Chemistry:Pikromycin
| |
| Names | |
|---|---|
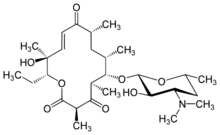
| IUPAC name
(3R,5R,6S,7S,9R,11E,13S,14R)-14-Ethyl-13-hydroxy-3,5,7,9,13-pentamethyl-6-[3,4,6-trideoxy-3-(dimethylamino)-β-D-xylo-hexopyranosyloxy]-1-oxacyclotetradec-11-ene-2,4,10-trione
| |
| Systematic IUPAC name
(3R,5R,6S,7S,9R,11E,13S,14R)-6-{[(2S,3R,4S,6R)-4-(Dimethylamino)-3-hydroxy-6-methyloxan-2-yl]oxy}-14-ethyl-13-hydroxy-3,5,7,9,13-pentamethyl-1-oxacyclotetradec-11-ene-2,4,10-trione | |
| Other names
Picromycin
| |
| Identifiers | |
3D model (JSmol)
|
|
| ChEBI | |
| ChemSpider | |
PubChem CID
|
|
| UNII | |
| |
| |
| Properties | |
| C28H47NO8 | |
| Molar mass | 525.683 g·mol−1 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| Infobox references | |
Pikromycin was studied by Brokmann and Hekel in 1951 and was the first antibiotic macrolide to be isolated.[1] Pikromycin is synthesized through a type I polyketide synthase system in Streptomyces venezuelae, a species of Gram-positive bacterium in the genus Streptomyces.[2] Pikromycin is derived from narbonolide, a 14-membered ring macrolide. [3] Along with the narbonolide backbone, pikromycin includes a desosamine sugar and a hydroxyl group. Although Pikromycin is not a clinically useful antibiotic, it can be used as a raw material to synthesize antibiotic ketolide compounds such as ertythromycins and new epothilones. [4]
Biosynthesis
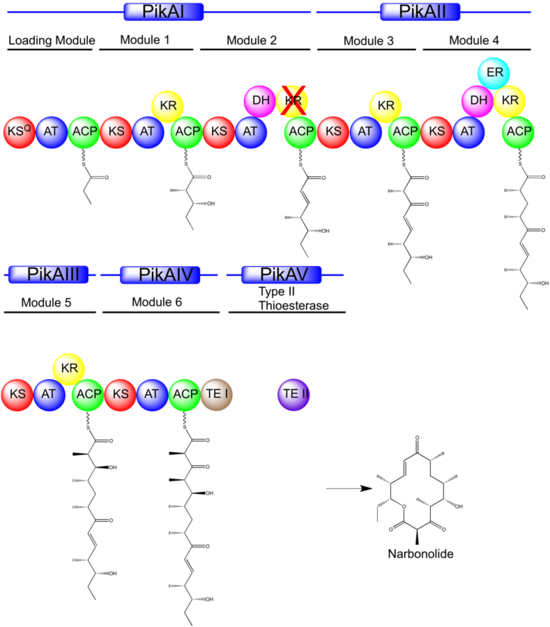
The pikromycin polyketide synthase of Streptomyces venezuelae contains four polypeptides: PikAI, PikAII, PikAIII, and PikAIV. These polypeptides contain a loading module, six extension molecules, and a thioesterase domain that terminated the biosynthetic procedure. [5] Recently electron cryo-microscopy have been used to determine sub-nanometre-resolution three- dimensional reconstructions of a full-length PKS module from the bacterium Streptomyces venezuelae that revealed an unexpectedly different architecture. [6] In Figure 1, each circle corresponds to a PKS mutilifuctional protein, where ACP is acyl carrier protein, KS is keto-ACP synthase, KSQ is a keto-ACP synthase like domain, AT is acyltransferase, KR is keto ACP reductase, KR with cross is inactive KR, DH is hydroxyl-thioester dehydratase, ER is enoyl reductase, TEI is thioesterase domain I, TEII is type II thioesterase. [7] Des corresponds to the enzymes utilized in desosamine biosynthesis and transfer, which include DesI-DesVIII.[citation needed]
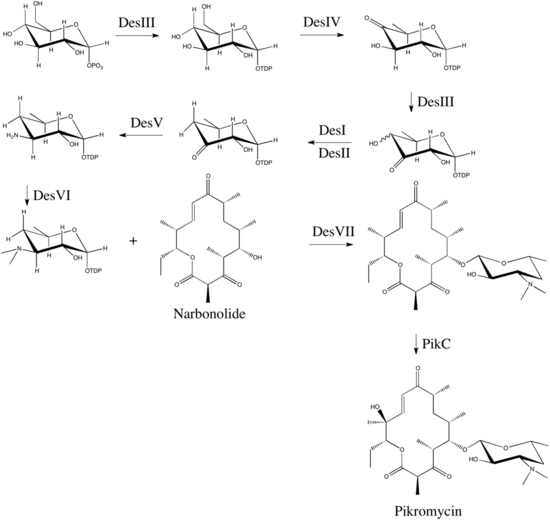
Figure 2 represents the desosamine deoxyamino sugar biosynthetic pathway. DesI-DesVI (des locus of pikromycin PKS) encodes all the enzymes needed to obtain TDP-desoamine from TDP-glucose. DesVII and DesVIII activities transfer desoamine to narbonolide and narbomycin is obtained. PikC cytochrome P450 hydrolase catalyzes the hydroxylation of narbomycin to obtain pikromycin. [2]
See also
References
- ↑ Brockmann, H.; Henkel, W. (1951). "Pikromycin, ein bitter schmeckendes Antibioticum aus Actinomyceten". Ntibiotica aus Actinomyceten 84: 184–288. doi:10.1002/cber.19510840306.
- ↑ 2.0 2.1 Y. Xue; D. Sherman (2001). "Biosynthesis and Combinatorial Biosynthesis of Pikromycin-Related Macrolides in Streptomyces venezuelae". Metabolic Engineering 3 (1): 15–26. doi:10.1006/mben.2000.0167. PMID 11162229.
- ↑ Maezawa, T. Hori, A. Kinumaki and M. Suzuki (1973). "Biological conversion of narbonolide to picromycin". The Journal of Antibiotics 26 (12): 771–775. doi:10.7164/antibiotics.26.771. PMID 4792390.
- ↑ J.D. Kittendorf; D.H. Sherman (2009). "The Methymycin/Pikromycin Biosynthetic Pathway: A Model for Metabolic Diversity in Natural Product". Bioorg Med Chem 17 (6): 2137–2146. doi:10.1016/j.bmc.2008.10.082. PMID 19027305.
- ↑ S. Guptaa; V. Lakshmanan; B.S. Kima; R. Fecik; K. A. Reynolds (2008). "Generation of Novel Pikromycin Antibiotic Products Through Mutasynthesis". ChemBioChem 9 (10): 1609–1616. doi:10.1002/cbic.200700635. PMID 18512859.
- ↑ S. Dutta; J. R. Whicher; D. A. Hansen; W. A. Hale; J. A. Chemler; G. R. Congdon; A. R. H. Narayan; K. Håkansson et al. (2014). "Structure of amodular polyketide synthase". Nature 510 (7506): 512–517. doi:10.1038/nature13423. PMID 24965652. Bibcode: 2014Natur.510..512D.
- ↑ D.L. Akey; J.D. Kittendorf; J.W. Giraldes; R.A. Fecik; D.H. Sherman; J.L. Smith (2006). "Structural basis for macrolactonization by the pikromycin thioesterase.". Nature Chemical Biology 2 (10): 537–542. doi:10.1038/nchembio824. PMID 16969372.
 |
