Biology:Methionine gamma-lyase
| methionine γ-lyase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Structure of a methionine gamma-lyase subunit, generated from 1GC2 | |||||||||
| Identifiers | |||||||||
| EC number | 4.4.1.11 | ||||||||
| CAS number | 42616-25-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
The enzyme methionine γ-lyase (EC 4.4.1.11, MGL) is in the γ-family of PLP-dependent enzymes. It degrades sulfur-containing amino acids to α-keto acids, ammonia, and thiols:
- L-methionine + H2O = methanethiol + NH3 + 2-oxobutanoate (overall reaction)
- (1a) L-methionine = methanethiol + 2-aminobut-2-enoate
- (1b) 2-aminobut-2-enoate = 2-iminobutanoate (spontaneous)
- (1c) 2-iminobutanoate + H2O = 2-oxobutanoate + NH3 (spontaneous)
Because sulfur-containing amino acids play a role in multiple biological processes, the regulation of these amino acids is essential. Additionally, it is crucial to maintain low homocysteine levels for the proper functioning of various pathways and for preventing the toxic effects of the cysteine homologue.[1] Methionine γ-lyase has been found in several bacteria (Clostridiums porogenes, Pseudomonas ovalis, Pseudomonas putida, Aeromonas sp., Citrobacter intermedius, Brevibacterium linens, Citrobacter freundii, Porphyromonas gingivalis, Treponema denticola), parasitic protozoa (Trichomonas vaginalis, Entamoeba histolytica), and plants (Arabidopsis thaliana).[2]
This enzyme belongs to the family of lyases, specifically the class of carbon-sulfur lyases. The systematic name of this enzyme class is L-methionine methanethiol-lyase (deaminating; 2-oxobutanoate-forming). Other names in common use include L-methioninase, methionine lyase, methioninase, methionine dethiomethylase, L-methionine γ-lyase, and L-methionine methanethiol-lyase (deaminating). This enzyme participates in selenoamino acid metabolism. It employs one cofactor, pyridoxal phosphate.
Structure
The enzyme is made up of 389-441 amino acids and forms four identical subunits. The active molecule is composed of two tightly associated dimers, the interface at which lies the active site. Each of the dimers has a pyridoxal 5’-phosphate (PLP) cofactor. Six amino acids located near the active site are involved in the reaction, namely Tyr59, Arg61, Tyr114, Cys116, Lys240, and Asp241.[2] Unlike the other amino acids, Cys116 is not typically found in PLP γ-family enzymes, which instead have glycine or proline. Although there is no direct contact between Cys116 and either MGL or the methionine substrate, studies show that the amino acid is involved in retaining substrate specificity.[3]
Reaction mechanism
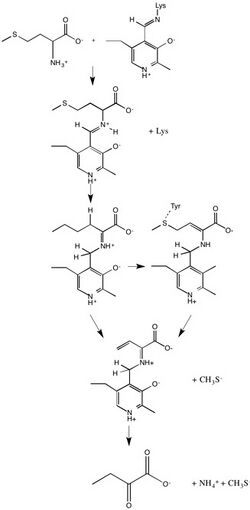
In enzymology, a methionine gamma-lyase (EC 4.4.1.11) is an enzyme that catalyzes the chemical reaction
- L-methionine + H2O [math]\displaystyle{ \rightleftharpoons }[/math] methanethiol + NH3 + 2-oxobutanoate
Thus, the two substrates of this enzyme are L-methionine and H2O, whereas its 3 products are methanethiol, NH3, and 2-oxobutanoate.
MGL also catalyzes α, β-elimination L-cysteine, degradation of O-substituted serine or homoserine, β- or γ-replacement, as well as deamination and γ-addition of L-vinylglycine. The reaction mechanism initially consists of the amino group of the substrate connected by a Schiff-base linkage to PLP. When a lysine residue replaces the amino group, an external aldimine is formed and hydrogens from the substrate are shifted to PLP. A neighboring tyrosine amino acid acts as an acid catalyst and attacks the substrate, consequently eliminating the thiol group from the substrate. Lastly, α-keto acid and ammonia are released from PLP.[2]
Function
Because MGL has differing substrate specificity among organisms, the enzyme also has varying physiological roles among organisms. In anaerobic bacteria and parasitic protozoa, MGL generates 2-oxobutyrate from methionine. 2-oxobutyrate is ultimately decomposed by acetate-CoA ligase and produces ATP, thus contributing to ATP metabolism. MGL also plays a role in the pathogenicity of periodontal bacterium such as P. gingivalis. A study finds a correlation between the presence of MGL and an increase in mice survival after subcutaneous injection of the bacteria. In B. linens, a cheese ripening bacterium, MGL activity is tightly linked with carbohydrate metabolism.[4]
In plants, MGL mRNA is found in dry seeds although the protein itself is not. However, the enzyme is highly expressed in wet seeds, suggesting that MGL is a vital part of early germination. MGL may also be involved in the formation of volatile sulfur compounds such as methanethiol on damaged plant leaves to defend against insects. However, it is undetermined whether MGL is present in guava, which was first discovered to have this protection mechanism, and whether other plants use a similar technique.[5]
Isozymes of MGL are only found in the parasitic protists E. histolytica and T. vaginalis. The isozymes differ in their ability to efficiently degrade methionine, homocysteine, and cysteine. E. histolytica MGL is derived from archaea MGL whereas T. vaginalis MGL share more similarities with bacterial MGL. Therefore, the inclusion of MGL in the genome of these two species occurred independently.[6]
Drug development
Trifluoromethionine (TFM) is a fluorinated methionine prodrug, which only presents its toxicity after degradation by MGL. Studies show that TFM is toxic to and slows the growth of anaerobic microorganisms (Mycobacterium smegmatis, Mycobacterium phlei, Candida lipolytica), periodontal bacteria (P. gingivalis, F. nucleatum), and parasitic protists (E. histolytica, T. vaginalis). Studies have shown that TFM is also efficacious in vivo. Furthermore, TFM has limited toxicity to mammalian cells, which do not have MGL. Therefore, TFM only exhibits toxic effects on pathogens that contain MGL.[7]
Cancer therapy
Some tumors, such as glioblastomas, medulloblastoma, and neuroblastoma, are much more sensitive to the methionine starvation than the normal tissues. Therefore, methionine depletion arises as a relevant therapeutical approach to treat cancer. For that reason, MGL has been studied to decrease the methionine levels in the blood serum and decrease the tumor growth and also to kill, by starvation, those malignant cells.[8]
References
- ↑ "Sulfur amino acid metabolism: pathways for production and removal of homocysteine and cysteine". Annual Review of Nutrition 24: 539–577. 2004. doi:10.1146/annurev.nutr.24.012003.132418. PMID 15189131.
- ↑ 2.0 2.1 2.2 2.3 "Methionine γ-lyase: the unique reaction mechanism, physiological roles, and therapeutic applications against infectious diseases and cancers". IUBMB Life 61 (11): 1019–1028. Nov 2009. doi:10.1002/iub.255. PMID 19859976.
- ↑ "The role of cysteine 116 in the active site of the antitumor enzyme L-methionine gamma-lyase from Pseudomonas putida". Bioscience, Biotechnology, and Biochemistry 72 (7): 1722–1730. Jul 2008. doi:10.1271/bbb.80015. PMID 18603802.
- ↑ Sulfur-containing amino acid metabolism in parasitic protozoa. 60. 2005. 1–99. doi:10.1016/S0065-308X(05)60001-2. ISBN 9780120317608.
- ↑ "Sulfur volatiles in guava (Psidium guajava L.) leaves: possible defense mechanism". Journal of Agricultural and Food Chemistry 56 (19): 8905–8910. Oct 2008. doi:10.1021/jf801735v. PMID 18778077.
- ↑ "Evolutionary relationships among pyridoxal-5'-phosphate-dependent enzymes. Regio-specific alpha, beta and gamma families". European Journal of Biochemistry 219 (3): 953–960. Feb 1994. doi:10.1111/j.1432-1033.1994.tb18577.x. PMID 8112347.
- ↑ "Trifluoromethionine, a prodrug designed against methionine gamma-lyase-containing pathogens, has efficacy in vitro and in vivo against Trichomonas vaginalis". Antimicrobial Agents and Chemotherapy 45 (6): 1743–1745. Jun 2001. doi:10.1128/AAC.45.6.1743-1745.2001. PMID 11353620.
- ↑ Fernandes, H. S.; Teixeira, C. S. Silva; Fernandes, P. A.; Ramos, M. J.; Cerqueira, N. M. F. S. A. (4 November 2016). "Amino acid deprivation using enzymes as a targeted therapy for cancer and viral infections". Expert Opinion on Therapeutic Patents 27 (3): 283–297. doi:10.1080/13543776.2017.1254194. ISSN 1354-3776. PMID 27813440.
Further reading
- "Current therapeutics, their problems, and sulfur-containing-amino-acid metabolism as a novel target against infections by "amitochondriate" protozoan parasites". Clinical Microbiology Reviews 20 (1): 164–187. Jan 2007. doi:10.1128/CMR.00019-06. PMID 17223627.
- "Diversity of L-methionine catabolism pathways in cheese-ripening bacteria". Applied and Environmental Microbiology 66 (12): 5514–5517. Dec 2000. doi:10.1128/aem.66.12.5514-5517.2000. PMID 11097940.
- "Isolation and purification of L-methionine-alpha-deamino-gamma-mercaptomethane-lyase (L-methioninase) from Clostridium sporogenes". Cancer Research 33 (8): 1862–5. Aug 1973. PMID 4720797.
- "Crystallization and preliminary X-ray analysis of L-methionine gamma-lyase 1 from Entamoeba histolytica". Acta Crystallographica Section F 64 (Pt 8): 697–699. Aug 2008. doi:10.1107/S1744309108018691. PMID 18678935.
- "Expression, purification and crystallization of L-methionine gamma-lyase 2 from Entamoeba histolytica". Acta Crystallographica Section F 62 (Pt 10): 1034–1036. Oct 2006. doi:10.1107/S1744309106036694. PMID 17012806.
- "Kinetic characterization of methionine gamma-lyases from the enteric protozoan parasite Entamoeba histolytica against physiological substrates and trifluoromethionine, a promising lead compound against amoebiasis". The FEBS Journal 275 (3): 548–560. Feb 2008. doi:10.1111/j.1742-4658.2007.06221.x. PMID 18199285.
- "Identification and characterization of two isoenzymes of methionine gamma-lyase from Entamoeba histolytica: a key enzyme of sulfur-amino acid degradation in an anaerobic parasitic protist that lacks forward and reverse trans-sulfuration pathways". The Journal of Biological Chemistry 278 (43): 42717–42727. Oct 2003. doi:10.1074/jbc.M212414200. PMID 12920135.
- "Homocysteine and cardiovascular disease: evidence on causality from a meta-analysis". BMJ 325 (7374): 1202–1206. Nov 2002. doi:10.1136/bmj.325.7374.1202. PMID 12446535.
 |

