Chemistry:Pseudouridine
| |
| Names | |
|---|---|
| IUPAC name
5-(β-D-Ribofuranosyl)pyrimidine-2,4(1H,3H)-dione
| |
| Systematic IUPAC name
5-[(2S,3R,4S,5R)-3,4-Dihydroxy-5-(hydroxymethyl)oxolan-2-yl]pyrimidine-2,4(1H,3H)-dione | |
| Other names
psi-Uridine, 5-Ribosyluracil, beta-D-Pseudouridine, 5-(beta-D-Ribofuranosyl)uracil
| |
| Identifiers | |
3D model (JSmol)
|
|
| 32779 | |
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| KEGG | |
PubChem CID
|
|
| UNII | |
| |
| |
| Properties | |
| C9H12N2O6 | |
| Molar mass | 244.20 g/mol |
| Appearance | White granular powder |
| Highly soluble in water. | |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| Infobox references | |
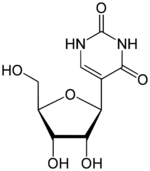
Pseudouridine (5-ribosyluracil, abbreviated by the Greek letter psi- Ψ)[1] is an isomer of the nucleoside uridine in which the uracil is attached via a carbon-carbon instead of a nitrogen-carbon glycosidic bond.
Pseudouridine is the most abundant RNA modification in cellular RNA.[2] After transcription and following synthesis, RNA can be modified with over 100 chemically distinct modifications. These can potentially regulate RNA expression post-transcriptionally, in addition to the four standard nucleotides and play a variety of roles in the cell including translation, localization and stabilization of RNA. Pseudouridine, being one of them, is the C5-glycoside isomer of uridine that contains a C-C bond between C1 of the ribose sugar and C5 of uracil, rather than usual C1-N1 bond found in uridine. Uridine is converted to pseudouridine by rotating the uridine molecule 180𝄪 across its N3-C6 axis.[3] The C-C bond gives it more rotational freedom and conformational flexibility.[4] In addition, pseudouridine has an extra hydrogen bond donor at the N1 position.
Pseudouridine is a ubiquitous constituent of structural RNA (transfer, ribosomal, small nuclear (snRNA) and small nucleolar), and present in coding RNA, across the three phylogenetic domains of life and was the first discovered. It accounts for 4% of the nucleotides in the yeast tRNA [citation needed]. This base modification is able to stabilize RNA and improve base-stacking by forming additional hydrogen bonds with water through its extra imino group.[5] There are 11 pseudouridines in the Escherichia coli rRNA, 30 in yeast cytoplasmic rRNA and a single modification in mitochondrial 21S rRNA and about 100 pseudouridines in human rRNA indicating that the extent of pseudouridylation increases with the complexity of an organism [citation needed]. Pseudouridine was also detected in Leishmania donovani genome. 18 pseudouridine modification sites were detected in the peptidyl transferase entry site and in the mRNA entry tunnel in protein translation. These modifications in the parasite lead to increased protein synthesis and growth rate.[6]
Pseudouridine in rRNA and tRNA has been shown to fine-tune and stabilize the regional structure and help maintain their functions in mRNA decoding, ribosome assembly, processing and translation.[4][7][8] Pseudouridine in snRNA has been shown to enhance spliceosomal RNA-pre-mRNA interaction to facilitate splicing regulation.[9]
Effects and modification on different RNA
tRNA
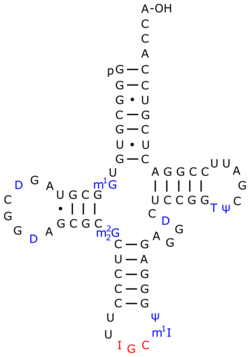
Ψ is ubiquitous in this class of RNAs and facilitates common tRNA structural motifs. One such structural motif is the TΨC stem loop which incorporates Ψ55. Ψ is commonly found in the D stem and anticodon stem and loop of tRNAs from each domain. In each structural motif the unique physicochemical properties of Ψ stabilize structures that would not be possible with the standard U.[4]
During translation Ψ modulates interactions of tRNA molecules with rRNAs and mRNAs. Ψ and other modified nucleotides affect the local structure of the tRNA domains they are found in without impacting the overall fold of the RNA. In the anticodon stem-loop (ASL) Ψ seems critical for proper binding of tRNAs to the ribosome. Ψ stabilizes the dynamic structure of the ASL and promotes stronger binding to the 30S ribosome. The stabilized conformation of the ASL helps maintain correct anticodon-codon pairings during translation. This stability may increase translational accuracy by decreasing the rate of peptide bond formation and allowing for more time for incorrect codon-anticodon pairs to be rejected. Despite Ψ’s role in local structure stabilization, pseudouridylation of tRNA is not essential for cell viability and is not usually required for aminoacylation.[4]
mRNA
Ψ is also found in mRNAs which are the template for protein synthesis. Ψ residues in mRNA can affect the coding specificity of stop codons UAA, UGA, and UAG. In these stop codons both a U→Ψ modification and a U→C mutation both promote nonsense suppression.[10] In the SARS-CoV2 vaccine from BioNTech/Pfizer, also known as BNT162b2, tozinameran or Comirnaty, all U's have been substituted with N1-methylpseudouridine,[11] a nucleoside related to Ψ that contains a methyl group added to N1 atom.
rRNA
Ψ is found in the large and small ribosomal subunits of all domains of life and their organelles. In the ribosome Ψ residues cluster in domains II, IV, and V and stabilize RNA-RNA and/or RNA-protein interactions. The stability afforded by Ψ may assist rRNA folding and ribosome assembly. Ψ may also influence the stability of local structures which impact the speed and accuracy of decoding and proofreading during translation.[4]
snRNA
Ψ is found in the major spliceosomal snRNAs of eukaryotes. Ψ residues in snRNA are often phylogenetically conserved, but have some variations across taxa and organisms. The Ψ residues in snRNAs are normally located in regions that participate in RNA-RNA and/or RNA-protein interactions involved in the assembly and function of the spliceosome. Ψ residues in snRNAS contribute to the proper folding and assembly of the spliceosome which is essential for pre-mRNA processing.[4]
Synthases
Pseudouridine are RNA modifications that are done post-transcription, so after the RNA is formed [citation needed]. The proteins that do this modification are called pseudouridine synthases (PUS) and are found in all kingdoms of life. Most research has been done on how PUS modify tRNA, so mechanisms involving snRNA, and mRNA are not clearly defined. PUS can vary on RNA specificity, structure, and isomerization mechanisms. The different structures of PUS are divided into five families which share the active sequence and important structural motifs.[1]
TruA
TruA domain modifies a variety of different places in tRNA, snRNA, and mRNA. The mechanism of isomerization of uridine is still being talked about in this family.[8][12]
PUS 1 is located in the nucleus and modifies tRNA at different locations, U44 of U2 snRNA, and U28 of U6 snRNA. Studies found that PUS 1 expression increased during environmental stress and is important for regulating the splicing of RNA. Also, that PUS 1 is necessary for taking the tRNA made in the nucleus and sending them to the cytoplasm.[8]
PUS 2 is very similar to PUS 1, but located in the mitochondria and only modifies U27 and U28 of mito-tRNA. This protein modifies the mitochondrial tRNA, which has a lesser amount of pseudouridine modifications compared to other tRNAs. Unlike most mitochondria located protein, PUS 2 has not been found to have a mitochondrial targeting signal or MTS.[8]
PUS 3 is a homolog to PUS 1, but modifies different places of the tRNA (U38/39) in the cytoplasm and mitochondrial. This protein is the most conserved of the TruA family. A decrease in modifications made by PUS 3 was found when the tRNA structure of improperly folded. Along with tRNA the protein targets ncRNA and mRNA, further research is still needed as to the importance of this modification. PUS 3 along with PUS 1 modify the steroid activator receptor in humans.[8]
TruB
The TruB family only contains PUS 4 located in the mitochondrial and nucleus. PUS 4 modification is heavily conserved located in the U55 in the elbow of the tRNA. The human form of PUS 4 is actually missing a binding domain called PUA or pseudouridine synthase and archaeosine trans-glycosylase. PUS 4 has a sequence specificity for T-loop part of the tRNA. Preliminary data of PUS4 modifying mRNA, but more research is needed to confirm. Also binds to a specific Brome Mosaic Virus, which is a plant-infecting RNA virus.[8][13]
TruD
TruD is able to modify a variety of RNA, and it is unclear how these different RNA substrates are recognized. PUS 7 modifies U2 snRNA at the position 35 and this modification will increase when the cells are in heat shock. Another modification is cytoplasmic tRNA in position 13, and position 35 in pre-tRNATyr. PUS 7 modifies almost specificity does not depend on the type of RNA as mRNA show pseudouridylated by PUS 7. Recognize this the sequence of the RNA, UGUAR with the second U being the nucleotide that will be modified. The pseudouridylation of mRNA by PUS 7 increases during heat shock, because the protein moves from the nucleus to the cytoplasm. The modification is thought to increase the stability of mRNA during heat shock before the RNA goes to the nucleus or mitochondria, but more studies are needed.[8][12]
RluA
The RluA domain of these proteins can identify the substrate through a different protein binding to the substrate and then particular bonds to the RluA domain.[1][12]
PUS 5 is not well studied and located pseudouridine synthase and similar to Pus 2 does not have a mitochondrial signal targeting sequence. The protein modifies U2819 of mitochondrial 21S rRNA. Also suspected that Pus 5 modifies some uridines in the mRNA, but again more data is needed to confirm.
PUS 6 has one that only modifies U31 of cytoplasmic and mitochondrial tRNA. Pus 6 is also known to modify mRNA.[8]
PUS 8 also known as Rib2 modifies cytoplasmic tRNA at position U32. On the C-terminus there is a DRAP-deaminase domain related to the biosynthesis of riboflavin. The RluA and DRAP or deaminase domain related to riboflavin synthase have completely separate functions in the protein and it is not known whether they interact with each other. PUS 8 is necessary in yeast, but that is suspected to be related to the riboflavin synthesis and not the pseudouridine modification.[8]
PUS 9 and PUS 8 catalyze the same position in mitochondrial tRNA instead of cytoplasmic. It is the only PUS protein that contains a mitochondrial targeting signal domain on the N-terminus. Studies suggest that PUS 9 can modify mRNAs, which would mean less substrate specificity.[8]
RsuA
Techniques in genome sequencing for pseudouridine
Pseudouridine can be identified through a multitude of different techniques. A common technique to identify modifications in RNA and DNA is Liquid Chromatography with Mass Spectrometry or LC-MS. Mass spectrometry separates molecules by the mass and charge. While uridine and pseudouridine have the same mass, they have different charges. Liquid chromatography works by retention time, which has to do with leaving the column.[14] A chemical way to identify pseudouridine uses a compound called CMC or N-cyclohexyl-N′-β-(4-methylmorpholinium) ethylcarbodiimide to specifically label and distinguish uridine from pseudouridine. CMC will bond both with pseudouridine and uridine, but holds tighter to the former, because of the third nitrogen able to form hydrogen bond. CMC bound to pseudouridine can then be imaged by tagging a signaling molecule. This method is still being worked on to become high-throughput.[15]
Medical relevance of pseudouridine
Pseudouridine exerts a subtle but significant influence on the nearby sugar-phosphate backbone and also enhances base stacking. These effects may underlie the biological role of most, but perhaps not all of the pseudouridine residues in RNA. Certain genetic mutants lacking specific pseudouridine residues in tRNA or rRNA exhibit difficulties in translation, display slow growth rates, and fail to compete effectively with wild-type strains in mixed culture. Pseudouridine modifications are also implicated in human diseases such as mitochondrial myopathy and sideroblastic anemia (MLASA) and Dyskeratosis congenita.[8] Dyskeratosis congenita and Hoyeraal-Hreidarsson syndrome are two rare inherited syndromes caused by mutations in DKC1, the gene encoding for the pseudouridine synthase dyskerin. Pseudouridines have been recognized as regulators of viral latency processes in human immunodeficiency virus (HIV) infections.[16] Pseudouridylation has also been associated with the pathogenesis of maternally inherited diabetes and deafness (MIDD). In particular, a point mutation in a mitochondrial tRNA seems to prevent the pseudouridylation of one nucleotide, thus altering the tRNA tertiary structure. This may lead to higher tRNA instability, causing deficiencies in mitochondrial translation and respiration.[16]
Vaccines
When pseudouridine is used in place of uridine in synthetic mRNA, the modified mRNA molecule arouses less response from Toll-like receptors, a part of the human immune system that would otherwise identify the mRNA as unwelcome. This makes pseudouridine useful in mRNA vaccines, including the mRNA COVID-19 vaccines. This property of pseudouridine was discovered by Katalin Karikó and Drew Weissman in 2005, for which they shared the 2023 Nobel Prize in Physiology or Medicine.[17][18]
N1-Methylpseudouridine provides even less innate immune response than Ψ, as well as improving translation capacity.[19] Both Pfizer-BioNTech and Moderna mRNA vaccines therefore use N1-Methylpseudouridine rather than Ψ.[19]
See also
- Pseudouridine kinase
- TRNA-pseudouridine synthase
- PUS1
References
- ↑ 1.0 1.1 1.2 Hamma, Tomoko; Ferré-D'Amaré, Adrian R. (November 2006). "Pseudouridine Synthases". Chemistry & Biology 13 (11): 1125–1135. doi:10.1016/j.chembiol.2006.09.009. ISSN 1074-5521. PMID 17113994.
- ↑ Penzo, Marianna; Guerrieri, Ania; Zacchini, Federico; Treré, Davide; Montanaro, Lorenzo (2017-11-01). "RNA Pseudouridylation in Physiology and Medicine: For Better and for Worse" (in en). Genes 8 (11): 301. doi:10.3390/genes8110301. ISSN 2073-4425. PMID 29104216.
- ↑ "Dyskerin: an essential pseudouridine synthase with multifaceted roles in ribosome biogenesis, splicing, and telomere maintenance". RNA 27 (12): 1441–1458. 2021. doi:10.1261/rna.078953.121. PMID 34556550.
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 Gray, Michael Charette, Michael W. (2000-05-01). "Pseudouridine in RNA: What, Where, How, and Why". IUBMB Life 49 (5): 341–351. doi:10.1080/152165400410182. ISSN 1521-6543. PMID 10902565.
- ↑ Davis, Darrell R. (1995). "Stabilization of RNA stacking by pseudouridine" (in en). Nucleic Acids Research 23 (24): 5020–5026. doi:10.1093/nar/23.24.5020. ISSN 0305-1048. PMID 8559660.
- ↑ Bussotti, Giovanni; Piel, Laura; Pescher, Pascale; Domagalska, Malgorzata A.; Rajan, K. Shanmugha; Cohen-Chalamish, Smadar; Doniger, Tirza; Hiregange, Disha-Gajanan et al. (21 December 2021). "Genome instability drives epistatic adaptation in the human pathogen Leishmania" (in en). Proceedings of the National Academy of Sciences 118 (51): e2113744118. doi:10.1073/pnas.2113744118. ISSN 0027-8424. PMID 34903666. Bibcode: 2021PNAS..11813744B.
- ↑ Ge, Junhui; Yu, Yi-Tao (April 2013). "RNA pseudouridylation: new insights into an old modification". Trends in Biochemical Sciences 38 (4): 210–218. doi:10.1016/j.tibs.2013.01.002. ISSN 0968-0004. PMID 23391857.
- ↑ 8.00 8.01 8.02 8.03 8.04 8.05 8.06 8.07 8.08 8.09 8.10 Rintala-Dempsey, Anne C.; Kothe, Ute (2017-01-03). "Eukaryotic stand-alone pseudouridine synthases – RNA modifying enzymes and emerging regulators of gene expression?". RNA Biology 14 (9): 1185–1196. doi:10.1080/15476286.2016.1276150. ISSN 1547-6286. PMID 28045575.
- ↑ Wu, Guowei; Radwan, Mohamed K.; Xiao, Mu; Adachi, Hironori; Fan, Jason; Yu, Yi-Tao (2016-06-07). "TheTORsignaling pathway regulates starvation-induced pseudouridylation of yeast U2 snRNA". RNA 22 (8): 1146–1152. doi:10.1261/rna.056796.116. ISSN 1355-8382. PMID 27268497.
- ↑ Adachi, Hironori; De Zoysa, Meemanage D.; Yu, Yi-Tao (March 2019). "Post-transcriptional pseudouridylation in mRNA as well as in some major types of noncoding RNAs". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 1862 (3): 230–239. doi:10.1016/j.bbagrm.2018.11.002. ISSN 1874-9399. PMID 30414851.
- ↑ "European medicines Agency Assessment report on Comirnaty (Common name: COVID-19 mRNA vaccine) (nucleoside-modified) Procedure No. EMEA/H/C/005735/0000". 2021-02-19. https://www.ema.europa.eu/en/documents/assessment-report/comirnaty-epar-public-assessment-report_en.pdf.
- ↑ 12.0 12.1 12.2 Penzo, M.; Guerrieri, A. N.; Zacchini, F.; Treré, D.; Montanaro, L. (2017-11-01). "RNA Pseudouridylation in Physiology and Medicine: For Better and for Worse". Genes 8 (11): 301. doi:10.3390/genes8110301. ISSN 2073-4425. PMID 29104216.
- ↑ Keffer-Wilkes, Laura Carole; Veerareddygari, Govardhan Reddy; Kothe, Ute (2016-11-14). "RNA modification enzyme TruB is a tRNA chaperone". Proceedings of the National Academy of Sciences 113 (50): 14306–14311. doi:10.1073/pnas.1607512113. ISSN 0027-8424. PMID 27849601. Bibcode: 2016PNAS..11314306K.
- ↑ Xu, J.; Gu, A. Y.; Thumati, N. R.; Wong JMY (2017-09-05). "Quantification of Pseudouridine Levels in Cellular RNA Pools with a Modified HPLC-UV Assay". Genes 8 (9): 219. doi:10.3390/genes8090219. ISSN 2073-4425. PMID 28872587.
- ↑ Kalsotra, Auinash (2016-11-02). Faculty of 1000 evaluation for Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA.. doi:10.3410/f.718875945.793524920.
- ↑ 16.0 16.1 Zhao, Yang; Karijolich, John; Glaunsinger, Britt; Zhou, Qiang (October 2016). "Pseudouridylation of 7 SK sn RNA promotes 7 SK sn RNP formation to suppress HIV ‐1 transcription and escape from latency". EMBO Reports 17 (10): 1441–1451. doi:10.15252/embr.201642682. ISSN 1469-221X. PMID 27558685.
- ↑ Dolgin, Elie (September 14, 2021). "The tangled history of mRNA vaccines". Nature. https://www.nature.com/articles/d41586-021-02483-w.
- ↑ "The Nobel Prize in Physiology or Medicine 2023" (in en-US). https://www.nobelprize.org/prizes/medicine/2023/press-release/.
- ↑ 19.0 19.1 "The Critical Contribution of Pseudouridine to mRNA COVID-19 Vaccines". Frontiers in Cell and Developmental Biology 9: 789427. 2021. doi:10.3389/fcell.2021.789427. PMID 34805188.
 |