Biology:Translatomics
Translatomics is the study of all open reading frames (ORFs) that are being actively translated in a cell or organism. This collection of ORFs is called the translatome. Characterizing a cell's translatome can give insight into the array of biological pathways that are active in the cell. According to the central dogma of molecular biology, the DNA in a cell is transcribed to produce RNA, which is then translated to produce a protein. Thousands of proteins are encoded in an organism's genome, and the proteins present in a cell cooperatively carry out many functions to support the life of the cell. Under various conditions, such as during stress or specific timepoints in development, the cell may require different biological pathways to be active, and therefore require a different collection of proteins. Depending on intrinsic and environmental conditions, the collection of proteins being made at one time varies. Translatomic techniques can be used to take a "snapshot" of this collection of actively translating ORFs, which can give information about which biological pathways the cell is activating under the present conditions.[1]
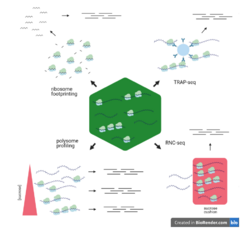
Usually, the ribosome profiling technique is used to acquire the translatome information.[2] Recent advancements, including single-cell ribosome profiling, have significantly improved the resolution of these studies, allowing researchers to gain insights into translation at the level of individual cells.[3] This is particularly important for heterogeneous cell populations, where overall bulk measurements may mask important cell-to-cell differences in protein synthesis. Other methods are polysome profiling, full-length translating mRNA profiling (RNC-seq), and translating ribosome affinity purification (TRAP-seq).[4] Unlike the transcriptome, the translatome is a more accurate approximation for estimating the expression level of some genes, since the correlation between the proteome and translatome is higher than the correlation between the transcriptome and proteome.[5]
History
Nearing the completion of the Human Genome Project the field of genetics was shifting its focus toward determining the functions of genes. This involved cataloguing other collections of biological materials, like RNA and proteins in cells. These collections of materials were called -omes, evoking the widespread excitement surrounding the sequencing of the human genome. The term translatome was first proposed in 2001 by Greenbaum et al.[6] The translatome was intended to describe the relative quantities of proteins in a proteome. The term translatome now generally refers to the collection of proteins actively being created in a cell. Translatomics, in combination with degradomics, aims to describe the net change to the proteome under different conditions.
Relevance and contribution to omics
The aim of genomics is to study the genome, or the collection of genetic material in an organism. Genomics subfields, or other -omics, such as Transcriptomics and proteomics aim to characterize genome function by quantifying products of the genome (such as RNA and proteins) under different conditions. In doing so, omics gain insight into different levels of regulation of gene expression and therefore genome function. However, these fields characterize biomolecules that have already been formed. In some cases, RNA or protein abundance does not reflect function because these biomolecules may be degraded rapidly, or they may remain in a cell long after they are initially synthesized. When using proteomics techniques to study the proteome, regulation of protein abundance at the level of post-translational modification and protein degradation may obscure earlier regulatory processes. Because cellular functions are often regulated at the level of translation, meaning the transcriptome does not always reflect genome function,[1] using translatomics techniques to study the translatome may allow one to observe regulation of genome function that would be obscured in transcriptomics or proteomics studies.
Methods
Translating mRNA Messenger RNA
Most translatomics techniques focus on characterizing the mRNAs that are complexed with ribosomes and therefore being translated.
Polysome profiling
Polysome profiling is a technique used to characterize the degree of translation of one or more mRNAs. A highly translated mRNA exists as a polysome, meaning it is complexed with multiple ribosomes. mRNAs translated at lower levels are complexed with fewer ribosomes. In polysome profiling, a sucrose gradient is used to separate molecular complexes in a cell lysate based on size.[7] The fractions from the column are analyzed by sequencing or other methods. The translation rate of mRNAs is determined based on its detection and abundance in the fractions of lower and higher molecular weight.
RNC-seq
The full length translating mRNA (RNC-seq) involves centrifugation of lysated sample on a sucrose cushion. This allows separation of the Ribosome-nascent chain complex(RNC) from free mRNAs and other cell components. The RNCs form a pellet in the centrifugation that is collected for further analysis. The mRNA being translated in these RNCs can be sequenced, allowing identification and quantification of the mRNAs being translated at the time.[1][8] However, RNC-mRNA complexes are fragile which can lead to ribosomes to dissociate from the mRNAs and degradation of mRNAs, potentially biasing the collected results.[1]
Ribo-seq/Ribosome profiling/Ribosome footprinting
In Ribosome profiling, cellular mRNA including polysomes is subjected to ribonucleases, enzymes that cleave RNA. Those positions in the RNA molecules that are bound by ribosomes are protected against digestion. After cessation of ribonuclease activity, these protected sites can be recovered and sequenced. The sequences obtained from ribo-seq therefore represent fragments of mRNAs that were being actively translated.[9]
TRAP-seq
TRAP-seq is used to identify mRNAs being actively translated in a specific cell type within a tissue or other assortment of cells. The cell type of interest is engineered to express a ribosomal subunit fused to an epitope tag such as green fluorescent protein.[10] After cell lysis, antibodies targeting the epitope are used to isolate mRNAs that are bound to the ribosomes containing the fusion proteins. This RNA is then converted to cDNA and sequenced. This technique specifically identifies mRNAs that were being translated in the cell type of interest.
Nascent Polypeptides
Folding States
Mass Spectrometry
Mass spectrometry methods are unable to determine folding of nascent polypeptides. No current methods examine the folding states of nascent polypeptides globally in the cell. There are methods for examining the folding state of individual nascent polypeptides.[1]
One method uses a nonspecific protease to cleave the nascent peptide at a low temperature. The protease can cleave the unfolded, flexible regions, but cannot cut tightly folded regions. The products of the cleavage can then be separated and studied to determine the folded regions of the nascent peptide.[1][11]
Nuclear Magnetic Resonance
To analyze the full structure of a nascent polypeptide, nuclear magnetic resonance (NMR) is used. NMR allows a dynamic view of molecules in solution and so can be used on ribosome-nascent chain complexes (RNC). Labelling of ribosomes allows the NMR data to be filtered for suspected ribosome signal and identify the signal of the nascent polypeptide.[12]
Identification and Quantification
While the structure of nascent polypeptides is currently unexaminable on a global scale, identification and quantification is not.
Stable Isotope Labeling by Amino Acids in Cell Culture
One method uses a variant of Stable isotope labeling by amino acids in cell culture (SILAC). SILAC labels proteins with stable isotopes to allow quantification, comparing labelled and unlabeled peptides for quantification. Pulse SILAC (pSILAC), only allows peptides created during the pulse to be labelled. In theory, this allows a capture of only nascent peptides for quantification. SILAC, however, requires similar levels of labelled and unlabelled proteins for accurate quantification. As such, pSILAC pulses have to run much longer than the translation process, making quantification of nascent peptides inaccurate.[1][13]
Bio-Orthogonal/Quantitative Non-Canonical Amino acid Tagging
Bio-Orthogonal/Quantitative Non-Canonical Amino acid Tagging (BONCAT/QuaNCAT) uses azidohomoalanine (AHA) to tag proteins. This allows isolation of newly created proteins for MS.[14][15] However, using AHA requires predepletion of intracellular methionine and introduction of AHA, stressing the cell and potentially altering translation dynamics within. Similar to pSILAC, AHA methods require longer pulses, thus limiting its efficacy in quantifying nascent peptides.[1][13]
Puromycin-associated nascent chain proteomics
Puromycin-associated nascent chain proteomics (PUNCH-P) uses a puromycin-biotin label to capture nascent polypeptides for MS. Though it does not disturb the cell process, it is less sensitive than other methods in detecting nascent peptides.[1][13] In addition to these quantification methods, other MS based methods are being worked on.
mRNA Degradation
Degradation of mRNA also plays an important part in regulating the translation process.
To explore mechanisms of decay, genome-wide mapping of uncapped and cleaved transcripts (GMUCT), parallel analysis of RNA ends (PARE), and degradome sequencing use the T4 ligase of the Illumina sequencing platform to sequence decapped mRNAs. T4 ligase ligates on RNA with a free 5’ monophosphate. As mature mRNAs have a 5’ cap, they are not bound as substrates, leaving decapped and degrading mRNAs to be bound.[16]
5′-monophosphorylated ends sequencing (5Pseq) captures both capped and decapped sequences to allow sequencing of both mature mRNA and degraded products. This helps identify mRNA degradation products, and has uses in studying ribosome stalling.[1][17]
These methods study 5’ to 3’ degradation, miRNA-mediated cleavage, and nonsense mediated mRNA decay, but cannot measure 3’ to 5’ degradation and other degradation mechanisms.
In Vivo Translation Tracking
The techniques above require lysis of cells and thus cannot be performed in living cells. Single molecule fluorescence resonance energy transfer (smFRET) and Nascent chain tracking (NCT) use fluorescence to track translational activity. Both methods track elongation rates of the polypeptides on single mRNAs.[18][19] Neither technique, however, is capable of high throughput.[1]
tRNAome
Transfer RNA (tRNA) is an important part of Translation. tRNAs read the mRNA, bringing the amino acids the ribosome assembles into a polypeptide. As such, the abundances and types of tRNAs has a large effect on the speed of protein synthesis.[20] tRNAs can be very similar to other tRNAs, with some tRNA species only differing by a single nucleotide. This, coupled with similar secondary and tertiary structures makes separating different tRNA species difficult.[1]
Gel Electrophoresis
2D-Gel electrophoresis is a classic method used in separating tRNAs. Initially, the tRNAs are denatured in 7M urea and separated in the first gel dimension. 4M urea allows partial refolding for additional separation in the second gel dimension. This method has allowed separation into 48 sets in E. coli and 30 in B. subtilis but has limited resolution. Large numbers of different tRNA species cannot be fully separated by 2D-gel electrophoresis, with only 62 spots found for the 269 rat tRNAs.[21]
Liquid Chromatography
High performance liquid chromatography can be used to separate tRNAs based on aminoacylated tRNA isoacceptors. This method cannot fully separate the tRNA species and cannot distinguish between codons, though it still can find quantitative differences between different cell lines.[21]
Mass Spectrometry
Mass spectrometry (MS) can be used to separate tRNAs based on unique endonuclease digestion products.[22] This, however, has limited resolution with mixtures of 30 tRNA species and needs fractionation prior to MS in larger groups of tRNA. It also cannot be used to identify deNovo tRNA species as it requires prior knowledge of the digestion patterns of tRNA species.[21]
Microarrays
Hybridization based microarrays use the 3’CCA conserved sequence in tRNAs to attach a fluorescent probe. 70-80 nucleotide long probes, covering the length of the tRNA, are then used to bind tRNAs. tRNAs with at least 8 base differences are able to be distinguished with microarrays, but tRNAs with smaller differences bind to the same probe.[21]
Quantitative Reverse Transcription
Quantitative reverse transcription, real time PCR (qRT-PCR) is another way for identifying and quantifying the tRNAome. A primer for the conserved 3’CCA sequence is used for priming reverse transcription for all tRNA species. Heavy modification of the tRNA nucleotides and the stability of the tRNA molecule can be an issue, so high temperatures and demethylation have been used to counter these. Demethylation has also been used to counteract errors heavy methylation induces in sequencing.[1]
Ribo-seq like
Ribo-tRNA-seq has been developed to observe the role of tRNA more closely to translation. Similar to Ribo-seq, Ribo-tRNA-seq captures tRNA molecules in ribosomes for library preparation before sequencing[23]
See also
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 "Translatomics: The Global View of Translation". International Journal of Molecular Sciences 20 (1): 212. January 2019. doi:10.3390/ijms20010212. PMID 30626072.
- ↑ "Translatome profiling: methods for genome-scale analysis of mRNA translation". Briefings in Functional Genomics 15 (1): 22–31. January 2016. doi:10.1093/bfgp/elu045. PMID 25380596.
- ↑ Ozadam H, Tonn T, Han CM, Segura A, Hoskins I, Rao S (2023). "Single-cell quantification of ribosome occupancy in early mouse development.". Nature 618 (7967): 1057–1064. doi:10.1038/s41586-023-06228-9. PMID 37344592.
- ↑ "Translatomics: The Global View of Translation". International Journal of Molecular Sciences 20 (1): 212. January 2019. doi:10.3390/ijms20010212. PMID 30626072.
- ↑ "Ribosome profiling reveals translation control as a key mechanism generating differential gene expression in Trypanosoma cruzi". BMC Genomics 16 (1): 443. June 2015. doi:10.1186/s12864-015-1563-8. PMID 26054634.
- ↑ "Interrelating Different Types of Genomic Data, from Proteome to Secretome: 'Oming in on Function". Genome Research 11 (1): 1463–8. 2001. doi:10.1101/gr.207401. PMID 11544189.
- ↑ "Polysome profiling followed by RNA-seq of cardiac differentiation stages in hESCs". Scientific Data 5 (1): 180287. December 2018. doi:10.1038/sdata.2018.287. PMID 30512016. Bibcode: 2018NatSD...580287P.
- ↑ Zhang, Gong; Hubalewska, Magdalena; Ignatova, Zoya (2009-02-08). "Transient ribosomal attenuation coordinates protein synthesis and co-translational folding". Nature Structural & Molecular Biology 16 (3): 274–280. doi:10.1038/nsmb.1554. ISSN 1545-9985. PMID 19198590. https://pubmed.ncbi.nlm.nih.gov/19198590.
- ↑ "Accurate detection of short and long active ORFs using Ribo-seq data". Bioinformatics 36 (7): 2053–2059. April 2020. doi:10.1093/bioinformatics/btz878. PMID 31750902.
- ↑ "Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP)". Nature Protocols 9 (1): 1282–91. May 2014. doi:10.1038/nprot.2014.085. PMID 24810037.
- ↑ Zhang, Gong; Hubalewska, Magdalena; Ignatova, Zoya (2009-02-08). "Transient ribosomal attenuation coordinates protein synthesis and co-translational folding". Nature Structural & Molecular Biology 16 (3): 274–280. doi:10.1038/nsmb.1554. ISSN 1545-9985. PMID 19198590. https://pubmed.ncbi.nlm.nih.gov/19198590/.
- ↑ Deckert, Annika; Waudby, Christopher A.; Wlodarski, Tomasz; Wentink, Anne S.; Wang, Xiaolin; Kirkpatrick, John P.; Paton, Jack F. S.; Camilloni, Carlo et al. (2016-05-03). "Structural characterization of the interaction of α-synuclein nascent chains with the ribosomal surface and trigger factor". Proceedings of the National Academy of Sciences of the United States of America 113 (18): 5012–5017. doi:10.1073/pnas.1519124113. ISSN 0027-8424. PMID 27092002. Bibcode: 2016PNAS..113.5012D.
- ↑ 13.0 13.1 13.2 Aviner, Ranen; Geiger, Tamar; Elroy-Stein, Orna (2013-08-15). "Novel proteomic approach (PUNCH-P) reveals cell cycle-specific fluctuations in mRNA translation". Genes & Development 27 (16): 1834–1844. doi:10.1101/gad.219105.113. ISSN 0890-9369. PMID 23934657.
- ↑ Howden, Andrew J.M.; Geoghegan, Vincent; Katsch, Kristin; Efstathiou, Georgios; Bhushan, Bhaskar; Boutureira, Omar; Thomas, Benjamin; Trudgian, David C. et al. (2013-04-10). "QuaNCAT: quantitating proteome dynamics in primary cells". Nature Methods 10 (4): 343–346. doi:10.1038/nmeth.2401. ISSN 1548-7091. PMID 23474466.
- ↑ Dieterich, Daniela C; Lee, Jennifer J; Link, A James; Graumann, Johannes; Tirrell, David A; Schuman, Erin M (2007-03-15). "Labeling, detection and identification of newly synthesized proteomes with bioorthogonal non-canonical amino-acid tagging" (in en). Nature Protocols 2 (3): 532–540. doi:10.1038/nprot.2007.52. ISSN 1754-2189. PMID 17406607. http://www.nature.com/articles/nprot.2007.52.
- ↑ Willmann, Matthew R.; Berkowitz, Nathan D.; Gregory, Brian D. (2014-05-01). "Improved genome-wide mapping of uncapped and cleaved transcripts in eukaryotes--GMUCT 2.0". Methods 67 (1): 64–73. doi:10.1016/j.ymeth.2013.07.003. ISSN 1095-9130. PMID 23867340. https://pubmed.ncbi.nlm.nih.gov/23867340/.
- ↑ Pelechano, Vicent; Alepuz, Paula (2017-07-07). "eIF5A facilitates translation termination globally and promotes the elongation of many non polyproline-specific tripeptide sequences". Nucleic Acids Research 45 (12): 7326–7338. doi:10.1093/nar/gkx479. ISSN 1362-4962. PMID 28549188.
- ↑ Morisaki, Tatsuya; Lyon, Kenneth; DeLuca, Keith F.; DeLuca, Jennifer G.; English, Brian P.; Zhang, Zhengjian; Lavis, Luke D.; Grimm, Jonathan B. et al. (2016-06-17). "Real-time quantification of single RNA translation dynamics in living cells" (in en). Science 352 (6292): 1425–1429. doi:10.1126/science.aaf0899. ISSN 0036-8075. PMID 27313040. Bibcode: 2016Sci...352.1425M. https://www.science.org/doi/10.1126/science.aaf0899.
- ↑ Stevens, Benjamin; Chen, Chunlai; Farrell, Ian; Zhang, Haibo; Kaur, Jaskiran; Broitman, Steven L.; Smilansky, Zeev; Cooperman, Barry S. et al. (2012). "FRET-based identification of mRNAs undergoing translation". PLOS ONE 7 (5): e38344. doi:10.1371/journal.pone.0038344. ISSN 1932-6203. PMID 22693619. Bibcode: 2012PLoSO...738344S.
- ↑ Zhong, Jiayong; Xiao, Chuanle; Gu, Wei; Du, Gaofei; Sun, Xuesong; He, Qing-Yu; Zhang, Gong (2015-06-19). "Transfer RNAs Mediate the Rapid Adaptation of Escherichia coli to Oxidative Stress". PLOS Genetics 11 (6): e1005302. doi:10.1371/journal.pgen.1005302. ISSN 1553-7390. PMID 26090660.
- ↑ 21.0 21.1 21.2 21.3 Czech, Andreas; Fedyunin, Ivan; Zhang, Gong; Ignatova, Zoya (2010-10-01). "Silent mutations in sight: co-variations in tRNA abundance as a key to unravel consequences of silent mutations" (in en). Molecular BioSystems 6 (10): 1767–1772. doi:10.1039/C004796C. ISSN 1742-2051. PMID 20617253. https://pubs.rsc.org/en/content/articlelanding/2010/mb/c004796c.
- ↑ Hossain, Mahmud; Limbach, Patrick A. (2007-02-01). "Mass spectrometry-based detection of transfer RNAs by their signature endonuclease digestion products" (in en). RNA 13 (2): 295–303. doi:10.1261/rna.272507. ISSN 1355-8382. PMID 17194720. PMC 1781365. http://rnajournal.cshlp.org/content/13/2/295.
- ↑ Chen, Chien-Wen; Tanaka, Motomasa (2018-04-10). "Genome-wide Translation Profiling by Ribosome-Bound tRNA Capture". Cell Reports 23 (2): 608–621. doi:10.1016/j.celrep.2018.03.035. ISSN 2211-1247. PMID 29642016.
 |
