Biology:Haplogroup O-M117
| Haplogroup O-M117 | |
|---|---|
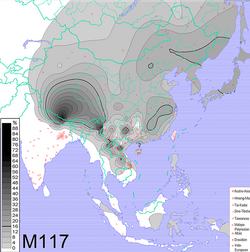 Interpolated frequency distribution[1] | |
| Possible time of origin | 18,203 [95% CI 16,626 <-> 19,783] years ago (Karmin 2015[2]) 17,430 ybp[3] 17,400 [95% CI 19,100 <-> 15,800] ybp (YFull[4]) |
| Coalescence age | 13,750 ybp[3] 12,600 [95% CI 11,300 <-> 14,000] ybp (YFull[4]) |
| Possible place of origin | probably East Asia or Southeast Asia[citation needed] |
| Ancestor | O-M134 |
| Descendants | O-M133 |
| Defining mutations | M117, Page23, CTS899/M1531, CTS1275/M1536, CTS3251, CTS5128/M1619, CTS6623/M1638, CTS11742/M1720, F141/M1564, F144, F235/M1587, F342/M1627, F373/M1636, F476/M1671, F579/M1692, F581, F584, F613/M1702, F649[citation needed] |
Haplogroup O2a2b1a1-M117 or Haplogroup O2a2b1a1-M117 (also defined by the phylogenetically equivalent mutation Page23/F8/F42) is a subclade of O2a2b1-M134 (and also a subclade of haplogroup O2-M122) that occurs frequently in China and in neighboring countries like Bhutan, Nepal, and Korea, also found among Sino-Tibetan language speaking people.
O2-M117 has been detected in samples of Tamang (38/45 = 84.4%), Tibetans (45/156 = 28.8% or 13/35 = 37.1%), Tharus (57/171 = 33.3%), Han Taiwanese (40/183 = 21.9%), Newars (14/66 = 21.2%), the general population of Kathmandu, Nepal (13/77 = 16.9%), Han Chinese (5/34 = 14.7% Chengdu, 5/35 = 14.3% Harbin, 4/35 = 11.4% Meixian, 3/30 = 10.0% Lanzhou, 2/32 = 6.3% Yili), Tungusic peoples from the PRC (7/45 = 15.6% Hezhe, 4/26 = 15.4% Evenki, 5/35 = 14.3% Manchu, 2/41 = 4.9% Xibe, 1/31 = 3.2% Oroqen), and Uyghurs (2/39 = 5.1% Yili, 1/31 = 3.2% Ürümqi) (Xue et al. 2006, Gayden et al. 2007, and Fornarino et al. 2009).
Like O-M7, O-M117 has been found with greatly varying frequency in many samples of Hmong-Mien-speaking peoples, such as Mienic peoples (7/20 = 35.0% Mountain Straggler Mien, 9/28 = 32.1% Blue Kimmun, 6/19 = 31.6% Flower Head Mien, 3/11 = 27.3% Top Board Mien, 3/11 = 27.3% Thin Board Mien, 11/47 = 23.4% Western Mien, 6/33 = 18.2% Northern Mien, 5/31 = 16.1% Lowland Yao, 5/35 = 14.3% Yao from Liannan, Guangdong, 5/37 = 13.5% Zaomin, 5/41 = 12.2% Lowland Kimmun, 3/41 = 7.3% Native Mien, 2/31 = 6.5% Southern Mien, 2/32 = 6.3% Mountain Kimmun, but 0/35 Yao from Bama, Guangxi), She (6/34 = 17.6% She, 4/56 = 7.1% Northern She), and Hmongic peoples (9/100 = 9.0% Miao from Hunan, 4/51 = 7.8% Hmong Daw from northern Laos, 3/49 = 6.1% Miao from Yunnan, 1/49 = 2.0% Miao from Guizhou, but 0/36 Bunu from Guangxi) (Cai et al. 2011 and Xue et al. 2006).
In Meghalaya, a predominantly tribal state of Northeast India, O-M133 has been found in 19.7% (14/71) of a sample of the Tibeto-Burman-speaking Garos, but in only 6.2% (22/353, ranging from 0/32 Bhoi to 6/44 = 13.6% Pnar) of a pool of eight samples of the neighboring Khasian-speaking tribes (Reddy et al. 2007).
Origin
The earliest attested genealogical split within haplogroup O-M117, that between O-M133 and O-M117(xM133), is estimated to have occurred approximately 12,600 [95% CI 11,300 <-> 14,000] ybp.[4] However, members of O-M117(xM133) are quite rare among extant humans. O-M117(xM133) has been observed in 2.2% (1/46) of the CHB (Han Chinese in Beijing, China) sample of the 1000 Genomes Project.[4] In commercial testing, O-MF1380 or O-CTS4960, which belongs to O-M117(xM133), has been found in China (Beijing, Henan, Shandong, Jiangsu, Anhui, Chongqing, Guangdong), Singapore, Indonesia, Saudi Arabia, and Japan.[4][5] O-M117(xM133) also has been found in 1.5% (2/133) of a sample collected in Daejeon, South Korea and in 1.0% (6/573) of a sample collected in Seoul, South Korea.[6] According to 23mofang, members of O-M117(xM133) comprise a subclade called O-CTS4960 (TMRCA 8,570 ybp), which is relatively concentrated in central, eastern, and northeastern areas of China and currently accounts for approximately 0.51% of the total population of males in China.[7]
The most recent common ancestor of all extant members of the O-M133 subclade, which predominates among extant members of O-M117, is estimated to have lived in a significantly less ancient era: 7,600 [95% CI 6,400 <-> 8,900] ybp according to YFull,[4] 7,455 [95% CI 6,514 <-> 8,500] years ago according to Karmin et al. 2015,[2] or 7,500 or 6,400 years ago (depending on which estimate of the mutation rate is used) according to Poznik et al. 2016.[8]
Distribution
China
Haplogroup O-M117 or O-M133 has been found often in samples of Han Chinese from various parts of China: 10/34 = 29.4% O-M133 Hakka in Taiwan,[9] 57/258 = 22.1% O-M133 miscellaneous Han volunteers in Taiwan,[9] 4/19 = 21.1% Fujian (CHS),[4] 12/60 = 20.0% O-M133 Minnan in Taiwan,[9] 29/167 = 17.4% East China,[10] 21/129 = 16.3% North China,[10] 7/46 = 15.2% Beijing (CHB),[8] 5/34 = 14.7% Chengdu,[11] 5/35 = 14.3% Harbin,[11] 9/65 = 13.8% South China,[10] 7/55 = 12.7% O-M133 Fujian,[9] 4/35 = 11.4% Meixian,[11] 75/689 = 10.9% Pudong,[12] 3/30 = 10.0% Lanzhou,[11] 50/530 = 9.4% Chongming Island,[12] 2/32 = 6.3% Yili,[11] 1/37 = 2.7% Hunan (CHS).[4]
Members of haplogroup O-M117 also have been found among various ethnic minorities in China, such as Tibetans (13/35 = 37.1%,[11] 45/156 = 28.8%[13]), Dai (13/52 = 25.0% CDX, or Chinese Dai in Xishuangbanna),[4] She people (6/34 = 17.6%[11]), Koreans (4/25 = 16.0% Koreans in the PRC[11]), Hezhe (7/45 = 15.6%[11]), Evenks (4/26 = 15.4%[11]), Manchu (5/35 = 14.3%[11]), Yao in Liannan, Guangdong (5/35 = 14.3%[11]), Mongols (5/45 = 11.1% Inner Mongolian[11]), Qiang (3/33 = 9.1%[11]), Daurs (3/39 = 7.7% Daur[11]), Hani (2/34 = 5.9%[11]), Xibe (2/41 = 4.9%[11]), Uyghurs (3/70 = 4.3%[11]), Oroqen (1/31 = 3.2%[11]), Buyi (1/35 = 2.9%[11]), and Hui (1/35 = 2.9%[11]).
Yan et al. (2014) have estimated that 16% of the present Han Chinese should be patrilineal descendants of a certain ancestor belonging to haplogroup O-M117 who has initiated a star-like population expansion dated to the Late Neolithic (5,400 [95% CI 4,100 <-> 6,700] years before present), which the authors have dubbed "Oα."[14]
According to 23mofang, haplogroup O-M117 (TMRCA 13,750 years) accounts for about 16.27% of the total male population of China,[15] with most members of O-M117 belonging to its O-F8 subclade (TMRCA 7,280 years), this latter subclade accounting for the Y-DNA of about 15.71% of all present-day Chinese males.[3][16]
India
In a study of the DNA of Adivasi populations in the state of Meghalaya, Reddy et al. (2007) found O-M133 in 19.7% (14/71) Garo, 13.6% (6/44) Pnar, 11.1% (2/18) Nongtrai, 8.3% (5/60) Lyngngam, 6.9% (2/29) War-Khasi, 6.3% (4/64) Maram, 5.3% (1/19) War-Jaintia, 2.3% (2/87) Khynriam, and 0% (0/32) Bhoi. The Garo natively speak the Garo language, whereas all the other studied populations natively speak Khasic languages.[17]
In another study that included populations in Meghalaya, Kumar et al. (2007) found O-M133 in 9.8% (9/92) Khasi and 9.1% (3/33) Garo.[18]
A study of populations of northern West Bengal and Sikkim published in 2011 found O-M117 in 57.7% (15/26) Rabha, 47.4% (9/19) Mech, 43.1% (22/51) Rajbanshi, 41.7% (15/36) Dhimal, and 7.4% (4/54) Bengali from the northern panhandle of West Bengal and in 9.1% (1/11) of a sample of Lachungpa from Sikkim. O-M117 was not found in this study's samples of Kol (0/62), Santhal (0/51), Kharia (0/34), or Oraon (0/31) from the northern panhandle of West Bengal.[19]
Japan
A study published in the year 2000 found O-M117 in 4.3% (1/23) of a sample representing Japan.[20] In a study published by Chinese researchers in the year 2006, O-M117 was found with high frequency (8/47 = 17.0%) in a sample of Japanese that should be from Kagawa Prefecture according to the geographical coordinates (134.0°E, 34.2°N) that have been provided (Xue et al. 2006). However, in a study published by Japanese researchers in the year 2007, the same haplogroup was found with much lower frequency (11/263 = 4.2%) in a larger sample of Japanese from various regions of Japan (Nonaka et al. 2007). (More precisely, Nonaka et al. have found O-M117 in 1/12 = 8.3% of a sample from Shizuoka, 4/52 = 7.7% of a sample from Tokyo, 2/44 = 4.5% of a sample from Chiba, 1/2 of a sample from Gifu, 1/2 of a sample from Yamanashi, 1/3 of a sample from Hiroshima, and 1/6 of a sample from Aichi.) O-M117 has been found in 8.8% (5/57) of the JPT (Japanese in Tokyo, Japan) sample of the 1000 Genomes Project.[8][21]
Korea
Between 11% and 15% of males in samples collected in South Korea have been found to belong to haplogroup O-M117 or O-M133 (20/133 = 15.0% Koreans in Daejeon,[6] 70/573 = 12.2% Koreans in Seoul,[6] 5/43 = 11.6% Koreans in South Korea,[11] 33/300 = 11.0% O-M133 Koreans[22]).
Mongolia
Haplogroup O-M117 has been found in about 5% of samples of Mongols in Mongolia: 4/20 = 20.0% NE Mongolia,[23] 1/18 = 5.6% central Mongolia,[23] 3/65 = 4.6% Outer Mongolian,[11] 1/23 = 4.3% SE Mongolia,[23] 3/97 = 3.1% NW Mongolia.[23]
Nepal
Haplogroup O-M117 has been found in 84.4% (38/45) of a sample of Tamang, 33.3% of sample of Tharu of Chitwan and Morang , 21.2% (14/66) of a sample of Newar, and 16.9% (13/77) of a sample of the general population of Kathmandu.[13]
Laos
In a study published in 2011, haplogroup O-M117 has been found in 7.8% (4/51) of a sample of Hmong Daw in Laos and in 5.1% (37/728) of a set of ethnic minorities who speak various Austroasiatic languages: 32.1% (9/28) Bit, 16.2% (6/37) Kataang, 14.0% (7/50) Mal, 13.7% (7/51) Khmu, 6.9% (2/29) Xinhmul, 3.3% (1/30) Alak, 2.94% (1/34) Inh, 2.86% (1/35) Talieng, 2.0% (1/50) Laven, 2.0% (1/50) Oy, 2.0% (1/50) So, 0% (0/28) Bo, 0% (0/32) Brau, 0% (0/32) Jeh, 0% (0/35) Lamet, 0% (0/35) Ngeq, 0% (0/38) Aheu, 0% (0/39) Suy, and 0% (0/45) Katu.[24]
Kutanan et al. 2019 found O-F8/F42, which is currently considered to be phylogenetically equivalent to O-M133, in 25.0% (5/20) of a sample of Laotians from Luang Prabang and 5.0% (1/20) of a sample of Laotians from Vientiane.[25]
Thailand
In a study published in 2014, haplogroup O-M133 has been found in 13.3% (10/75) of a sample of the general population of Bangkok and in 3.7% (1/27) of a sample of Akka from Chiang Mai.[9]
Brunelli et al. (2017) have found O-M117 in 35.0% (7/20) of Shan, 22.4% (46/205) of Khon Mueang, 22.2% (4/18) of Mon, 20.0% (5/25) of Western Lawa, 17.6% (16/91) of Tai Lue, 16.7% (4/24) of Tai Khuen, 13.6% (9/66) of Tai Yuan, and 11.5% (3/26) of Tai Yong in Northern Thailand and in 31.6% (6/19) of Tai Yuan in Central Thailand.[26] However, in the same study, haplogroup O-M117 was not observed in a sample of 25 Eastern Lawa in Northern Thailand.[26]
Kutanan et al. (2019) have found O-F8/F42 (equivalent to O-M133) in 14.75% (131/888) of a pool of samples from Thailand, including 50.0% (9/18) Palaung in Northern Thailand, 38.9% (7/18) Shan in Northern Thailand, 33.3% (20/60) Khon Mueang in Northern Thailand, 31.0% (13/42) Karen in Northern Thailand, 28.6% (6/21) Nyahkur in Northeast Thailand, 23.5% (4/17) Kaleun, 17.1% (22/129) Thai (Siamese), 16.7% (5/30) Tai Lue in Northern Thailand, 16.7% (3/18) Nyaw in Northeast Thailand, 16.7% (3/18) Blang in Northern Thailand, 15.4% (4/26) Tai Yuan, 14.3% (15/105) Mon, 14.3% (5/35) Phuan, 11.8% (2/17) Soa, 11.8% (2/17) Tai Khün, 9.4% (3/32) Western Lawa, 8.3% (3/36) Black Tai, 6.5% (4/62) Lao Isan, and 5.6% (1/18) Khmu.[25]
Vietnam
Haplogroup O-M133 has been found in 4/46 = 8.7% of the KHV (Kinh in Ho Chi Minh City, Vietnam) sample of the 1000 Genomes Project.[8][4] Haplogroup O-M133 has been found in 1/24 = 4.17% of a sample of people in Hanoi, Vietnam.[9] A study published in 2011 found haplogroup O-M117 in 1/15 = 6.67% Kinh and 1/12 = 8.33% Muong.[24]
Macholdt et al. 2020 have found O-F8, which is currently considered to be phylogenetically equivalent to O-M1706 or O-M133, in 36.4% (12/33) of a sample of Hanhi from Mường Tè District, 22.2% (8/36) of a sample of Lachi from Hoàng Su Phì District, 14.9% (7/47) of a sample of Tay, 14.3% (3/21) of a sample of Phula from Xín Mần District, 12.9% (4/31) of a sample of Lahu from Mường Tè District, 8.3% (2/24) of a sample of Thai, 4.8% (2/42) of a sample of Kinh from Hanoi, 3.7% (1/27) of a sample of Giarai, 2.8% (1/36) of a sample of Pathen, and 2.7% (1/37) of a sample of Nung from Vietnam.[27] All members of O-F8 among the Hanhi and Lahu of Mường Tè District belonged to the O-F2137 subclade. One Tay individual from Mường Khương District belonged to the O-F155 subclade and one Tay individual from Tràng Định District belonged to the O-F317 subclade. All other members of O-F8 belonged to the O-F8(xF155, F2137, F317) paragroup. Only one individual in this study (a Tay from Đức Trọng District) has been assigned to O-P164(xF8, F46, F4110) and therefore potentially might belong to O-M117(xF8/M133).
Subclades
According to the ISOGG experiental tree, the subclades of O2ab1a1-M117 are shown below (Owen Lu et al. 2016):
- O2a2b1a1 (M117/Page23)
- O2a2b1a1a (M133)
- O2a2b1a1a1 (F438)
- O2a2b1a1a1a (Y17728)
- O2a2b1a1a1a1 (F155)
- O2a2b1a1a1a2 (F1754)
- O2a2b1a1a1a2a (F2137)
- O2a2b1a1a1a3 (Z25907)
- O2a2b1a1a2 (FGC23469)
- O2a2b1a1a2a (F310)
- O2a2b1a1a2a1 (F402)
- O2a2b1a1a2a1a (F1531)
- O2a2b1a1a2a1 (F402)
- O2a2b1a1a2a (F310)
- O2a2b1a1a1a (Y17728)
- O2a2b1a1a3 (CTS7634)
- O2a2b1a1a3a (F317)
- O2a2b1a1a3a1 (F3039)
- O2a2b1a1a3b (CTS5488)
- O2a2b1a1a3a (F317)
- O2a2b1a1a4 (Z25853)
- O2a2b1a1a4a (CTS5492)
- O2a2b1a1a4a1 (CTS6987)
- O2a2b1a1a4a (CTS5492)
- O2a2b1a1a5 (CTS10738/M1707)
- O2a2b1a1a5a (CTS9678)
- O2a2b1a1a5a1 (Z39663)
- O2a2b1a1a5b (A9457)
- O2a2b1a1a5a (CTS9678)
- O2a2b1a1a6 (CTS4658)
- O2a2b1a1a6a (CTS5308)
- O2a2b1a1a6b (Z25928)
- O2a2b1a1a6b1 (SK1730)
- O2a2b1a1a6b1a (Z26030)
- O2a2b1a1a6b1b (Z26010)
- O2a2b1a1a6b2 (A9462)
- O2a2b1a1a6b3 (B456)
- O2a2b1a1a6b1 (SK1730)
- O2a2b1a1a1 (F438)
- O2a2b1a1b (CTS4960)
- O2a2b1a1a (M133)
References
Citations
- ↑ O'Rourke, Dennis; Cai, Xiaoyun; Qin, Zhendong; Wen, Bo; Xu, Shuhua; Wang, Yi; Lu, Yan; Wei, Lanhai et al. (2011). "Human Migration through Bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum Revealed by Y Chromosomes". PLOS ONE 6 (8): e24282. doi:10.1371/journal.pone.0024282. ISSN 1932-6203. PMID 21904623. Bibcode: 2011PLoSO...624282C.
- ↑ 2.0 2.1 Karmin, MonikaExpression error: Unrecognized word "etal". (2015). "A recent bottleneck of Y chromosome diversity coincides with a global change in culture". Genome Research 25 (4): 459–466. doi:10.1101/gr.186684.114. PMID 25770088.
- ↑ 3.0 3.1 3.2 Phylogenetic tree of human Y-DNA at 23mofang
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 4.7 4.8 4.9 YFull Haplogroup YTree v6.03.46 at 31 July 2018
- ↑ Phylogenetic tree of haplogroup O-M117/O-Page23 according to Family Tree DNA
- ↑ 6.0 6.1 6.2 Jin Park, Myung; Young Lee, Hwan; Ick Yang, Woo; Shin, Kyoung-Jin (2012). "Understanding the Y chromosome variation in Korea—relevance of combined haplogroup and haplotype analyses". International Journal of Legal Medicine 126 (4): 589–599. doi:10.1007/s00414-012-0703-9. PMID 22569803.
- ↑ "O-Cts4960单倍群详情". https://www.23mofang.com/ancestry/ytree/O-CTS4960/detail.
- ↑ 8.0 8.1 8.2 8.3 Poznik, G. DavidExpression error: Unrecognized word "etal". (June 2016). "Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences". Nature Genetics 48 (6): 593–599. doi:10.1038/ng.3559. PMID 27111036.
- ↑ 9.0 9.1 9.2 9.3 9.4 9.5 Trejaut, Jean A; Poloni, Estella S; Yen, Ju-Chen; Lai, Ying-Hui; Loo, Jun-Hun; Lee, Chien-Liang; He, Chun-Lin; Lin, Marie (2014). "Taiwan Y-chromosomal DNA variation and its relationship with Island Southeast Asia". BMC Genetics 2014 (15): 77. doi:10.1186/1471-2156-15-77. PMID 24965575.
- ↑ 10.0 10.1 10.2 Yan, Shi; Wang, Chuan-Chao; Li, Hui; Li, Shi-Lin; Jin, Li (2011). "An updated tree of Y-chromosome Haplogroup O and revised phylogenetic positions of mutations P164 and PK4". European Journal of Human Genetics 19 (9): 1013–1015. doi:10.1038/ejhg.2011.64. PMID 21505448.
- ↑ 11.00 11.01 11.02 11.03 11.04 11.05 11.06 11.07 11.08 11.09 11.10 11.11 11.12 11.13 11.14 11.15 11.16 11.17 11.18 11.19 11.20 11.21 11.22 Xue, Yali; Zerjal, Tatiana; Bao, Weidong; Zhu, Suling; Shu, Qunfang; Xu, Jiujin; Du, Ruofu; Fu, Songbin et al. (April 2006). "Male Demography in East Asia: A North–South Contrast in Human Population Expansion Times". Genetics 172 (4): 2431–2439. doi:10.1534/genetics.105.054270. PMID 16489223.
- ↑ 12.0 12.1 Zhang, X.; Tang, Z.; Wang, B.; Zhou, X.; Zhou, L.; Zhang, G.; Tian, J.; Zhao, Y.; Yao, Z.; Tian, L.; et al. "Forensic Analysis and Genetic Structure Construction of Chinese Chongming Island Han Based on Y Chromosome STRs and SNPs." Genes 2022, 13, 1363. https://doi.org/10.3390/genes13081363
- ↑ 13.0 13.1 Gayden, Tenzin; Cadenas, Alicia M.; Regueiro, Maria; Singh, Nanda B.; Zhivotovsky, Lev A.; Underhill, Peter A.; Cavalli-Sforza, Luigi L.; Herrera, Rene J. (2007). "The Himalayas as a Directional Barrier to Gene Flow". American Journal of Human Genetics 80 (5): 884–894. doi:10.1086/516757. PMID 17436243.
- ↑ "Y chromosomes of 40% Chinese descend from three Neolithic super-grandfathers". PLOS ONE 9 (8): e105691. 2014. doi:10.1371/journal.pone.0105691. PMID 25170956. Bibcode: 2014PLoSO...9j5691Y.
- ↑ https://www.23mofang.com/ancestry/ytree/O-M117/detail
- ↑ https://www.23mofang.com/ancestry/ytree/O-F8/detail
- ↑ Reddy, BMExpression error: Unrecognized word "etal". (2007). "Austro-Asiatic Tribes of Northeast India Provide Hitherto Missing Genetic Link between South and Southeast Asia". PLOS ONE 2 (11): e1141. doi:10.1371/journal.pone.0001141. PMID 17989774. Bibcode: 2007PLoSO...2.1141R.
- ↑ Kumar, VikrantExpression error: Unrecognized word "etal". (2007). "Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations". BMC Evolutionary Biology 2007 (7): 47. doi:10.1186/1471-2148-7-47. PMID 17389048. Bibcode: 2007BMCEE...7...47K.
- ↑ Debnath, Monojit; Palanichamy, Malliya G; Mitra, Bikash; Jin, Jie-Qiong; Chaudhuri, Tapas K; Zhang, Ya-Ping (2011). "Y-chromosome haplogroup diversity in the sub-Himalayan Terai and Duars populations of East India". Journal of Human Genetics 56 (11): 765–771. doi:10.1038/jhg.2011.98. PMID 21900945.
- ↑ Peter A. Underhill, Peidong Shen, Alice A. Lin et al., "Y chromosome sequence variation and the history of human populations," Nature Genetics • Volume 26 • November 2000
- ↑ YFull Haplogroup YTree v5.08 at 14 November 2017
- ↑ Jin Park, Myung; Young Lee, Hwan; Young Kim, Na; Young Lee, Eun; Ick Yang, Woo; Shin, Kyoung-Jin (2013). "Y-SNP miniplexes for East Asian Y-chromosomal haplogroup determination in degraded DNA". Forensic Science International: Genetics 7 (1): 75–81. doi:10.1016/j.fsigen.2012.06.014. PMID 22818129.
- ↑ 23.0 23.1 23.2 23.3 Di Cristofaro, JExpression error: Unrecognized word "etal". (2013). "Afghan Hindu Kush: Where Eurasian Sub-Continent Gene Flows Converge". PLOS ONE 8 (10): e76748. doi:10.1371/journal.pone.0076748. PMID 24204668. Bibcode: 2013PLoSO...876748D.
- ↑ 24.0 24.1 Cai, XExpression error: Unrecognized word "etal". (2011). "Human Migration through Bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum Revealed by Y Chromosomes". PLOS ONE 6 (8): e24282. doi:10.1371/journal.pone.0024282. PMID 21904623. Bibcode: 2011PLoSO...624282C.
- ↑ 25.0 25.1 Wibhu Kutanan, Jatupol Kampuansai, Metawee Srikummool, Andrea Brunelli, Silvia Ghirotto, Leonardo Arias, Enrico Macholdt, Alexander Hübner, Roland Schröder, and Mark Stoneking (2019), "Contrasting paternal and maternal genetic histories of Thai and Lao populations."
- ↑ 26.0 26.1 Brunelli, AExpression error: Unrecognized word "etal". (2017). "Y chromosomal evidence on the origin of northern Thai people". PLOS ONE 12 (7): e0181935. doi:10.1371/journal.pone.0181935. PMID 28742125. Bibcode: 2017PLoSO..1281935B.
- ↑ Enrico Macholdt, Leonardo Arias, Nguyen Thuy Duong, et al. (2020), "The paternal and maternal genetic history of Vietnamese populations." European Journal of Human Genetics (2020) 28:636–645. https://doi.org/10.1038/s41431-019-0557-4
Sources
- Journal articles
- Black, M. L.; Dufall, K.; Wise, C.; Sullivan, S.; Bittles, A. H. (2006). "Genetic ancestries in northwest Cambodia". Annals of Human Biology 33 (5–6): 620–7. doi:10.1080/03014460600882561. PMID 17381059.
- Cai, Xiaoyun; Qin, Zhendong; Wen, Bo; Xu, Shuhua; Wang, Yi; Lu, Yan; Wei, Lanhai; Wang, Chuanchao et al. (2011). O'Rourke, Dennis. ed. "Human Migration through Bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum Revealed by Y Chromosomes". PLOS ONE 6 (8): e24282. doi:10.1371/journal.pone.0024282. PMID 21904623. Bibcode: 2011PLoSO...624282C.
- Cordaux, R.; Weiss, G; Saha, N; Stoneking, M (2004). "The Northeast Indian Passageway: A Barrier or Corridor for Human Migrations?". Molecular Biology and Evolution 21 (8): 1525–33. doi:10.1093/molbev/msh151. PMID 15128876.
- Gan, Rui-Jing; Pan, Shang-Ling; Mustavich, Laura F.; Qin, Zhen-Dong; Cai, Xiao-Yun; Qian, Ji; Liu, Cheng-Wu; Peng, Jun-Hua et al. (2008). "Pinghua population as an exception of Han Chinese's coherent genetic structure". Journal of Human Genetics 53 (4): 303–13. doi:10.1007/s10038-008-0250-x. PMID 18270655.
- Gayden, Tenzin; Cadenas, Alicia M.; Regueiro, Maria; Singh, Nanda B.; Zhivotovsky, Lev A.; Underhill, Peter A.; Cavalli-Sforza, Luigi L.; Herrera, Rene J. (2007). "The Himalayas as a Directional Barrier to Gene Flow". The American Journal of Human Genetics 80 (5): 884–94. doi:10.1086/516757. PMID 17436243.
- Hammer, Michael F.; Karafet, Tatiana M.; Park, Hwayong; Omoto, Keiichi; Harihara, Shinji; Stoneking, Mark; Horai, Satoshi (2005). "Dual origins of the Japanese: Common ground for hunter-gatherer and farmer Y chromosomes". Journal of Human Genetics 51 (1): 47–58. doi:10.1007/s10038-005-0322-0. PMID 16328082.
- He, Jun-Dong; Peng, Min-Sheng; Quang, Huy Ho; Dang, Khoa Pham; Trieu, An Vu; Wu, Shi-Fang; Jin, Jie-Qiong; Murphy, Robert W. et al. (2012). Kayser, Manfred. ed. "Patrilineal Perspective on the Austronesian Diffusion in Mainland Southeast Asia". PLOS ONE 7 (5): e36437. doi:10.1371/journal.pone.0036437. PMID 22586471. Bibcode: 2012PLoSO...736437H.
- Hurles, M; Sykes, B; Jobling, M; Forster, P (2005). "The Dual Origin of the Malagasy in Island Southeast Asia and East Africa: Evidence from Maternal and Paternal Lineages". The American Journal of Human Genetics 76 (5): 894–901. doi:10.1086/430051. PMID 15793703.
- Jin, Han-Jun; Tyler-Smith, Chris; Kim, Wook (2009). Batzer, Mark A. ed. "The Peopling of Korea Revealed by Analyses of Mitochondrial DNA and Y-Chromosomal Markers". PLOS ONE 4 (1): e4210. doi:10.1371/journal.pone.0004210. PMID 19148289. Bibcode: 2009PLoSO...4.4210J.
- Jing, Chen; Hui, LI; Zhen-Dong, QIN; Wen-Hong, LIU; Wei-Xiong, LIN; Rui-Xing, YIN; Li, JIN; Shang-Ling, PAN (2006). "Y-chromosome Genotyping and Genetic Structure of Zhuang Populations". Acta Genetica Sinica 33 (12): 1060–72. doi:10.1016/S0379-4172(06)60143-1. PMID 17185165.
- Karafet, Tatiana; Xu, Liping; Du, Ruofu; Wang, William; Feng, Shi; Wells, R.S.; Redd, Alan J.; Zegura, Stephen L. et al. (2001). "Paternal Population History of East Asia: Sources, Patterns, and Microevolutionary Processes". The American Journal of Human Genetics 69 (3): 615–28. doi:10.1086/323299. PMID 11481588.
- Karafet, Tatiana M.; Lansing, J. S.; Redd, Alan J.; Watkins, Joseph C.; Surata, S. P. K.; Arthawiguna, W. A.; Mayer, Laura; Bamshad, Michael et al. (2005). "Balinese Y-Chromosome Perspective on the Peopling of Indonesia: Genetic Contributions from Pre-Neolithic Hunter-Gatherers, Austronesian Farmers, and Indian Traders". Human Biology 77 (1): 93–114. doi:10.1353/hub.2005.0030. PMID 16114819.
- Katoh, Toru; Munkhbat, Batmunkh; Tounai, Kenichi; Mano, Shuhei; Ando, Harue; Oyungerel, Ganjuur; Chae, Gue-Tae; Han, Huun et al. (2005). "Genetic features of Mongolian ethnic groups revealed by Y-chromosomal analysis". Gene 346: 63–70. doi:10.1016/j.gene.2004.10.023. PMID 15716011.
- Kayser, M.; Brauer, S; Cordaux, R; Casto, A; Lao, O; Zhivotovsky, LA; Moyse-Faurie, C; Rutledge, RB et al. (2006). "Melanesian and Asian Origins of Polynesians: MtDNA and Y Chromosome Gradients Across the Pacific". Molecular Biology and Evolution 23 (11): 2234–44. doi:10.1093/molbev/msl093. PMID 16923821.
- Kharkov, V. N.; Stepanov, V. A.; Medvedeva, O. F.; Spiridonova, M. G.; Voevoda, M. I.; Tadinova, V. N.; Puzyrev, V. P. (2007). "Gene pool differences between Northern and Southern Altaians inferred from the data on Y-chromosomal haplogroups". Russian Journal of Genetics 43 (5): 551–562. doi:10.1134/S1022795407050110. PMID 17633562.
- Kim, Wook; Yoo, Tag-Keun; Kim, Sung-Joo; Shin, Dong-Jik; Tyler-Smith, Chris; Jin, Han-Jun; Kwak, Kyoung-Don; Kim, Eun-Tak et al. (2007). Blagosklonny, Mikhail. ed. "Lack of Association between Y-Chromosomal Haplogroups and Prostate Cancer in the Korean Population". PLOS ONE 2 (1): e172. doi:10.1371/journal.pone.0000172. PMID 17245448. Bibcode: 2007PLoSO...2..172K.
- Kumar, Vikrant; Reddy, Arimanda NS; Babu, Jagedeesh P; Rao, Tipirisetti N; Langstieh, Banrida T; Thangaraj, Kumarasamy; Reddy, Alla G; Singh, Lalji et al. (2007). "Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations". BMC Evolutionary Biology 7 (1): 47. doi:10.1186/1471-2148-7-47. PMID 17389048. Bibcode: 2007BMCEE...7...47K.
- Li, Hui; Wen, Bo; Chen, Shu-Juo; Su, Bing; Pramoonjago, Patcharin; Liu, Yangfan; Pan, Shangling; Qin, Zhendong et al. (2008). "Paternal genetic affinity between western Austronesians and Daic populations". BMC Evolutionary Biology 8 (1): 146. doi:10.1186/1471-2148-8-146. PMID 18482451. Bibcode: 2008BMCEE...8..146L.
- Nonaka, I.; Minaguchi, K.; Takezaki, N. (2007). "Y-chromosomal Binary Haplogroups in the Japanese Population and their Relationship to 16 Y-STR Polymorphisms". Annals of Human Genetics 71 (4): 480–95. doi:10.1111/j.1469-1809.2006.00343.x. PMID 17274803. http://ir.tdc.ac.jp/irucaa/bitstream/10130/491/1/j.1469-1809.2006.00343.x.pdf.
- Reddy, B. Mohan; Langstieh, B. T.; Kumar, Vikrant; Nagaraja, T.; Reddy, A. N. S.; Meka, Aruna; Reddy, A. G.; Thangaraj, K. et al. (2007). Awadalla, Philip. ed. "Austro-Asiatic Tribes of Northeast India Provide Hitherto Missing Genetic Link between South and Southeast Asia". PLOS ONE 2 (11): e1141. doi:10.1371/journal.pone.0001141. PMID 17989774. Bibcode: 2007PLoSO...2.1141R.
- Shi, Simona; Pala, Maria; Battaglia, Vincenza; Maranta, Ramona; Achilli, Alessandro; Modiano, Guido; Torroni, Antonio; Semino, Ornella et al. (2009). "Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): A reservoir of genetic variation". BMC Evolutionary Biology 9 (1): 154. doi:10.1186/1471-2148-9-154. PMID 19573232. Bibcode: 2009BMCEE...9..154F.
- Su, B.; Jin, L.; Underhill, P.; Martinson, J.; Saha, N.; McGarvey, S. T.; Shriver, M. D.; Chu, J. et al. (2000). "Polynesian origins: Insights from the Y chromosome". Proceedings of the National Academy of Sciences 97 (15): 8225–8228. doi:10.1073/pnas.97.15.8225. PMID 10899994. Bibcode: 2000PNAS...97.8225S. "*H6 (=O-M122(xO-M7, O-M134)) in 18/73=24.7% *H8 (=O-M134) in 2/73=2.7% for a total of 20/73=27.4% O-M122 in a pool of seven samples from Micronesia. *13/40=32.5% O-M122(xM7,M134) in a pool of three samples from Polynesia. *9/27=33.3% H6 (=O-M122(xM7,M134)) *6/27=22.2% H8 (=O-M134) for a total of 15/27=55.6% O-M122 in "Malay" sample *2/19=10.5% H6 (=O-M122(xM7,M134)) in "Kota Kinabalu" sample.".
- Su, Bing; Xiao, Junhua; Underhill, Peter; Deka, Ranjan; Zhang, Weiling; Akey, Joshua; Huang, Wei; Shen, Di et al. (1999). "Y-Chromosome Evidence for a Northward Migration of Modern Humans into Eastern Asia during the Last Ice Age". The American Journal of Human Genetics 65 (6): 1718–24. doi:10.1086/302680. PMID 10577926.
- Tajima, Atsushi; Hayami, Masanori; Tokunaga, Katsushi; Juji, Takeo; Matsuo, Masafumi; Marzuki, Sangkot; Omoto, Keiichi; Horai, Satoshi (2004). "Genetic origins of the Ainu inferred from combined DNA analyses of maternal and paternal lineages". Journal of Human Genetics 49 (4): 187–93. doi:10.1007/s10038-004-0131-x. PMID 14997363.
- Wang, Wei; Wise, Cheryl; Baric, Tom; Black, Michael L.; Bittles, Alan H. (August 1, 2003). "The origins and genetic structure of three co-resident Chinese Muslim populations: the Salar, Bo'an and Dongxiang". Human Genetics 113 (3): 244–52. doi:10.1007/s00439-003-0948-y. ISSN 0340-6717. PMID 12759817.
- Wells, R. S.; Yuldasheva, N.; Ruzibakiev, R.; Underhill, P. A.; Evseeva, I.; Blue-Smith, J.; Jin, L.; Su, B. et al. (2001). "The Eurasian Heartland: A continental perspective on Y-chromosome diversity". Proceedings of the National Academy of Sciences 98 (18): 10244–9. doi:10.1073/pnas.171305098. PMID 11526236. Bibcode: 2001PNAS...9810244W.
- Wen, Bo; Li, Hui; Lu, Daru; Song, Xiufeng; Zhang, Feng; He, Yungang; Li, Feng; Gao, Yang et al. (2004). "Genetic evidence supports demic diffusion of Han culture". Nature 431 (7006): 302–5. doi:10.1038/nature02878. PMID 15372031. Bibcode: 2004Natur.431..302W.
- Wen, Bo; Xie, Xuanhua; Gao, Song; Li, Hui; Shi, Hong; Song, Xiufeng; Qian, Tingzhi; Xiao, Chunjie et al. (2004). "Analyses of Genetic Structure of Tibeto-Burman Populations Reveals Sex-Biased Admixture in Southern Tibeto-Burmans". The American Journal of Human Genetics 74 (5): 856–865. doi:10.1086/386292. PMID 15042512.
- Wen, Bo; Hong, S; Ling, R; Huifeng, X; Kaiyuan, L; Wenyi, Z; Bing, S; Shiheng, S et al. (2004). "The origin of Mosuo people as revealed by mtDNA and Y chromosome variation". Science China Life Sciences 47 (1): 1–10. doi:10.1360/02yc0207. PMID 15382670.
- Xie, Xuan-Hua (2004). "Genetic Structure of Tujia as Revealed by Y Chromosomes". Yi Chuan Xue Bao = Acta Genetica Sinica 31 (10): 1023–9. PMID 15552034. http://en.cnki.com.cn/Article_en/CJFDTOTAL-YCXB200410000.htm.
- Xue, Y.; Zerjal, T; Bao, W; Zhu, S; Shu, Q; Xu, J; Du, R; Fu, S et al. (2005). "Male Demography in East Asia: A North-South Contrast in Human Population Expansion Times". Genetics 172 (4): 2431–9. doi:10.1534/genetics.105.054270. PMID 16489223.
- Yang, Zhili; Dong, Yongli; Gao, Lu; Cheng, Baowen; Yang, Jie; Zeng, Weimin; Lu, Jing; Su, Yanhua et al. (2005). "The distribution of Y chromosome haplogroups in the nationalities from Yunnan Province of China". Annals of Human Biology 32 (1): 80–7. doi:10.1080/03014460400027557. PMID 15788357.
- Zhou, Ruixia; Yang, Daqun; Zhang, Hua; Yu, Weiping; An, Lizhe; Wang, Xilong; Li, Hong; Xu, Jiujin et al. (2008). "Origin and evolution of two Yugur sub-clans in Northwest China: A case study in paternal genetic landscape". Annals of Human Biology 35 (2): 198–211. doi:10.1080/03014460801922927. PMID 18428013.
 |

