Biology:Cohesin
Cohesin is a protein complex that mediates sister chromatid cohesion, homologous recombination, and DNA looping. Cohesin is formed of SMC3, SMC1, SCC1 and SCC3 (SA1 or SA2 in humans). Cohesin holds sister chromatids together after DNA replication until anaphase when removal of cohesin leads to separation of sister chromatids. The complex forms a ring-like structure and it is believed that sister chromatids are held together by entrapment inside the cohesin ring. Cohesin is a member of the SMC family of protein complexes which includes Condensin, MukBEF and SMC-ScpAB.
Cohesin was separately discovered in budding yeast (Saccharomyces cerevisiae) both by Douglas Koshland[1] and Kim Nasmyth in 1997.[2]
Structure
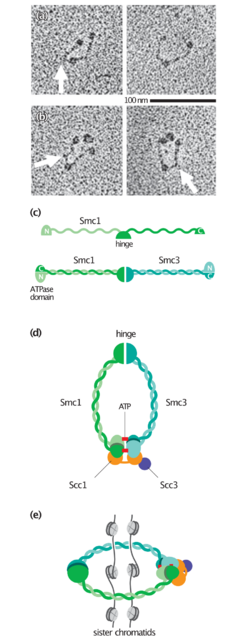
Cohesin is a multi-subunit protein complex, made up of SMC1, SMC3, RAD21 and SCC3 (SA1 or SA2).[3] SMC1 and SMC3 are members of the Structural Maintenance of Chromosomes (SMC) family. SMC proteins have two main structural characteristics: an ATP-binding cassette-like 'head' domain with ATPase activity (formed by the interaction of the N- and C- terminals) and a hinge domain that allows dimerization of SMCs. The head and the hinge domains are connected to each other via long anti-parallel coiled coils. The dimer is present in a V-shaped form, connected by the hinges.
The N-terminal domain of RAD21 contains two α-helices which forms a three helix bundle with the coiled coil of SMC3.[4] The central region of RAD21 is thought to be largely unstructured but contains several binding sites for regulators of cohesin. This includes a binding site for SA1 or SA2,[5] recognition motifs for separase cleavage[6] and a region that is competitively bound by PDS5A, PDS5B or NIPBL.[7][8][9] The C-terminal domain of RAD21 forms a winged helix that binds two β-sheets in the Smc1 head domain.[10]
Once RAD21 binds the SMC proteins, SCC3 can also associate with RAD21. When RAD21 binds on both SMC1 and SMC3, the cohesin complex forms a closed ring structure. The interfaces between the SMC subunits and RAD21 can open to allow DNA to pass in and out of the cohesin ring.[11]
While structures are available for many of the subunits and their interfaces, a structure of the entire cohesin complex has not been solved. Our knowledge of the conformation of cohesin comes largely from electron microscopy. These studies have revealed cohesin in numerous conformations including rings, elongated rods and most recently in a folded conformations. It is not known which conformation is predominant inside the cell and whether some are induced by sample preparation.[12]
Function
The cohesin complex regulates multiple vital cellular processes, such as:
- Sister chromatids cohesion during Mitosis and meiosis. It keeps the sister chromatids connected with each other during metaphase ensuring that, during cell division, each sister chromatid segregates to opposite poles. Without cohesin, the cell would be unable to control sister chromatid segregation since there would be no way of ensuring whether the spindle fiber attached on each sister chromatid is from a different pole.[13][14] Other proteins also regulate this process together with cohesin. These are PDS5A, PDS5B, NIPBL and ESCO1 in mammalian cells.[14]
- Bipolar spindle apparatus assembly during mitosis. Cohesin ensures the attachment of spindle microtubules and sister kinetochores onto chromosomes. This is tighly related with the correct sisters chromatids seggregation towards the two spindle poles. Dysregulation in this process leads to premature chromosomes separation and multipolar spindles formation.[15][16] The proteins Shugoshin 1 (or SGO1), Rae1 and NuMA are associated with cohesin in this assembly process.[17][18]
- DNA damage checkpoint and repair. It participates in repairing double-strand breaks in DNA via homologus recombination, where the sister chromatid is used as a template for sequence reconstruction.[19]
- Recently many novel functions of cohesin have been discovered in many different cellular processes. Cohesin has been shown to be responsible for transcription regulation, DNA double strand break repair, chromosome condensation, pairing of homologous chromosomes during meiosis I, mono-orientation of sister kinetochores during meiosis I, non-homologous centromere coupling, chromosome architecture and rearrangement, DNA replication etc.[20]
Dissociation of sister chromatid cohesion
The anaphase promoting complex associated to Cdc20 (APC/C-cdc20) marks Securin (anaphase inhibitor) for degradation by the proteasome. Securin is cleaved at anaphase, following APC/C-cdc20 mediated degradation, and it renders separase (a protease, inhibited by the association with securin) to cleave the kleisin subunit. An alpha-kleisin is associated with the cohesin complex, linking both SMC 3 and SMC 1 together, with the exact kleisin varying between mitosis and meiosis (Scc1 and Rec8 respectively), and its cleavage ultimately leads to the removal of cohesin from chromosomes.[21]
Dissociation of sister chromatids cohesion defines anaphase onset, which establishes two sets of identical chromosomes at each pole of the cell (telophase). Then the two daughter cells separate, and a new round of the cell cycle freshly starts in each one, at the stage of G0. When cells are ready to divide, because cell size is big enough or because they receive the appropriate stimulus,[22] they activate the mechanism to enter into the G1 stage of cell cycle, and they duplicate most organelles during S (synthesis) phase, including their centrosome. Therefore, when the cell division process will end, each daughter cell will receive a complete set of organelles. At the same time, during S phase all cells must duplicate their DNA very precisely, a process termed DNA replication. Once DNA replication has finished, in eukaryotes the DNA molecule is compacted and condensed, to form the mitotic chromosomes, each one constituted by two sister chromatids, which stay held together by the establishment of cohesion between them; each chromatid is a complete DNA molecule, attached via microtubules to one of the two centrosomes of the dividing cell, located at opposed poles of the cell. To avoid premature sister chromatid separation, the APC/C is maintained in an inactive state bound to different molecules, which are part of a complex mechanism termed the spindle assembly checkpoint.
Mechanism of Sister Chromatid Cohesion
It is not clear how the cohesin ring links sister chromatids together. There are two possible scenarios:
- Cohesin subunits bind to each sister chromatid and form a bridge between the two.
- Since cohesin has a ring structure, it is able to encircle both sister chromatids.
Current evidence suggests that the second scenario is the most likely. Proteins that are essential for sister chromatid cohesion, such as Smc3 and Scc1, do not regulate the formation of covalent bonds between cohesin and DNA, indicating that DNA interaction is not sufficient for cohesion.[11] In addition, disturbing the ring structure of cohesin through cleavage of Smc3 or Scc1 triggers premature sister chromatid segregation in vivo.[23] This shows that the ring structure is important for cohesin's function.
Early studies suggested various ways in which cohesin may entrap DNA,[24] including as a monomer that holds both homologues together, and a "hand-cuff" model where two intertwining cohesin complexes each hold one sister chromatid. While some studies support the idea of a hand-cuff model,[24] the model is inconsistent with a number of experimental observations,[25] and is generally considered to entrap chromatin as a monomer.
Even though the ring hypothesis appears to be valid, there are still questions about the number of rings required to hold sister chromatids together. One possibility is that one ring surrounds the two chromatids. Another possibility involves the creation of a dimer where each ring surrounds one sister chromatid. The two rings are connected to each other through formation of a bridge that holds the two sister chromatids together.
The topology and structure of these subunits has been best characterized in budding yeast,[26][27] but the sequence conservation of these proteins and biochemical and electron microscopic observations imply that cohesin complexes in other species are very similar in their structure, [1].
The cohesin complex is established during the initial stages of S-phase. The complexes associate with chromosomes before DNA replication occurs. Once cells start replicating their DNA, cohesin rings close and link the sister chromatids together.[11] Cohesin complexes must be present during S-phase in order for cohesion to take place. It is unclear, however, how cohesin is loaded on the chromosomes during G1. There are two proposed hypotheses so far:
- The ATPase domain of the SMC proteins interacts with DNA and this interaction initially mediates the loading of cohesin complexes on chromosomes.
- Several proteins aid in the loading process. For example, Scc2 and Scc4 are both required for cohesin to load in budding yeast.
Localization of cohesin rings
Cohesin binding along the chromosomal DNA is considered to be dynamic and its location changes based on gene transcription, specific DNA sequence and presence of chromosome-associated proteins. There are three possible scenarios:
- Cohesin location is influenced by the orientation of neighboring genes and it is most frequently located in areas of convergent transcription. Gene orientation depends on the direction of transcription and can be of three types: head-to-head, head-to-tail and tail-to-tail. The tail-to-tail configuration results in the convergence of transcription machinery. One hypothesis states that the RNA polymerase “pushes” cohesin along the DNA, causing them to move towards the direction of the RNA polymerases. Changing the transcription pattern of genes changes the location of cohesin indicating that localization of cohesin may depend on transcription.[28]
- In another model, chromatin loop extrusion is pushed by transcription generated supercoiling ensuring also that cohesin relocalizes quickly and loops grow with reasonable speed and in a good direction. In addition, the supercoiling-driven loop extrusion mechanism is consistent with earlier explanations proposing why topologically associating domains (TADs) flanked by convergent CTCF binding sites form more stable chromatin loops than TADs flanked by divergent CTCF binding sites. In this model, the supercoiling also stimulates enhancer promoter contacts and it is proposed that transcription of eRNA sends the first wave of supercoiling that can activate mRNA transcription in a given TAD.[29]
- A few cohesin rings are found in chromosome arms that have AT-rich DNA sequences indicating that DNA sequence may be an independent factor of cohesin binding.[28]
- Cohesin rings, especially in budding yeast, are also located in the region surrounding the centromere.[28] Two hypotheses may explain this: the presence of repetitive heterochromatic DNA in centromeres and the presence of chromosome-associated proteins. For example, Schizosaccharomyces pombe have multiple copies of specific heterochromatic DNA whose involvement in cohesion binding has been proven. Budding yeast lacks repetitive sequences and, therefore, requires a different mechanism for cohesion binding. Evidence suggests that binding of cohesin to the budding yeast centromere region depends on chromosome-associated proteins of the kinetochore that mediate cohesion association to pericentric regions (the kinetochore is an enhancer of pericentric cohesin binding).[30]
Cohesin and CTCF
Many chromatin loops are formed by so-called loop extrusion mechanism, when the cohesin ring moves actively along the two DNA double helices, translocating one of them with respect to the other. Thus, the loop can become smaller or larger. The loop extrusion process stops when cohesin encounters the architectural chromatin protein CTCF. The CTCF site needs to be in a proper orientation to stop cohesin.
Meiosis
Cohesin proteins SMC1β, SMC3, REC8 and STAG3 appear to participate in cohesion of sister chromatids throughout the meiotic process in human oocytes.[31] SMC1β, REC8 and STAG3 proteins are meiosis specific cohesins.
The STAG3 protein appears to be essential for female meiosis. A homozygous frameshift mutation in the Stag3 gene was identified in a large consanguineous family with premature ovarian failure.[32] Also, female mice deficient in STAG3 are sterile, and their fetal oocytes arrest at early prophase 1.
Evolution
Cohesin structure and function has been conserved in evolution. The SMC proteins are found in prokaryotes and have been conserved through evolution. The coils of SMC1 and SMC3 are conserved with an amino acid divergence of less than 0.5%.[33]
| Name | Saccharomyces cerevisiae | Schizosaccharomyces pombe | Drosophila | Vertebrates |
|---|---|---|---|---|
| Smc1 | Smc1 | Psm1 | DmSmc1 | Smc1 |
| Smc3 | Smc3 | Psm3 | DmSmc3 | Smc3 |
| Scc1 | Mcd1/Pds3 | Rad21 | DmRad21 | Rad21 |
| Scc3 | Scc3 | Psc3 | DmSA | SA1 and SA2 |
Clinical significance
The term "cohesinopathy" has been used to describe conditions affecting the cohesin complex.[34][35][36]
These conditions include:
- Cornelia de Lange Syndrome
- Roberts syndrome
- Warsaw breakage syndrome
- Many types of malignancies
See also
References
- ↑ Guacci, V; Koshland, D; Strunnikov, A (3 October 1997). "A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae.". Cell 91 (1): 47–57. doi:10.1016/s0092-8674(01)80008-8. PMID 9335334.
- ↑ Michaelis, C; Ciosk, R; Nasmyth, K (3 October 1997). "Cohesins: chromosomal proteins that prevent premature separation of sister chromatids.". Cell 91 (1): 35–45. doi:10.1016/s0092-8674(01)80007-6. PMID 9335333.
- ↑ Losada A, Hirano M, Hirano T (1998). "Identification of Xenopus SMC protein complexes required for sister chromatid cohesion". Genes Dev. 12 (13): 1986–1997. doi:10.1101/gad.12.13.1986. PMID 9649503.
- ↑ Gligoris, TG; Scheinost, JC; Bürmann, F; Petela, N; Chan, KL; Uluocak, P; Beckouët, F; Gruber, S et al. (21 November 2014). "Closing the cohesin ring: structure and function of its Smc3-kleisin interface.". Science 346 (6212): 963–7. doi:10.1126/science.1256917. PMID 25414305. Bibcode: 2014Sci...346..963G.
- ↑ Hara, K; Zheng, G; Qu, Q; Liu, H; Ouyang, Z; Chen, Z; Tomchick, DR; Yu, H (October 2014). "Structure of cohesin subcomplex pinpoints direct shugoshin-Wapl antagonism in centromeric cohesion.". Nature Structural & Molecular Biology 21 (10): 864–70. doi:10.1038/nsmb.2880. PMID 25173175.
- ↑ Uhlmann, F; Lottspeich, F; Nasmyth, K (1 July 1999). "Sister-chromatid separation at anaphase onset is promoted by cleavage of the cohesin subunit Scc1.". Nature 400 (6739): 37–42. doi:10.1038/21831. PMID 10403247. Bibcode: 1999Natur.400...37U.
- ↑ Petela, NJ; Gligoris, TG; Metson, J; Lee, BG; Voulgaris, M; Hu, B; Kikuchi, S; Chapard, C et al. (21 June 2018). "Scc2 Is a Potent Activator of Cohesin's ATPase that Promotes Loading by Binding Scc1 without Pds5.". Molecular Cell 70 (6): 1134–1148.e7. doi:10.1016/j.molcel.2018.05.022. PMID 29932904.
- ↑ Kikuchi, S; Borek, DM; Otwinowski, Z; Tomchick, DR; Yu, H (1 November 2016). "Crystal structure of the cohesin loader Scc2 and insight into cohesinopathy.". Proceedings of the National Academy of Sciences of the United States of America 113 (44): 12444–12449. doi:10.1073/pnas.1611333113. PMID 27791135. Bibcode: 2016PNAS..11312444K.
- ↑ Muir, KW; Kschonsak, M; Li, Y; Metz, J; Haering, CH; Panne, D (8 March 2016). "Structure of the Pds5-Scc1 Complex and Implications for Cohesin Function.". Cell Reports 14 (9): 2116–2126. doi:10.1016/j.celrep.2016.01.078. PMID 26923589.
- ↑ Haering, CH; Schoffnegger, D; Nishino, T; Helmhart, W; Nasmyth, K; Löwe, J (24 September 2004). "Structure and stability of cohesin's Smc1-kleisin interaction.". Molecular Cell 15 (6): 951–64. doi:10.1016/j.molcel.2004.08.030. PMID 15383284. http://www2.mrc-lmb.cam.ac.uk/groups/JYL/PDF/Haering%20Mol%20Cell%202004.pdf.
- ↑ 11.0 11.1 11.2 "Chromosomal cohesin forms a ring". Cell 112 (6): 765–77. March 2003. doi:10.1016/S0092-8674(03)00162-4. PMID 12654244.
- ↑ Yatskevich, S; Rhodes, J; Nasmyth, K (3 December 2019). "Organization of Chromosomal DNA by SMC Complexes.". Annual Review of Genetics 53: 445–482. doi:10.1146/annurev-genet-112618-043633. PMID 31577909.
- ↑ Nasmyth, Kim; Haering, Christian H. (2009). "Cohesin: its roles and mechanisms". Annual Review of Genetics 43: 525–558. doi:10.1146/annurev-genet-102108-134233. ISSN 1545-2948. PMID 19886810. https://pubmed.ncbi.nlm.nih.gov/19886810/.
- ↑ 14.0 14.1 Brooker, Amanda S.; Berkowitz, Karen M. (2014). "The roles of cohesins in mitosis, meiosis, and human health and disease". Cell Cycle Control. Methods in Molecular Biology (Clifton, N.J.). 1170. pp. 229–266. doi:10.1007/978-1-4939-0888-2_11. ISBN 978-1-4939-0887-5.
- ↑ Wong, Richard W.; Blobel, Günter (2008-10-07). "Cohesin subunit SMC1 associates with mitotic microtubules at the spindle pole". Proceedings of the National Academy of Sciences of the United States of America 105 (40): 15441–15445. doi:10.1073/pnas.0807660105. ISSN 1091-6490. PMID 18832153. Bibcode: 2008PNAS..10515441W.
- ↑ McNally, Karen P.; Beath, Elizabeth A.; Danlasky, Brennan M.; Barroso, Consuelo; Gong, Ting; Li, Wenzhe; Martinez-Perez, Enrique; McNally, Francis J. (October 2022). "Cohesin is required for meiotic spindle assembly independent of its role in cohesion in C. elegans". PLOS Genetics 18 (10): e1010136. doi:10.1371/journal.pgen.1010136. ISSN 1553-7404. PMID 36279281.
- ↑ McGuinness, Barry E.; Hirota, Toru; Kudo, Nobuaki R.; Peters, Jan-Michael; Nasmyth, Kim (March 2005). "Shugoshin prevents dissociation of cohesin from centromeres during mitosis in vertebrate cells". PLOS Biology 3 (3): e86. doi:10.1371/journal.pbio.0030086. ISSN 1545-7885. PMID 15737064.
- ↑ Kong, Xiangduo; Ball, Alexander R.; Sonoda, Eiichiro; Feng, Jie; Takeda, Shunichi; Fukagawa, Tatsuo; Yen, Tim J.; Yokomori, Kyoko (March 2009). "Cohesin associates with spindle poles in a mitosis-specific manner and functions in spindle assembly in vertebrate cells". Molecular Biology of the Cell 20 (5): 1289–1301. doi:10.1091/mbc.e08-04-0419. ISSN 1939-4586. PMID 19116315.
- ↑ Litwin, Ireneusz; Pilarczyk, Ewa; Wysocki, Robert (2018-11-28). "The Emerging Role of Cohesin in the DNA Damage Response". Genes 9 (12): 581. doi:10.3390/genes9120581. ISSN 2073-4425. PMID 30487431.
- ↑ "Cohesin: functions beyond sister chromatid cohesion". FEBS Letters 587 (15): 2299–312. August 2013. doi:10.1016/j.febslet.2013.06.035. PMID 23831059.
- ↑ "Cohesin: a guardian of genome integrity". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1823 (8): 1324–42. August 2012. doi:10.1016/j.bbamcr.2012.05.027. PMID 22677545.
- ↑ "Size control in animal development". Cell 96 (2): 235–44. January 1999. doi:10.1016/S0092-8674(00)80563-2. PMID 9988218.
- ↑ "The cohesin complex and its roles in chromosome biology". Genes & Development 22 (22): 3089–114. November 2008. doi:10.1101/gad.1724308. PMID 19056890.
- ↑ 24.0 24.1 "A handcuff model for the cohesin complex". The Journal of Cell Biology 183 (6): 1019–31. December 2008. doi:10.1083/jcb.200801157. PMID 19075111.
- ↑ "Cohesin: a catenase with separate entry and exit gates?". Nature Cell Biology 13 (10): 1170–7. October 2011. doi:10.1038/ncb2349. PMID 21968990.
- ↑ Haering, CH; Löwe, J; Hochwagen, A; Nasmyth, K (April 2002). "Molecular architecture of SMC proteins and the yeast cohesin complex.". Molecular Cell 9 (4): 773–88. doi:10.1016/s1097-2765(02)00515-4. PMID 11983169.
- ↑ Haering, CH; Farcas, AM; Arumugam, P; Metson, J; Nasmyth, K (17 July 2008). "The cohesin ring concatenates sister DNA molecules.". Nature 454 (7202): 297–301. doi:10.1038/nature07098. PMID 18596691. Bibcode: 2008Natur.454..297H. http://wrap.warwick.ac.uk/207/1/WRAP_Arumugam_resubmitted_cover_delete.pdf.
- ↑ 28.0 28.1 28.2 "Molecular biology: cohesins slip sliding away". Nature 430 (6999): 520–1. July 2004. doi:10.1038/430520b. PMID 15282594. Bibcode: 2004Natur.430..520R. https://zenodo.org/record/1233231.
- ↑ "Transcription-induced supercoiling as the driving force of chromatin loop extrusion during formation of TADs in interphase chromosomes". Nucleic Acids Res. 46 (4): 1648–1660. 13 November 2017. doi:10.1093/nar/gkx1123. PMID 29140466.
- ↑ "The kinetochore is an enhancer of pericentric cohesin binding". PLOS Biology 2 (9): E260. September 2004. doi:10.1371/journal.pbio.0020260. PMID 15309047.
- ↑ "Dynamics of cohesin proteins REC8, STAG3, SMC1 beta and SMC3 are consistent with a role in sister chromatid cohesion during meiosis in human oocytes". Human Reproduction 25 (9): 2316–27. September 2010. doi:10.1093/humrep/deq180. PMID 20634189.
- ↑ "Mutant cohesin in premature ovarian failure". The New England Journal of Medicine 370 (10): 943–949. March 2014. doi:10.1056/NEJMoa1309635. PMID 24597867.
- ↑ "The coiled coils of cohesin are conserved in animals, but not in yeast". PLOS ONE 4 (3): e4674. 2009. doi:10.1371/journal.pone.0004674. PMID 19262687. Bibcode: 2009PLoSO...4.4674W.
- ↑ "Cohesinopathy mutations disrupt the subnuclear organization of chromatin". The Journal of Cell Biology 187 (4): 455–62. November 2009. doi:10.1083/jcb.200906075. PMID 19948494.
- ↑ "Warsaw breakage syndrome, a cohesinopathy associated with mutations in the XPD helicase family member DDX11/ChlR1". American Journal of Human Genetics 86 (2): 262–6. February 2010. doi:10.1016/j.ajhg.2010.01.008. PMID 20137776.
- ↑ Warburton, Peter E., ed (September 2009). "The cellular phenotype of Roberts syndrome fibroblasts as revealed by ectopic expression of ESCO2". PLOS ONE 4 (9): e6936. doi:10.1371/journal.pone.0006936. PMID 19738907. Bibcode: 2009PLoSO...4.6936V.
Further reading
- "Cohesin: a guardian of genome integrity". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1823 (8): 1324–42. August 2012. doi:10.1016/j.bbamcr.2012.05.027. PMID 22677545.
- "Cohesin: functions beyond sister chromatid cohesion". FEBS Letters 587 (15): 2299–312. August 2013. doi:10.1016/j.febslet.2013.06.035. PMID 23831059.
- "Cohesins: chromosomal proteins that prevent premature separation of sister chromatids". Cell 91 (1): 35–45. October 1997. doi:10.1016/S0092-8674(01)80007-6. PMID 9335333.
- "A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae". Cell 91 (1): 47–57. October 1997. doi:10.1016/S0092-8674(01)80008-8. PMID 9335334.
- "Yeast cohesin complex requires a conserved protein, Eco1p(Ctf7), to establish cohesion between sister chromatids during DNA replication". Genes & Development 13 (3): 320–33. February 1999. doi:10.1101/gad.13.3.320. PMID 9990856.
- "Sister-chromatid separation at anaphase onset is promoted by cleavage of the cohesin subunit Scc1". Nature 400 (6739): 37–42. July 1999. doi:10.1038/21831. PMID 10403247. Bibcode: 1999Natur.400...37U.
External links
- cohesin at the US National Library of Medicine Medical Subject Headings (MeSH)
 |