Two-state trajectory
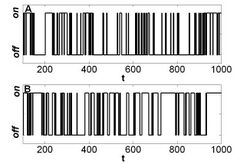
A two-state trajectory (also termed two-state time trajectory or a trajectory with two states) is a dynamical signal that fluctuates between two distinct values: ON and OFF, open and closed, , etc. Mathematically, the signal has, for every either the value or .
In most applications, the signal is stochastic; nevertheless, it can have deterministic ON-OFF components. A completely deterministic two-state trajectory is a square wave. There are many ways one can create a two-state signal, e.g. flipping a coin repeatedly.
A stochastic two-state trajectory is among the simplest stochastic processes. Extensions include: three-state trajectories, higher discrete state trajectories, and continuous trajectories in any dimension.[1]
Two state trajectories in biophysics, and related fields
Two state trajectories are very common. Here, we focus on relevant trajectories in scientific experiments: these are seen in measurements in chemistry, physics, and the biophysics of individual molecules[2][3] (e.g. measurements of protein dynamics and DNA and RNA dynamics,[4][5][6][7][8] activity of ion channels,[9][10] enzyme activity,[11][12][13][14][15] quantum dots[16][17][18][19][20][21]). From these experiments, one aims at finding the correct model explaining the measured process.[22][23][24][25][26][27][28][29][30][31][32] We explain about various relevant systems in what follows.
Ion channels
Since the ion channel is either opened or closed, when recording the number of ions that go through the channel when time elapses, observed is a two-state trajectory of the current versus time.
Enzymes
Here, there are several possible experiments on the activity of individual enzymes with a two-state signal. For example, one can create substrate that only upon the enzymatic activity shines light when activated (with a laser pulse). So, each time the enzyme acts, we see a burst of photons during the time period that the product molecule is in the laser area.
Dynamics of biological molecules
Structural changes of molecules are viewed in various experiments' type. Förster resonance energy transfer is an example. In many cases one sees a time trajectory that fluctuates among several cleared defined states.
Quantum dots
Another system that fluctuates among an on state and an off state is a quantum dot. Here, the fluctuations are since the molecule is either in a state that emits photons or in a dark state that does not emit photons (the dynamics among the states are influenced also from its interactions with the surroundings).
See also
References
- ↑ Erhan Cinlar (1975). Introduction to Stochastic Processes. Prentice Hall Inc, New Jersey. ISBN 978-0-486-49797-6.
- ↑ Moerner, W. E.; Orrit, M (1999). "Illuminating Single Molecules in Condensed Matter". Science 283 (5408): 1670–6. doi:10.1126/science.283.5408.1670. PMID 10073924. Bibcode: 1999Sci...283.1670M.
- ↑ Weiss, Shimon (1999). "Fluorescence Spectroscopy of Single Biomolecules". Science 283 (5408): 1676–83. doi:10.1126/science.283.5408.1676. PMID 10073925. Bibcode: 1999Sci...283.1676W.
- ↑ Schuler, Benjamin; Lipman, Everett A.; Eaton, William A. (2002). "Probing the free-energy surface for protein folding with single-molecule fluorescence spectroscopy". Nature 419 (6908): 743–7. doi:10.1038/nature01060. PMID 12384704. Bibcode: 2002Natur.419..743S. https://zenodo.org/record/1233253.
- ↑ Yang, Haw; Luo, Guobin; Karnchanaphanurach, Pallop; Louie, Tai-Man; Rech, Ivan; Cova, Sergio; Xun, Luying; Xie, X. Sunney (2003). "Protein Conformational Dynamics Probed by Single-Molecule Electron Transfer". Science 302 (5643): 262–6. doi:10.1126/science.1086911. PMID 14551431. Bibcode: 2003Sci...302..262Y.
- ↑ Min, Wei; Luo, Guobin; Cherayil, Binny J.; Kou, S. C.; Xie, X. Sunney (2005). "Observation of a Power-Law Memory Kernel for Fluctuations within a Single Protein Molecule". Physical Review Letters 94 (19). doi:10.1103/PhysRevLett.94.198302. PMID 16090221. Bibcode: 2005PhRvL..94s8302M.
- ↑ Rhoades, Elizabeth; Gussakovsky, Eugene; Haran, Gilad (2003). "Watching proteins fold one molecule at a time". Proceedings of the National Academy of Sciences 100 (6): 3197–202. doi:10.1073/pnas.2628068100. PMID 12612345. Bibcode: 2003PNAS..100.3197R.
- ↑ Zhuang, X.; Kim, H; Pereira, MJ; Babcock, HP; Walter, NG; Chu, S (2002). "Correlating Structural Dynamics and Function in Single Ribozyme Molecules". Science 296 (5572): 1473–6. doi:10.1126/science.1069013. PMID 12029135. Bibcode: 2002Sci...296.1473Z.
- ↑ Neher, Erwin; Sakmann, Bert (1976). "Single-channel currents recorded from membrane of denervated frog muscle fibres". Nature 260 (5554): 799–802. doi:10.1038/260799a0. PMID 1083489. Bibcode: 1976Natur.260..799N.
- ↑ Kasianowicz, John J.; Brandin, Eric; Branton, Daniel; Deamer, David W. (1996). "Characterization of individual polynucleotide molecules using a membrane channel". Proceedings of the National Academy of Sciences 93 (24): 13770–3. doi:10.1073/pnas.93.24.13770. PMID 8943010. Bibcode: 1996PNAS...9313770K.
- ↑ Lu, H. P.; Xun, L; Xie, XS (1998). "Single-Molecule Enzymatic Dynamics". Science 282 (5395): 1877–82. doi:10.1126/science.282.5395.1877. PMID 9836635. Bibcode: 1998Sci...282.1877P.
- ↑ Edman, Lars; Földes-Papp, Zeno; Wennmalm, Stefan; Rigler, Rudolf (1999). "The fluctuating enzyme: A single molecule approach". Chemical Physics 247 (1): 11–22. doi:10.1016/S0301-0104(99)00098-1. Bibcode: 1999CP....247...11E.
- ↑ Velonia, Kelly; Flomenbom, Ophir; Loos, Davey; Masuo, Sadahiro; Cotlet, Mircea; Engelborghs, Yves; Hofkens, Johan; Rowan, Alan E. et al. (2005). "Single-Enzyme Kinetics of CALB-Catalyzed Hydrolysis". Angewandte Chemie International Edition 44 (4): 560–4. doi:10.1002/anie.200460625. PMID 15619259.
- ↑ Flomenbom, O.; Velonia, K; Loos, D; Masuo, S; Cotlet, M; Engelborghs, Y; Hofkens, J; Rowan, AE et al. (2005). "Stretched exponential decay and correlations in the catalytic activity of fluctuating single lipase molecules". Proceedings of the National Academy of Sciences 102 (7): 2368–72. doi:10.1073/pnas.0409039102. PMID 15695587. Bibcode: 2005PNAS..102.2368F.
- ↑ English, Brian P; Min, Wei; Van Oijen, Antoine M; Lee, Kang Taek; Luo, Guobin; Sun, Hongye; Cherayil, Binny J; Kou, S C et al. (2005). "Ever-fluctuating single enzyme molecules: Michaelis-Menten equation revisited". Nature Chemical Biology 2 (2): 87–94. doi:10.1038/nchembio759. PMID 16415859.
- ↑ Nie, S; Chiu, D.; Zare, R. (1994). "Probing individual molecules with confocal fluorescence microscopy". Science 266 (5187): 1018–21. doi:10.1126/science.7973650. PMID 7973650. Bibcode: 1994Sci...266.1018N.
- ↑ Schmidt, Ulrich; Weiss, Matthias (2011). "Anomalous diffusion of oligomerized transmembrane proteins". The Journal of Chemical Physics 134 (16): 165101. doi:10.1063/1.3582336. PMID 21528980. Bibcode: 2011JChPh.134p5101S.
- ↑ Zumofen, Gert; Hohlbein, Johannes; Hübner, Christian (2004). "Recurrence and Photon Statistics in Fluorescence Fluctuation Spectroscopy". Physical Review Letters 93 (26). doi:10.1103/PhysRevLett.93.260601. PMID 15697961. Bibcode: 2004PhRvL..93z0601Z.
- ↑ Cohen, Adam E.; Moerner, WE (2006). "Suppressing Brownian motion of individual biomolecules in solution". Proceedings of the National Academy of Sciences 103 (12): 4362–5. doi:10.1073/pnas.0509976103. PMID 16537418. Bibcode: 2006PNAS..103.4362C.
- ↑ Moerner, W. E.; Dickson, Robert M.; Cubitt, Andrew B.; Tsien, Roger Y. (1997). "On/off blinking and switching behaviour of single molecules of green fluorescent protein". Nature 388 (6640): 355–8. doi:10.1038/41048. PMID 9237752. Bibcode: 1997Natur.388..355D.
- ↑ Chung, Inhee; Bawendi, Moungi (2004). "Relationship between single quantum-dot intermittency and fluorescence intensity decays from collections of dots". Physical Review B 70 (16). doi:10.1103/PhysRevB.70.165304. Bibcode: 2004PhRvB..70p5304C.
- ↑ Bauer, R.J.; Bowman, B.F.; Kenyon, J.L. (1987). "Theory of the kinetic analysis of patch-clamp data". Biophysical Journal 52 (6): 961–78. doi:10.1016/S0006-3495(87)83289-7. PMID 2447973. Bibcode: 1987BpJ....52..961B.
- ↑ Kienker, P. (1989). "Equivalence of Aggregated Markov Models of Ion-Channel Gating". Proceedings of the Royal Society B: Biological Sciences 236 (1284): 269–309. doi:10.1098/rspb.1989.0024. PMID 2471201. Bibcode: 1989RSPSB.236..269K.
- ↑ Fredkin, Donald R.; Rice, John A. (1986). "On Aggregated Markov Processes". Journal of Applied Probability 23 (1): 208–14. doi:10.2307/3214130.
- ↑ Colquhoun, D.; Hawkes, A. G. (1982). "On the Stochastic Properties of Bursts of Single Ion Channel Openings and of Clusters of Bursts". Philosophical Transactions of the Royal Society B: Biological Sciences 300 (1098): 1–59. doi:10.1098/rstb.1982.0156. PMID 6131450. Bibcode: 1982RSPTB.300....1C.
- ↑ Song, L.; Magleby, K.L. (1994). "Testing for microscopic reversibility in the gating of maxi K+ channels using two-dimensional dwell-time distributions". Biophysical Journal 67 (1): 91–104. doi:10.1016/S0006-3495(94)80458-8. PMID 7919030. Bibcode: 1994BpJ....67...91S.
- ↑ Qin, Feng; Auerbach, Anthony; Sachs, Frederick (2000). "Hidden Markov Modeling for Single Channel Kinetics with Filtering and Correlated Noise". Biophysical Journal 79 (4): 1928–44. doi:10.1016/S0006-3495(00)76442-3. PMID 11023898. Bibcode: 2000BpJ....79.1928Q.
- ↑ Bruno, W. J.; Yang, J; Pearson, JE (2005). "Using independent open-to-closed transitions to simplify aggregated Markov models of ion channel gating kinetics". Proceedings of the National Academy of Sciences 102 (18): 6326–31. doi:10.1073/pnas.0409110102. PMID 15843461. Bibcode: 2005PNAS..102.6326B.
- ↑ Flomenbom, O.; Silbey, RJ (2006). "Utilizing the information content in two-state trajectories". Proceedings of the National Academy of Sciences 103 (29): 10907–10. doi:10.1073/pnas.0604546103. PMID 16832051. Bibcode: 2006PNAS..10310907F.
- ↑ Flomenbom, Ophir; Klafter, Joseph; Szabo, Attila (2005). "What Can One Learn from Two-State Single-Molecule Trajectories?". Biophysical Journal 88 (6): 3780–3. doi:10.1529/biophysj.104.055905. PMID 15764653. Bibcode: 2005BpJ....88.3780F.
- ↑ Flomenbom, O.; Silbey, R. J. (2008). "Toolbox for analyzing finite two-state trajectories". Physical Review E 78 (6). doi:10.1103/PhysRevE.78.066105. PMID 19256903. Bibcode: 2008PhRvE..78f6105F.
- ↑ Flomenbom, Ophir (2011). "Making it Possible: Constructing a Reliable Mechanism from a Finite Trajectory". in Komatsuzaki, Tamiki; Kawakami, Masaru; Takahashi, Satoshi et al.. Single-Molecule Biophysics: Experiment and Theory, Volume 146. Advances in Chemical Physics. 146. pp. 367–93. doi:10.1002/9781118131374.ch13. ISBN 978-1-118-13137-4. https://books.google.com/books?id=vRCrDHyyNUgC&pg=PT372.
 |
