Chemistry:Dynamic combinatorial chemistry
This article only describes one highly specialized aspect of its associated subject. (October 2014) |
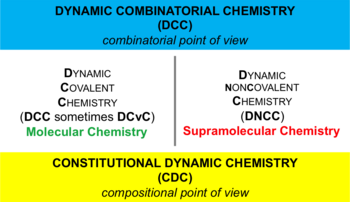
Dynamic combinatorial chemistry (DCC); also known as constitutional dynamic chemistry (CDC) is a method to the generation of new molecules formed by reversible reaction of simple building blocks under thermodynamic control.[3][4] The library[further explanation needed] of these reversibly interconverting building blocks is called a dynamic combinatorial library (DCL).[5][6] All constituents in a DCL are in equilibrium, and their distribution is determined by their thermodynamic stability within the DCL. The interconversion of these building blocks may involve covalent or non-covalent interactions. When a DCL is exposed to an external influence (such as proteins or nucleic acids), the equilibrium shifts and those components that interact with the external influence are stabilised and amplified, allowing more of the active compound to be formed.
History
By modern definition, dynamic combinatorial chemistry is generally considered to be a method of facilitating the generation of new chemical species by the reversible linkage of simple building blocks, under thermodynamic control.[4] This principle is known to select the most thermodynamically stable product from an equilibrating mixture of a number of components, a concept commonly utilised in synthetic chemistry to direct the control of reaction selectivity.[7] Although this approach was arguably utilised in the work of Fischer[8] and Werner[9] as early as the 19th century, their respective studies of carbohydrate and coordination chemistry were restricted to rudimentary speculation, requiring the rationale of modern thermodynamics.[10][11] It was not until supramolecular chemistry revealed early concepts of molecular recognition, complementarity and self-organisation that chemists could begin to employ strategies for the rational design and synthesis of macromolecular targets.[12] The concept of template synthesis was further developed and rationalised through the pioneering work of Busch in the 1960s, which clearly defined the role of a metal ion template in stabilising the desired ‘thermodynamic’ product, allowing for its isolation from the complex equilibrating mixture.[13][14] Although the work of Busch helped to establish the template method as a powerful synthetic route to stable macrocyclic structures, this approach remained exclusively within the domain of inorganic chemistry until the early 1990s, when Sanders et al. first proposed the concept of dynamic combinatorial chemistry.[4] Their work combined thermodynamic templation in tandem with combinatorial chemistry, to generate an ensemble complex porphyrin and imine macrocycles using a modest selection of simple building blocks.
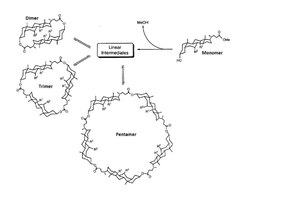
Sanders then developed this early manifestation of dynamic combinatorial chemistry as a strategy for organic synthesis; the first example being the thermodynamically-controlled macrolactonisation of oligocholates to assemble cyclic steroid-derived macrocycles capable of interconversion via component exchange.[15] Early work by Sanders et al. employed transesterification to generate dynamic combinatorial libraries. In retrospect, it was unfortunate that esters were selected for mediating component exchange, as transesterification processes are inherently slow and require vigorous anhydrous conditions.[4] However, their subsequent investigations identified that both the disulfide and hydrazone covalent bonds exhibit effective component exchange processes and so present a reliable means of generating dynamic combinatorial libraries capable of thermodynamic templation. This chemistry now forms the basis of much research in the developing field of dynamic covalent chemistry, and has in recent years emerged as a powerful tool for the discovery of molecular receptors.
Protein-directed
One of the key developments within the field of DCC is the use of proteins (or other biological macromolecules, such as nucleic acids) to influence the evolution and generation of components within a DCL.[16][17][18][19][20][21] Protein-directed DCC provides a way to generate, identify and rank novel protein ligands, and therefore have huge potential in the areas of enzyme inhibition and drug discovery.[22]

Reversible covalent reactions
The development of protein-directed DCC has not been straightforward because the reversible reactions employed must occur in aqueous solution at biological pH and temperature, and the components of the DCL must be compatible with proteins.[16][22]
Several reversible reactions have been proposed and/or applied in protein-directed DCC. These included boronate ester formation,[23][24][25] diselenides-disulfides exchange,[26] disulphide formation,[27][28][29] hemithiolacetal formation,[30][31] hydrazone formation,[32][33] imine formation[34][35][36] and thiol-enone exchange.[37]
Pre-equilibrated DCL
For reversible reactions that do not occur in aqueous buffers, the pre-equilibrated DCC approach can be used. The DCL was initially generated (or pre-equilibrated) in organic solvent, and then diluted into aqueous buffer containing the protein target for selection. Organic based reversible reactions, including Diels-Alder[38] and alkene cross metathesis reactions,[39] have been proposed or applied to protein-directed DCC using this method.
Reversible non-covalent reactions
Reversible non-covalent reactions, such as metal-ligand coordination,[40][41] has also been applied in protein-directed DCC. This strategy is useful for the investigation of the optimal ligand stereochemistry to the binding site of the target protein.[42]
Enzyme-catalysed reversible reactions
Enzyme-catalysed reversible reactions, such as protease-catalysed amide bond formation/hydrolysis reactions[43] and the aldolase-catalysed aldol reactions,[44][45] have also been applied to protein-directed DCC.
Analytical methods
Protein-directed DCC system must be amenable to efficient screening.[16][22] Several analytical techniques have been applied to the analysis of protein-directed DCL. These include HPLC,[27][31][32][35] mass spectrometry,[24][28][29][33] NMR spectroscopy,[23][25][30] and X-ray crystallography.[46]
Multi-protein approach
Although most applications of protein-directed DCC to date involved the use of single protein in the DCL, it is possible to identify protein ligands by using multiple proteins simultaneously, as long as a suitable analytical technique is available to detect the protein species that interact with the DCL components.[47] This approach may be used to identify specific inhibitors or broad-spectrum enzyme inhibitors.
Other applications
DCC is useful in identifying molecules with unusual binding properties, and provides synthetic routes to complex molecules that aren't easily accessible by other means. These include smart materials, foldamers, self-assembling molecules with interlocking architectures and new soft materials.[4] The application of DCC to detect volatile bioactive compounds, i.e. the amplification and sensing of scent, was proposed in a concept paper.[48] Recently, DCC was also used to study the abiotic origins of life.[49]
See also
- Combinatorial biology
- Combinatorial chemistry
- Drug discovery
- Fragment-based lead discovery
- High-throughput screening
References
- ↑ Lehn, Jean-Marie (2007). "From supramolecular chemistry towards constitutional dynamic chemistry and adaptive chemistry" (in en). Chem. Soc. Rev. 36 (2): 151–160. doi:10.1039/B616752G. ISSN 0306-0012. PMID 17264919. http://xlink.rsc.org/?DOI=B616752G.
- ↑ Lehn, Jean-Marie (2011). "Constitutional dynamic chemistry: Bridge from supramolecular chemistry to adaptive chemistry". in Barboiu, Mihail. Constitutional Dynamic Chemistry. Topics in Current Chemistry. 322. Springer Berlin Heidelberg. pp. 1–32. doi:10.1007/128_2011_256. ISBN 978-3-642-28343-7.
- ↑ Schaufelberger, F.; Timmer, B. J. J.; Ramström, O. Principles of Dynamic Covalent Chemistry. In Dynamic Covalent Chemistry: Principles, Reactions, and Applications; Zhang, W.; Jin, Y., Eds.; John Wiley & Sons: Chichester, 2018; Chapter 1, pp 1–30.
- ↑ 4.0 4.1 4.2 4.3 4.4 Corbett, P. T.; Leclaire, J.; Vial, L.; West, K. R.; Wietor, J.-L.; Sanders, J. K. M.; Otto, S. (Sep 2006). "Dynamic combinatorial chemistry". Chem. Rev. 106 (9): 3652–3711. doi:10.1021/cr020452p. PMID 16967917.
- ↑ Komáromy, D.; Nowak, P.; Otto, S. Dynamic Combinatorial Libraries. In Dynamic Covalent Chemistry: Principles, Reactions, and Applications; Zhang, W.; Jin, Y., Eds.; John Wiley & Sons: Chichester, 2018; Chapter 2, pp 31–119.
- ↑ Lehn, J.-M.; Ramström, O. Generation and screening of a dynamic combinatorial library. PCT. Int. Appl. WO 20010164605, 2001.
- ↑ Rowan, Stuart J.; Cantrill, Stuart J.; Cousins, Graham R. L.; Sanders, Jeremy K. M.; Stoddart, J. Fraser (2002-03-15). "Dynamic Covalent Chemistry". Angewandte Chemie International Edition 41 (6): 898–952. doi:10.1002/1521-3773(20020315)41:6<898::AID-ANIE898>3.0.CO;2-E. ISSN 1521-3773. PMID 12491278.
- ↑ Kunz, Horst (2002-12-02). "Emil Fischer—Unequalled Classicist, Master of Organic Chemistry Research, and Inspired Trailblazer of Biological Chemistry". Angewandte Chemie International Edition 41 (23): 4439–4451. doi:10.1002/1521-3773(20021202)41:23<4439::AID-ANIE4439>3.0.CO;2-6. ISSN 1521-3773. PMID 12458504.
- ↑ Constable, Edwin C.; Housecroft, Catherine E. (2013-01-28). "Coordination chemistry: the scientific legacy of Alfred Werner". Chem. Soc. Rev. 42 (4): 1429–1439. doi:10.1039/c2cs35428d. PMID 23223794.
- ↑ Anderson, Sally; Anderson, Harry L.; Sanders, Jeremy K. M. (1993-09-01). "Expanding roles for templates in synthesis". Accounts of Chemical Research 26 (9): 469–475. doi:10.1021/ar00033a003. ISSN 0001-4842.
- ↑ Hoss, Ralf; Vögtle, Fritz (1994-03-03). "Template Syntheses". Angewandte Chemie International Edition in English 33 (4): 375–384. doi:10.1002/anie.199403751. ISSN 1521-3773.
- ↑ Lehn, Jean-Marie (2007-01-30). "From supramolecular chemistry towards constitutional dynamic chemistry and adaptive chemistry". Chem. Soc. Rev. 36 (2): 151–160. doi:10.1039/b616752g. PMID 17264919.
- ↑ Thompson, Major C.; Busch, Daryle H. (1964-01-01). "Reactions of Coordinated Ligands. VI. Metal Ion Control in the Synthesis of Planar Nickel(II) Complexes of α-Diketo-bis-mercaptoimines". Journal of the American Chemical Society 86 (2): 213–217. doi:10.1021/ja01056a021. ISSN 0002-7863.
- ↑ Thompson, Major C.; Busch, Daryle H. (1962-05-01). "Reactions of Coördinated Ligands. II. Nickel(II) Complexes of Some Novel Tetradentate Ligands". Journal of the American Chemical Society 84 (9): 1762–1763. doi:10.1021/ja00868a073. ISSN 0002-7863.
- ↑ Brady, Paul A.; Bonar-Law, Richard P.; Rowan, Stuart J.; Suckling, Christopher J.; Sanders, Jeremy K. M. (January 1996). "?Living? macrolactonisation: thermodynamically-controlled cyclisation and interconversion of oligocholates". Chemical Communications (3): 319–320. doi:10.1039/cc9960000319.
- ↑ 16.0 16.1 16.2 Greaney, M. F.; Bhat, V. T. Protein-directed dynamic combinatorial chemistry. In Dynamic combinatorial chemistry: in drug discovery, bioinorganic chemistry, and materials sciences; Miller, B. L., Ed.; John Wiley & Sons: New Jersey, 2010; Chapter 2, pp 43–82.
- ↑ Huang, R.; Leung, I. K. H. (Jul 2016). "Protein-directed dynamic combinatorial chemistry: a guide to protein ligand and inhibitor discovery". Molecules 21 (7): 910. doi:10.3390/molecules21070910. PMID 27438816.
- ↑ Frei, P.; Hevey, R.; Ernst, B. (Sep 2018). "Dynamic Combinatorial Chemistry: A New Methodology Comes of Age". Chem. Eur. J. 25 (1): 60–73. doi:10.1002/chem.201803365. PMID 30204930.
- ↑ Jaegle, M.; Wong, E. L.; Tauber, C.; Nawrotzky, E.; Arkona, C.; Rademann, J. (Jan 2017). "Protein-templated fragment ligations - from molecular recognition to drug discovery". Angew. Chem. Int. Ed. 56 (26): 7358–7378. doi:10.1002/anie.201610372. PMID 28117936.
- ↑ Mondal, M.; Hirsch, A. K. (Apr 2015). "Dynamic combinatorial chemistry: a tool to facilitate the identification of inhibitors for protein targets". Chem. Soc. Rev. 44 (8): 2455–2488. doi:10.1039/c4cs00493k. PMID 25706945.
- ↑ Herrmann, A. (Mar 2014). "Dynamic combinatorial/covalent chemistry: a tool to read, generate and modulate the bioactivity of compounds and compound mixtures". Chem. Soc. Rev. 43 (6): 1899–1933. doi:10.1039/c3cs60336a. PMID 24296754.
- ↑ 22.0 22.1 22.2 Hochgürtel, M.; Lehn, J.-M. Dynamic combinatorial diversity in drug discovery. In Fragment-based approaches in drug discovery; Jahnke, W., Erlanson, D. A., Ed.; Wiley-VCH: Weinheim, 2006; Chapter 16, pp 341–364.
- ↑ 23.0 23.1 23.2 Leung, I. K. H.; Demetriades, M.; Hardy, A. P.; Lejeune, C.; Smart, T. J.; Szöllössi, A.; Kawamura, A.; Schofield, C. J. et al. (Jan 2013). "NMR reporter ligand screening for inhibitors of 2OG oxygenases". J. Med. Chem. 56 (2): 547–555. doi:10.1021/jm301583m. PMID 23234607.
- ↑ 24.0 24.1 Demetriades, M.; Leung, I. K. H.; Chowdhury, R.; Chan, M. C.; Yeoh, K. K.; Tian, Y.-M.; Claridge, T. D. W.; Ratcliffe, P. J. et al. (Jul 2012). "Dynamic combinatorial chemistry employing boronic acids/boronate esters leads to potent oxygenase inhibitors". Angew. Chem. Int. Ed. 51 (27): 6672–6675. doi:10.1002/anie.201202000. PMID 22639232.
- ↑ 25.0 25.1 Leung, I. K. H.; Brown Jr, T.; Schofield, C. J.; Claridge, T. D. W. (May 2011). "An approach to enzyme inhibition employing reversible boronate ester formation". Med. Chem. Commun. 2 (5): 390–395. doi:10.1039/C1MD00011J.
- ↑ Rasmussen, B.; Sørensen, A.; Gotfredsen, H.; Pittelkow, M. (Feb 2014). "Dynamic combinatorial chemistry with diselenides and disulfides in water". Chem. Commun. 50 (28): 3716–3718. doi:10.1039/C4CC00523F. PMID 24577496.
- ↑ 27.0 27.1 Ramström, O.; Lehn, J.-M. (Jul 2000). "In situ generation and screening of a dynamic combinatorial carbohydrate library against concanavalin A". ChemBioChem 1 (1): 41–48. doi:10.1002/1439-7633(20000703)1:1<41::AID-CBIC41>3.0.CO;2-L. PMID 11828397.
- ↑ 28.0 28.1 Liénard, B. M. R.; Selevsek, N.; Oldham, N. J.; Schofield, C. J. (Feb 2007). "Combined mass spectrometry and dynamic chemistry approach to identify metalloenzyme inhibitors". ChemMedChem 2 (2): 175–179. doi:10.1002/cmdc.200600250. PMID 17206734.
- ↑ 29.0 29.1 Liénard, B. M. R.; Hüting, R.; Lassaux, P.; Galleni, M.; Frére, J.-M.; Schofield, C. J. (Feb 2008). "Dynamic combinatorial mass spectrometry leads to metallo-β-lactamase inhibitors". J. Med. Chem. 51 (3): 684–688. doi:10.1021/jm070866g. PMID 18205296.
- ↑ 30.0 30.1 Caraballo, R.; Dong, H.; Ribeiro, J. P.; Jiménez-Barbero, J.; Ramström, O. (Jan 2010). "Direct STD NMR identification of β-galactosidase inhibitors from a virtual dynamic hemithioacetal system". Angew. Chem. Int. Ed. 49 (3): 589–593. doi:10.1002/anie.200903920. PMID 20013972.
- ↑ 31.0 31.1 Clipson, A. J.; Bhat, V. T.; McNae, I.; Caniard, A. M.; Campopiano, D. J.; Greaney, M. F. (Aug 2012). "Bivalent enzyme inhibitors discovered using dynamic covalent chemistry". Chem. Eur. J. 18 (34): 10562–10570. doi:10.1002/chem.201201507. PMID 22782854. https://www.pure.ed.ac.uk/ws/files/11097675/Bivalent_Enzyme_Inhibitors_Discovered_Using_Dynamic_Covalent_Chemistry.pdf.
- ↑ 32.0 32.1 Hochgürtel, M.; Niesinger, R.; Kroth, H.; Piecha, D.; Hofmann, M. W.; Krause, S.; Schaaf, O.; Nicolau, C. et al. (Jan 2003). "Ketones as building blocks for dynamic combinatorial libraries: highly active neuraminidase inhibitors generated via selective pressure of the biological target". J. Med. Chem. 46 (3): 356–358. doi:10.1021/jm025589m. PMID 12540234.
- ↑ 33.0 33.1 Sindelar, M.; Lutz, T. A.; Petrera, M.; Wanner, K. T. (Feb 2013). "Focused pseudostatic hydrazone libraries screened by mass spectrometry binding assay: optimizing affinities toward γ-aminobutyric acid transporter 1". J. Med. Chem. 56 (3): 1323–1340. doi:10.1021/jm301800j. PMID 23336362.
- ↑ Yang, Z.; Fang, Z.; He, W.; Wang, Z.; Gang, H.; Tian, Q.; Guo, K. (Apr 2016). "Identification of inhibitors for vascular endothelial growth factor receptor by using dynamic combinatorial chemistry". Bioorg. Med. Chem. Lett. 26 (7): 1671–1674. doi:10.1016/j.bmcl.2016.02.063. PMID 26920800.
- ↑ 35.0 35.1 Zameo, S.; Vauzeilles, B.; Beau, J.-M. (Dec 2006). "Direct composition analysis of a dynamic library of imines in an aqueous medium". Eur. J. Org. Chem. 2006 (24): 5441–5444. doi:10.1002/ejoc.200600859.
- ↑ Herrmann, A. (Aug 2009). "Dynamic mixtures and combinatorial libraries: imines as probes for molecular evolution at the interface between chemistry and biology". Org. Biomol. Chem. 7 (16): 3195–3204. doi:10.1039/B908098H. PMID 19641772.
- ↑ Shi, B.; Stevenson, R.; Campopiano, D. J.; Greaney, M. F. (Jul 2006). "Discovery of glutathione S-transferase inhibitors using dynamic combinatorial chemistry". J. Am. Chem. Soc. 128 (26): 8459–8467. doi:10.1021/ja058049y. PMID 16802811.
- ↑ Boul, P. J.; Reutenauer, P.; Lehn, J.-M. (Jan 2005). "Reversible Diels-Alder reactions for the generation of dynamic combinatorial libraries". Org. Lett. 7 (1): 15–18. doi:10.1021/ol048065k. PMID 15624966.
- ↑ Poulsen, S.-A.; Bornaghi, L. F. (May 2006). "Fragment-based drug discovery of carbonic anhydrase II inhibitors by dynamic combinatorial chemistry utilizing alkene cross metathesis". Bioorg. Med. Chem. 14 (10): 3275–3284. doi:10.1016/j.bmc.2005.12.054. PMID 16431113.
- ↑ Sakai, S.; Shigemasa, Y.; Sasaki, T. (Nov 1997). "A self-adjusting carbohydrate ligand for GalNAc specific lectins". Tetrahedron Lett. 38 (47): 8145–8148. doi:10.1016/S0040-4039(97)10187-3.
- ↑ Sakai, S.; Shigemasa, Y.; Sasaki, T. (1999). "Iron(II)-assisted assembly of trivalent GalNAc clusters and their interactions with GalNAc-specific lectins". Bull. Chem. Soc. Jpn. 72 (6): 1313–1319. doi:10.1246/bcsj.72.1313. http://ci.nii.ac.jp/naid/130004150385.
- ↑ Kilpin, K. J.; Dyson, P. J. (Feb 2013). "Enzyme inhibition by metal complexes: concepts, strategies and applications". Chem. Sci. 4 (4): 1410–1419. doi:10.1039/C3SC22349C.
- ↑ Swann, P. G.; Casanova, R. A.; Desai, A.; Frauenhoff, M. M.; Urbancic, M.; Slomczynska, U.; Hopfinger, A. J.; Le Breton, G. C. et al. (1996). "Nonspecific protease-catalyzed hydrolysis/synthesis of a mixture of peptides: product diversity and ligand amplification by a molecular trap". Biopolymers 40 (6): 617–625. doi:10.1002/(sici)1097-0282(1996)40:6<617::aid-bip3>3.0.co;2-z. PMID 9140201.
- ↑ Lins, R. J.; Flitsch, S. L.; Turner, N. J.; Irving, E.; Brown, S. A. (Sep 2002). "Enzymatic generation and in situ screening of a dynamic combinatorial library of sialic acid analogues". Angew. Chem. Int. Ed. 41 (18): 3405–3407. doi:10.1002/1521-3773(20020916)41:18<3405::AID-ANIE3405>3.0.CO;2-P. PMID 12298046.
- ↑ Lins, R. J.; Flitsch, S. L.; Turner, N. J.; Irving, E.; Brown, S. A. (Jan 2004). "Generation of a dynamic combinatorial library using sialic acid aldolase and in situ screening against wheat germ agglutinin". Tetrahedron 60 (3): 771–780. doi:10.1016/j.tet.2003.11.062.
- ↑ Valade, A.; Urban, D.; Beau, J.-M. (Jan–Feb 2007). "Two galatosyltransferases' selection of different binders from the same uridine-based dynamic combinatorial library". J. Comb. Chem. 9 (1): 1–4. doi:10.1021/cc060033w. PMID 17206823.
- ↑ Das, M.; Tianming, Y.; Jinghua, D.; Prasetya, F.; Yiming, X.; Wong, K.; Cheong, A.; Woon, E. C. Y. (Jun 2018). "Multi-Protein Dynamic Combinatorial Chemistry: A Novel Strategy that Leads to Simultaneous Discovery of Subfamily-Selective Inhibitors for Nucleic Acid Demethylases FTO and ALKBH3". Chem. Asian J. 13 (19): 2854–2867. doi:10.1002/asia.201800729. PMID 29917331. https://discovery.ucl.ac.uk/id/eprint/10107020/.
- ↑ Herrmann, A. (Jul 2012). "Dynamic Mixtures: Challenges and Opportunities for the Amplification and Sensing of Scents". Chem. Eur. J. 18 (28): 8568–8577. doi:10.1002/chem.201200668. PMID 22588709.
- ↑ Chandru, Kuhan; Guttenberg, Nicholas; Giri, Chaitanya; Hongo, Yayoi; Butch, Christopher; Mamajanov, Irena; Cleaves, H. James (31 May 2018). "Simple prebiotic synthesis of high diversity dynamic combinatorial polyester libraries" (in En). Communications Chemistry 1 (1). doi:10.1038/s42004-018-0031-1. ISSN 2399-3669.
External links
- University of North Carolina at Chapel Hill: Center for Dynamic Combinatorial Chemistry [citation needed]
- University of Cambridge: Dynamic Combinatorial Chemistry
 |