Physics:Quantitative proteomics
Quantitative proteomics is an analytical chemistry technique for determining the amount of proteins in a sample.[1][2][3][4][5] The methods for protein identification are identical to those used in general (i.e. qualitative) proteomics, but include quantification as an additional dimension. Rather than just providing lists of proteins identified in a certain sample, quantitative proteomics yields information about the physiological differences between two biological samples. For example, this approach can be used to compare samples from healthy and diseased patients. Quantitative proteomics is mainly performed by two-dimensional gel electrophoresis (2-DE), preparative native PAGE, or mass spectrometry (MS). However, a recent developed method of quantitative dot blot (QDB) analysis is able to measure both the absolute and relative quantity of an individual proteins in the sample in high throughput format, thus open a new direction for proteomic research. In contrast to 2-DE, which requires MS for the downstream protein identification, MS technology can identify and quantify the changes.
Quantification using spectrophotometry
The concentration of a certain protein in a sample may be determined using spectrophotometric procedures.[6] The concentration of a protein can be determined by measuring the OD at 280 nm on a spectrophotometer, which can be used with a standard curve assay to quantify the presence of tryptophan, tyrosine, and phenylalanine.[7] However, this method is not the most accurate because the composition of proteins can vary greatly and this method would not be able to quantify proteins that do not contain the aforementioned amino acids. This method is also inaccurate due to the possibility of nucleic acid contamination. Other more accurate spectrophotometric procedures for protein quantification include the Biuret, Lowry, BCA, and Bradford methods. An alternative method for label free protein quantification in clear liquid is cuvette-based SPR technique, that simultaneously measures the refractive index ranging 1.0 to 1.6 nD and concentration of the protein ranging from 0.5 μL to 2 mL in volume. This system consists of the calibrated optical filter with very high angular resolution and the interaction of light with this crystal forms a resonance at a wavelength which correlates to concentration and refractive index near the crystal.[8]
Quantification using two dimensional electrophoresis
Two-dimensional gel electrophoresis (2-DE) represents one of the main technologies for quantitative proteomics with advantages and disadvantages. 2-DE provides information about the protein quantity, charge, and mass of the intact protein. It has limitations for the analysis of proteins larger than 150 kDa or smaller than 5kDa and low solubility proteins. Quantitative MS has higher sensitivity but does not provide information about the intact protein.
Classical 2-DE based on post-electrophoretic dye staining has limitations: at least three technical replicates are required to verify the reproducibility. Difference gel electrophoresis (DIGE) uses fluorescence-based labeling of the proteins prior to separation has increased the precision of quantification as well as the sensitivity in the protein detection.{{citation needed|date=July 2014} urrent main approach for the 2-DE based study of proteomes.[citation needed]
Quantification using mass spectrometry

Mass spectrometry (MS) represents one of the main technologies for quantitative proteomics with advantages and disadvantages.[5] Quantitative MS has higher sensitivity but can provide only limited information about the intact protein. Quantitative MS has been used for both discovery and targeted proteomic analysis to understand global proteomic dynamics in populations of cells (bulk analysis)[10] or in individual cells (single-cell analysis).[11][12]
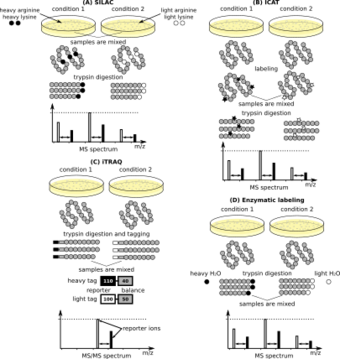
Early approaches developed in the 1990s applied isotope-coded affinity tags (ICAT), which uses two reagents with heavy and light isotopes, respectively, and a biotin affinity tag to modify cysteine containing peptides. This technology has been used to label whole Saccharomyces cerevisiae cells,[13] and, in conjunction with mass spectrometry, helped lay the foundation of quantitative proteomics. This approach has been superseded by isobaric mass tags,[10] which are also used for single-cell protein analysis.[14]
Relative and absolute quantification
Mass spectrometry is not inherently quantitative because of differences in the ionization efficiency and/or detectability of the many peptides in a given sample, which has sparked the development of methods to determine relative and absolute abundance of proteins in samples.[4][5] The intensity of a peak in a mass spectrum is not a good indicator of the amount of the analyte in the sample, although differences in peak intensity of the same analyte between multiple samples accurately reflect relative differences in its abundance.
Stable isotope labeling in mass spectrometry
Stable isotope labels
An approach for relative quantification that is more costly and time-consuming, though less sensitive to experimental bias than label-free quantification, entails labeling the samples with stable isotope labels that allow the mass spectrometer to distinguish between identical proteins in separate samples. One type of label, isotopic tags, consist of stable isotopes incorporated into protein crosslinkers that causes a known mass shift of the labeled protein or peptide in the mass spectrum. Differentially labeled samples are combined and analyzed together, and the differences in the peak intensities of the isotope pairs accurately reflect difference in the abundance of the corresponding proteins.
Absolute proteomic quantification using isotopic peptides entails spiking known concentrations of synthetic, heavy isotopologues of target peptides into an experimental sample and then performing LC-MS/MS. As with relative quantification using isotopic labels, peptides of equal chemistry co-elute and are analyzed by MS simultaneously. Unlike relative quantification, though, the abundance of the target peptide in the experimental sample is compared to that of the heavy peptide and back-calculated to the initial concentration of the standard using a pre-determined standard curve to yield the absolute quantification of the target peptide.
Relative quantification methods include isotope-coded affinity tags (ICAT), isobaric labeling (tandem mass tags (TMT) and isobaric tags for relative and absolute quantification (iTRAQ)), label-free quantification metal-coded tags (MeCAT), N-terminal labelling, stable isotope labeling with amino acids in cell culture (SILAC), and terminal amine isotopic labeling of substrates (TAILS). A mathematically rigorous approach that integrates peptide intensities and peptide-measurement agreement into confidence intervals for protein ratios has emerged.[15]
Absolute quantification is performed using selected reaction monitoring (SRM).
Metal-coded tags
Metal-coded tags (MeCAT) method is based on chemical labeling, but rather than using stable isotopes, different lanthanide ions in macrocyclic complexes are used. The quantitative information comes from inductively coupled plasma MS measurements of the labeled peptides. MeCAT can be used in combination with elemental mass spectrometry ICP-MS allowing first-time absolute quantification of the metal bound by MeCAT reagent to a protein or biomolecule. Thus it is possible to determine the absolute amount of protein down to attomole range using external calibration by metal standard solution. It is compatible to protein separation by 2D electrophoresis and chromatography in multiplex experiments. Protein identification and relative quantification can be performed by MALDI-MS/MS and ESI-MS/MS.
Mass spectrometers have a limited capacity to detect low-abundance peptides in samples with a high dynamic range. The limited duty cycle of mass spectrometers also restricts the collision rate, resulting in an undersampling.[16] Sample preparation protocols represent sources of experimental bias.
Stable isotope labeling with amino acids in cell culture
Stable isotope labeling with amino acids in cell culture (SILAC) is a method that involves metabolic incorporation of "heavy" C- or N-labeled amino acids into proteins followed by MS analysis. SILAC requires growing cells in specialized media supplemented with light or heavy forms of essential amino acids, lysine or arginine. One cell population is grown in media containing light amino acids while the experimental condition is grown in the presence of heavy amino acids. The heavy and light amino acids are incorporated into proteins through cellular protein synthesis. Following cell lysis, equal amounts of protein from both conditions are combined and subjected to proteotypic digestion. Arginine and lysine amino acids were chosen, because trypsin, the predominant enzyme used to generate proteotypic peptides for MS analysis, cleaves at the C-terminus of lysine and arginine. Following digestion with trypsin, all the tryptic peptides from cells grown in SILAC media would have at least one labeled amino acid, resulting in a constant mass shift from the labeled sample over non-labeled. Because the peptides containing heavy and light amino acids are chemically identical, they co-elute during reverse-phase column fractionation and are detected simultaneously during MS analysis. The relative protein abundance is determined by the relative peak intensities of the isotopically distinct peptides.
Traditionally the level of multiplexing in SILAC was limited due to the number of SILAC isotopes available. Recently, a new technique called NeuCode SILAC,[17] has augmented the level of multiplexing achievable with metabolic labeling (up to 4). The NeuCode amino acid method is similar to SILAC but differs in that the labeling only utilizes heavy amino acids. The use of only heavy amino acids eliminates the need for 100% incorporation of amino acids needed for SILAC. The increased multiplexing capability of NeuCode amino acids is from the use of mass defects from extra neutrons in the stable isotopes. These small mass differences however need to be resolved on high resolution mass spectrometers.
One of the main benefits of SILAC is the level of quantitation bias from processing errors is low because heavy and light samples are combined before sample preparation for MS analysis. SILAC and NeuCode SILAC are excellent techniques for detecting small changes in protein levels or post-translational modifications between experimental groups.
Isobaric labeling
Isobaric mass tags (tandem mass tags) are tags that have identical mass and chemical properties that allow heavy and light isotopologues to co-elute together. All mass tags consist of a mass reporter that has a unique number of 13C substitutions, a mass normalizer that has a unique mass that balances the mass of the tag to make all the tags equal in mass and a reactive moiety that crosslinks to the peptides. These tags are designed to cleave at a specific linker region upon high-energy CID, yielding different-sized tags that are then quantitated by LC-MS/MS. Protein or peptide samples prepared from cells, tissues or biological fluids are labeled in parallel with the isobaric mass tags and combined for analysis. Protein quantitation is accomplished by comparing the intensities of the reporter ions in the MS/MS spectra. Three types of tandem mass tags are available with different reactivity: (1) reactive NHS ester which provides high-efficiency, amine-specific labeling (TMTduplex, TMTsixplex, TMT10plex and TMT11plex), (2) reactive iodacetyl function group which labels sulfhydryl-(-SH) groups (iodoTMT) and (3) reactive alkoxyamine functional group which provides covalent labeling of carbonyl-containing compounds (aminoxyTMT).
A key benefit of isobaric labeling over other quantification techniques (e.g. SILAC, ICAT, Label-free) is the increased multiplex capabilities and thus increased throughput potential. The ability to combine and analyze several samples simultaneously in one LC-MS run eliminates the need to analyze multiple data sets and eliminates run-to-run variation. Multiplexing reduces sample processing variability, improves specificity by quantifying the proteins from each condition simultaneously, and reduces turnaround time for multiple samples. The current available isobaric chemical tags facilitate the simultaneous analysis of up to 11 experimental samples.
Label-free quantification in mass spectrometry
One approach for relative quantification is to separately analyze samples by MS and compare the spectra to determine peptide abundance in one sample relative to another, as in label-free strategies. It is generally accepted, that while label-free quantification is the least accurate of the quantification paradigms, it is also inexpensive and reliable when put under heavy statistical validation. There are two different methods of quantification in label-free quantitative proteomics: AUC (area under the curve) and spectral counting.
Methods of label-free quantification
AUC is a method by which for a given peptide spectrum in an LC-MS run, the area under the spectral peak is calculated. AUC peak measurements are linearly proportional to the concentration of protein in a given analyte mixture. Quantification is achieved through ion counts, the measurement of the amount of an ion at a specific retention time.[18] Discretion is required for the standardization of the raw data.[19] High-resolution spectrometer can alleviate problems that arise when trying to make data reproducible, however much of the work regarding normalizing data can be done through software such as OpenMS, and MassView.[20]
Spectral counting involves counting the spectra of an identified protein and then standardizing using some form of normalization.[21] Typically this is done with an abundant peptide mass selection (MS) that is then fragmented and then MS/MS spectra are counted.[18] Multiple samplings of the protein peak is required for accurate estimation of the protein abundance because of the complex physiochemical nature of peptides. Thus, optimization for MS/MS experiments is a constant concern. One alternative to get around this problems is use a data independent technique that cycles between high and low collision energies. Thus a large survey of all possible precursor and product ions is collected. This is limited, however, by the mass spectrometry software's ability to recognize and match peptide patterns of associations between the precursor and product ions.
Applications
File:Journal.pone.0095194.g001.TIF
Biomedical applications
Quantitative proteomics has distinct applications in the medical field. Especially in the fields of drug and biomarker discovery. LC-MS/MS techniques have started to over take more traditional methods like the western blot and ELISA due to the cumbersome nature of labeling different and separating proteins using these methods and the more global analysis of protein quantification. Mass spectrometry methods are more sensitive to difference in protein structure like post-translational modification and thus can quantify differing modifications to proteins. Quantitative proteomics can circumvent these issues, only needing sequence information to be performed. It can be applied on a global proteome level, or on specifically isolating binding partners in pull-down or affinity purification experiments.[5][23] Disadvantages, however, in sensitivity and analysis time must be kept in consideration.[24]
Drug discovery
Quantitative proteomics has the largest applications in the protein target identification, protein target validation, and toxicity profiling of drug discovery.[25] Drug discovery has been used to investigate protein-protein interaction and, more recently, drug-small molecule interactions, a field of study called chemoproteomics. Thus, it has shown great promise in monitoring side-effects of small drug-like molecules and understanding the efficacy and therapeutic effect of one drug target over another.[26][27] One of the more typical methodologies for absolute protein quantification in drug discovery is the use of LC-MS/MS with multiple reaction monitoring (MRM). The mass spectrometry is typically done by a triple quadrupole MS.[25]
See also
References
- ↑ Kastenholz, Bernd (2004). "Preparative Native Continuous Polyacrylamide Gel Electrophoresis (PNC-PAGE): An Efficient Method for Isolating Cadmium Cofactors in Biological Systems" (in en). Analytical Letters 37 (4): 657–665. doi:10.1081/AL-120029742. ISSN 0003-2719. http://www.tandfonline.com/doi/full/10.1081/AL-120029742.
- ↑ "Mass spectrometry-based proteomics turns quantitative". Nature Chemical Biology 1 (5): 252–62. October 2005. doi:10.1038/nchembio736. PMID 16408053. Bibcode: 2005NatCB...1..252O.
- ↑ "Quantitative mass spectrometry in proteomics: a critical review". Analytical and Bioanalytical Chemistry 389 (4): 1017–31. October 2007. doi:10.1007/s00216-007-1486-6. PMID 17668192. Bibcode: 2007ABiCh.389.1017B.
- ↑ 4.0 4.1 "Quantitative mass spectrometry-based proteomics: an overview". Quantitative Methods in Proteomics. Methods in Molecular Biology. 893. Humana Press. 2012. pp. 85–100. doi:10.1007/978-1-61779-885-6_7. ISBN 978-1-61779-884-9.
- ↑ 5.0 5.1 5.2 5.3 Rozanova, Svitlana; Barkovits, Katalin; Nikolov, Miroslav; Schmidt, Carla; Urlaub, Henning; Marcus, Katrin (2021). "Quantitative Mass Spectrometry-Based Proteomics: An Overview". in Marcus, Katrin; Eisenacher, Martin; Sitek, Barbara. Quantitative Methods in Proteomics. Methods in Molecular Biology. 2228. New York, NY: Springer US. pp. 85–116. doi:10.1007/978-1-0716-1024-4_8. ISBN 978-1-0716-1023-7.
- ↑ Ninfa, Ballou, Benore. Fundamental Approaches to Biochemistry and Biotechnology, 2nd edition 2010. Fitzgerald Science Press, Bethesda, MD.
- ↑ "An absolute method for protein determination based on difference in absorbance at 235 and 280 nm". Analytical Biochemistry 109 (1): 156–159. November 1980. doi:10.1016/0003-2697(80)90024-x. PMID 7469012. Bibcode: 1980AnBio.109..156W.
- ↑ Hermannsson, Pétur G.; Vannahme, Christoph; Smith, Cameron L. C.; Sørensen, Kristian T.; Kristensen, Anders (2015). "Refractive index dispersion sensing using an array of photonic crystal resonant reflectors". Applied Physics Letters 107 (6): 061101. doi:10.1063/1.4928548. Bibcode: 2015ApPhL.107f1101H. https://backend.orbit.dtu.dk/ws/files/118745939/1.4928548.pdf.
- ↑ "Technologies and challenges in large-scale phosphoproteomics". Proteomics 13 (6): 910–931. March 2013. doi:10.1002/pmic.201200484. PMID 23404676.
- ↑ 10.0 10.1 "Mass-spectrometric exploration of proteome structure and function". Nature 537 (7620): 347–355. September 2016. doi:10.1038/nature19949. PMID 27629641. Bibcode: 2016Natur.537..347A. http://www.nature.com/articles/nature19949.
- ↑ Specht, Harrison; Emmott, Edward; Petelski, Aleksandra; Huffman, R. Gray; Perlman, David H; Serra, Marco; Kharchenko, Peter; Koller, Antonius et al. (2021-01-27). "Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2". Genome Biology 22 (1): 50. doi:10.1186/s13059-021-02267-5. PMID 33504367.
- ↑ "Single-cell protein analysis by mass spectrometry". Current Opinion in Chemical Biology 60: 1–9. June 2020. doi:10.1016/j.cbpa.2020.04.018. PMID 32599342.
- ↑ "Accurate quantitation of protein expression and site-specific phosphorylation". Proceedings of the National Academy of Sciences of the United States of America 96 (12): 6591–6. June 1999. doi:10.1073/pnas.96.12.6591. PMID 10359756. Bibcode: 1999PNAS...96.6591O.
- ↑ "Unpicking the proteome in single cells". Science 367 (6477): 512–513. January 2020. doi:10.1126/science.aaz6695. PMID 32001644. Bibcode: 2020Sci...367..512S.
- ↑ "Bayesian Confidence Intervals for Multiplexed Proteomics Integrate Ion-statistics with Peptide Quantification Concordance*[S]". Molecular & Cellular Proteomics 18 (10): 2108–2120. 2019. doi:10.1074/mcp.TIR119.001317. PMID 31311848.
- ↑ "Assessing bias in experiment design for large scale mass spectrometry-based quantitative proteomics". Molecular & Cellular Proteomics 6 (10): 1741–8. October 2007. doi:10.1074/mcp.M600470-MCP200. PMID 17617667.
- ↑ "NeuCode labels for relative protein quantification". Molecular & Cellular Proteomics 13 (9): 2503–12. September 2014. doi:10.1074/mcp.M114.040287. PMID 24938287.
- ↑ 18.0 18.1 "Less label, more free: approaches in label-free quantitative mass spectrometry". Proteomics 11 (4): 535–53. February 2011. doi:10.1002/pmic.201000553. PMID 21243637.
- ↑ "Comparative LC-MS: a landscape of peaks and valleys". Proteomics 8 (4): 731–49. February 2008. doi:10.1002/pmic.200700694. PMID 18297651.
- ↑ "Quantification of proteins and metabolites by mass spectrometry without isotopic labeling or spiked standards". Analytical Chemistry 75 (18): 4818–26. September 2003. doi:10.1021/ac026468x. PMID 14674459. Bibcode: 2003AnaCh..75.4818W.
- ↑ "Role of spectral counting in quantitative proteomics". Expert Review of Proteomics 7 (1): 39–53. February 2010. doi:10.1586/epr.09.69. PMID 20121475.
- ↑ "Quantitative-proteomic comparison of alpha and Beta cells to uncover novel targets for lineage reprogramming". PLOS ONE 9 (4). 2014-04-23. doi:10.1371/journal.pone.0095194. PMID 24759943. Bibcode: 2014PLoSO...995194C.
- ↑ Shema-Yaacoby, Efrat; Nikolov, Miroslav; Haj-Yahya, Mahmood; Siman, Peter; Allemand, Eric; Yamaguchi, Yuki; Muchardt, Christian; Urlaub, Henning et al. (2013-08-15). "Systematic Identification of Proteins Binding to Chromatin-Embedded Ubiquitylated H2B Reveals Recruitment of SWI/SNF to Regulate Transcription" (in en). Cell Reports 4 (3): 601–608. doi:10.1016/j.celrep.2013.07.014. PMID 23933260.
- ↑ "Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present". Analytical and Bioanalytical Chemistry 404 (4): 939–965. September 2012. doi:10.1007/s00216-012-6203-4. PMID 22772140. Bibcode: 2012ABiCh.404..939B.
- ↑ 25.0 25.1 "Quantitative targeted absolute proteomics-based ADME research as a new path to drug discovery and development: methodology, advantages, strategy, and prospects". Journal of Pharmaceutical Sciences 100 (9): 3547–59. September 2011. doi:10.1002/jps.22612. PMID 21560129. Bibcode: 2011JPhmS.100.3547O.
- ↑ "Target profiling of small molecules by chemical proteomics". Nature Chemical Biology 5 (9): 616–24. September 2009. doi:10.1038/nchembio.216. PMID 19690537. Bibcode: 2009NatCB...5..616R.
- ↑ "Target identification and mechanism of action in chemical biology and drug discovery". Nature Chemical Biology 9 (4): 232–40. April 2013. doi:10.1038/nchembio.1199. PMID 23508189. Bibcode: 2013NatCB...9..232S.
 |
