound to the active site. The nucleophilic cysteine is required for acyl-enzyme formation, and is used in the overall condensation activity. His 337 is also used for the condensation activity, specifically the stabilization of the negative charge on the malonyl thioester carbonyl in the transition state. His303 is used to accelerate catalysis by deprotonating the water molecule to allow for a nucleophilic attack on malonate, thereby releasing bicarbonate. Phe396 acts as a gatekeeper controlling the order of substrate addition.[16]
There are a number of molecules known to inhibit this enzyme. For example, cerulenin inhibits synthase II in Spinacia oleracea, Allium ampelprasum, Escherichia coli, and Streptoccoccus pneumonia. In Escherichia coli, platensimycin, thiolactomycin, and iodoacetamide are also known inhibitors.[15]
Optimal pH range will vary depending on the organism. In Escherichia coli, the range is 5.5–6.1. In Streptoccoccus pneumoniae, 6.8–7, in Plasmodium falciparum 7.5, and in Spinacia oleracea, 8.1–8.5. Optimal Temperature will vary, but for the most part remain in the range of 30–37 °C.[15]
Ketoacyl synthase family 4
A majority of KS4 enzymes exist in eukaryotic organisms, while the remainder are from bacteria. These enzymes are normally classified as either chalcone synthases, stilbene synthases, or type III PKSs. Overall, there are 10 different subfamilies within KS4. Typically, KS4 members will have a Cys-His-Asn catalytic triad. Both chalcone synthases and stilbene synthases will catalyze the same acyl transfer, decarboxylation, and condensation steps as in KS1. However, they will also further cyclize and aromatize the reactions before the final chalcone product is formed.[1]
Chalcone synthase
Chalcone synthase (E.C. 2.3.1.74), also known as naringenin-chalcone synthase, is responsible for the reaction:
- 3 malonyl-CoA + 4-coumaroyl-CoA → 4 CoA + naringenin chalcone + 3 CO2
In Medicago sativa, for example, the reaction occurs over the course of a loading step, a decarboxylation step, and finally, an elongation step.[17]
A number of inhibitors include cerulenin in Sinapis alba, Daucus carota, and Phaseolus vulgaris, apigenin in Secale cereale and Avena sativa, and eriodictyol in Decale cereal, Daucus carota, and Xanthisma gracile.[17]
The optimum pH at which this enzyme can function varies between organisms, but typically is placed somewhere between 6 and 8. The same goes for optimal temperature at 30-45 °C.[17]
Ketoacyl synthase family 5
KS5 family members are all present in eukaryotic cells, mostly animals. Most of these enzymes can be classified as fatty acid elongases. These enzymes are known to be used in the elongation of very long-chain fatty acids. KS5 has 11 subfamilies. Little is yet known about the KS5 family. Currently, none of the specific enzymes have E.C. numbers. No catalytic triad residues have been confirmed. Conserved histidine and asparagine residues have been found, with the histidine in a membrane-spanning region. However, there are not yet conserved cysteine residues known.[1]
References
- ↑ Jump up to: 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Chen, Yingfei; Kelly, Erin E.; Masluk, Ryan P.; Nelson, Charles L.; Cantu, David C.; Reilly, Peter J. (2011-10-01). "Structural classification and properties of ketoacyl synthases" (in en). Protein Science 20 (10): 1659–1667. doi:10.1002/pro.712. ISSN 1469-896X. PMID 21830247.
- ↑ Schweizer, Eckhart; Hofmann, Jörg (2004-09-01). "Microbial Type I Fatty Acid Synthases (FAS): Major Players in a Network of Cellular FAS Systems". Microbiology and Molecular Biology Reviews 68 (3): 501–517. doi:10.1128/MMBR.68.3.501-517.2004. ISSN 1092-2172. PMID 15353567.
- ↑ "Polyketide synthases". http://www.rasmusfrandsen.dk/polyketide_synthases.htm.
- ↑ Jump up to: 4.0 4.1 Hertweck, Christian (2009-06-15). "The Biosynthetic Logic of Polyketide Diversity" (in en). Angewandte Chemie International Edition 48 (26): 4688–4716. doi:10.1002/anie.200806121. ISSN 1521-3773. PMID 19514004.
- ↑ "ENZYME entry 2.3.1.180". expasy.org. http://enzyme.expasy.org/EC/2.3.1.180.
- ↑ Khandekar, SS; Gentry, DR; Van Aller, GS; Warren, P; Xiang, H; Silverman, C; Doyle, ML; Chambers, PA et al. (10 August 2001). "Identification, substrate specificity, and inhibition of the Streptococcus pneumoniae beta-ketoacyl-acyl carrier protein synthase III (FabH).". The Journal of Biological Chemistry 276 (32): 30024–30. doi:10.1074/jbc.M101769200. PMID 11375394.
- ↑ "BRENDA - Information on EC 2.3.1.180 - beta-ketoacyl-[acyl-carrier-protein synthase III"]. http://www.brenda-enzymes.org/enzyme.php?ecno=2.3.1.180.
- ↑ Todd, J.; Post-Beittenmiller, D.; Jaworski, J. G. (1999-01-01). "KCS1 encodes a fatty acid elongase 3-ketoacyl-CoA synthase affecting wax biosynthesis in Arabidopsis thaliana". The Plant Journal 17 (2): 119–130. doi:10.1046/j.1365-313x.1999.00352.x. ISSN 0960-7412. PMID 10074711.
- ↑ "KCS1 - 3-ketoacyl-CoA synthase 1 - Arabidopsis thaliana (Mouse-ear cress) - KCS1 gene & protein". https://www.uniprot.org/uniprot/Q9MAM3.
- ↑ Blacklock, Brenda J.; Jaworski, Jan G. (2006-07-28). "Substrate specificity of Arabidopsis 3-ketoacyl-CoA synthases". Biochemical and Biophysical Research Communications 346 (2): 583–590. doi:10.1016/j.bbrc.2006.05.162. PMID 16765910.
- ↑ Tresch, Stefan; Heilmann, Monika; Christiansen, Nicole; Looser, Ralf; Grossmann, Klaus (2012-04-01). "Inhibition of saturated very-long-chain fatty acid biosynthesis by mefluidide and perfluidone, selective inhibitors of 3-ketoacyl-CoA synthases". Phytochemistry 76: 162–171. doi:10.1016/j.phytochem.2011.12.023. ISSN 1873-3700. PMID 22284369.
- ↑ Jump up to: 12.0 12.1 "BRENDA - Information on EC 2.3.1.41 - beta-ketoacyl-[acyl-carrier-protein synthase I"]. http://www.brenda-enzymes.org/enzyme.php?ecno=2.3.1.41.
- ↑ von Wettstein-Knowles, Penny; Olsen, Johan G.; McGuire, Kirsten A.; Henriksen, Anette (2006-02-01). "Fatty acid synthesis" (in en). FEBS Journal 273 (4): 695–710. doi:10.1111/j.1742-4658.2005.05101.x. ISSN 1742-4658. PMID 16441657.
- ↑ "ENZYME entry 2.3.1.179". expasy.org. http://enzyme.expasy.org/EC/2.3.1.179.
- ↑ Jump up to: 15.0 15.1 15.2 "BRENDA - Information on EC 2.3.1.179 - beta-ketoacyl-[acyl-carrier-protein synthase II"]. http://www.brenda-enzymes.org/enzyme.php?ecno=2.3.1.179.
- ↑ Zhang, Yong-Mei; Hurlbert, Jason; White, Stephen W.; Rock, Charles O. (2006-06-23). "Roles of the Active Site Water, Histidine 303, and Phenylalanine 396 in the Catalytic Mechanism of the Elongation Condensing Enzyme of Streptococcus pneumoniae" (in en). Journal of Biological Chemistry 281 (25): 17390–17399. doi:10.1074/jbc.M513199200. ISSN 0021-9258. PMID 16618705.
- ↑ Jump up to: 17.0 17.1 17.2 "BRENDA - Information on EC 2.3.1.74 - naringenin-chalcone synthase". http://www.brenda-enzymes.org/enzyme.php?ecno=2.3.1.74.
 |