Biology:Fragmentation (cell biology)
This article has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
(Learn how and when to remove this template message)
|
Fragmentation describes the process of splitting into several pieces or fragments. In cell biology, fragmentation is useful for a cell during both DNA cloning and apoptosis. DNA cloning is important in asexual reproduction or creation of identical DNA molecules, and can be performed spontaneously by the cell or intentionally by laboratory researchers. Apoptosis is the programmed destruction of cells, and the DNA molecules within them, and is a highly regulated process. These two ways in which fragmentation is used in cellular processes describe normal cellular functions and common laboratory procedures performed with cells. However, problems within a cell can sometimes cause fragmentation that results in irregularities such as red blood cell fragmentation and sperm cell DNA fragmentation.
DNA Cloning
DNA cloning can be performed spontaneously by the cell for reproductive purposes. This is a form of asexual reproduction where an organism splits into fragments and then each of these fragments develop into mature, fully grown individuals that are clones of the original organism (See reproductive fragmentation). DNA cloning can also be performed intentionally by laboratory researchers. Here, DNA fragmentation is a molecular genetic technique that permits researchers to use recombinant DNA technology to prepare large numbers of identical DNA molecules. In order for DNA cloning to be completed, it is necessary to obtain discrete, small regions of an organism's DNA that constitute specific genes. Only relatively small DNA molecules can be cloned in any available vector. Therefore, the long DNA molecules that compose an organism's genome must be cleaved into fragments that can be inserted into the vector DNA.[1] Two enzymes facilitate the production of such recombinant DNA molecules:
- 1. Restriction Enzymes
- Restriction enzymes are endonucleases produced by bacteria that typically recognize small base pair sequences (called restriction sites) and then cleave both strands of DNA at this site.[2] A restriction site is typically a palindromic sequence, which means that the restriction-site sequence is the same on each strand of DNA when read in the 5' to 3' direction.
- For each restriction enzyme, bacteria also produce a modification enzyme so that a host bacterium's own DNA is protected from cleavage. This is done by modifying the host DNA at or near each potential cleavage site. The modification enzyme adds a methyl group to one or two bases, and the presence of this methyl group prevents the restriction endonuclease from cutting the DNA.[3]
- Restriction enzymes are endonucleases produced by bacteria that typically recognize small base pair sequences (called restriction sites) and then cleave both strands of DNA at this site.[2] A restriction site is typically a palindromic sequence, which means that the restriction-site sequence is the same on each strand of DNA when read in the 5' to 3' direction.
- Many restriction enzymes make staggered cuts in the two DNA strands at their recognition site, which generates fragments with a single stranded "tail" that overhangs at both ends, called a sticky end. Restriction enzymes can also make straight cuts in the two DNA strands at their recognition site, which generates blunt ends.[4]
- 2. DNA ligase
- During normal DNA replication, DNA ligase catalyzes end-to-end joining (ligation) of short fragments of DNA, called Okazaki fragments. For the purposes of DNA cloning, purified DNA ligase is used to covalently join the ends of a restriction fragment and vector DNA that have complementary ends. They are covalently ligated together through the standard 3' to 5' phosphodiester bonds of DNA.[5]
- DNA ligase can ligate complementary sticky and blunt ends, but blunt-end ligation is inefficient and requires a higher concentration of both DNA and DNA ligase than the ligation of sticky ends does.[6] For this reason, most restriction enzymes used in DNA cloning make staggered cuts in the DNA strands to create sticky ends.
- During normal DNA replication, DNA ligase catalyzes end-to-end joining (ligation) of short fragments of DNA, called Okazaki fragments. For the purposes of DNA cloning, purified DNA ligase is used to covalently join the ends of a restriction fragment and vector DNA that have complementary ends. They are covalently ligated together through the standard 3' to 5' phosphodiester bonds of DNA.[5]
The key to cloning a DNA fragment is to link it to a vector DNA molecule that can replicate within a host cell. After a single recombinant DNA molecule (composed of a vector plus an inserted DNA fragment) is introduced into a host cell, the inserted DNA can be replicated along with the vector, generating a large number of identical DNA molecules.[7] The basic scheme for this can be summarized as follows:
- Vector + DNA Fragment
- ↓
- Recombinant DNA
- ↓
- Replication of recombinant DNA within host cell
- ↓
- Isolation, sequencing, and manipulation of purified DNA fragment
- Vector + DNA Fragment
There are numerous experimental variations to this scheme, but these steps are essential to DNA cloning in a laboratory.[8]
Apoptosis
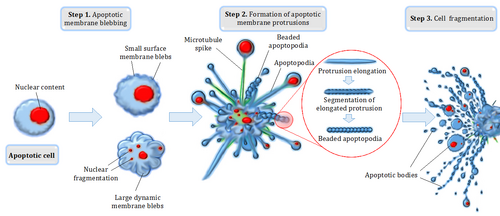
Apoptosis refers to the demise of cells by a specific form of programmed cell death, characterized by a well-defined sequence of morphological changes.[10] Cellular and nuclear shrinkage, chromatin condensation and fragmentation, formation of apoptotic bodies and phagocytosis by neighboring cells characterize the main morphological changes in the apoptosis process.[11] Extensive morphological and biochemical changes during apoptosis ensure that dying cells leave minimal impact on neighboring cells and/or tissues.
Genes involved in controlling cell death encode proteins with three distinct functions:[12]
- "Killer" proteins are required for a cell to begin the apoptotic process
- "Destruction" proteins do things such as digest DNA in a dying cell
- "Engulfment" proteins are required for phagocytosis of the dying cell by another cell
The cleavage of chromosomal DNA into smaller fragments is an integral part, and biochemical hallmark, of apoptosis. Apoptosis involves the activation of endonucleases with subsequent cleavage of chromatin DNA into fragments of 180 base pairs or multiples of 180 base pairs (e.g. 360, 540). This pattern of fragmentation can be used to detect apoptosis in tests such as a DNA laddering assay with gel electrophoresis, a TUNEL assay, or a Nicoletti assay.[13] Apoptotic DNA fragmentation relies on an enzyme called Caspase-Activated DNase (CAD).[14] CAD is usually inhibited by another protein in the cell, called Inhibitor of caspase-activated DNase (ICAD).[15] In order for apoptosis to begin, an enzyme called caspase 3 cleaves ICAD so that CAD becomes activated. CAD then cleaves the DNA between nucleosomes, which occur in chromatin at 180 base pair intervals. The sites between nucleosomes are the only parts of the DNA that are exposed and accessible to CAD.[16]
Irregularities
DNA fragmentation can occur under certain conditions in a few different cell types. This can lead to problems for a cell, or it may lead to a cell receiving a signal to undergo apoptosis. Below are a couple of examples of irregular fragmentation that can occur in cells.
- 1. Red blood cell fragmentation
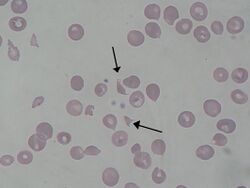
- A fragmented red blood cell is known as a schistocyte and is generally the result of an intracellular mechanical injury to the red blood cell.[17] A wide variety of schistocytes may be observed. Schistocytes are usually seen in relatively low numbers and are associated with conditions in which the normally smooth endothelial lining, or endothelium, is roughened or irregular, and/or the vascular lumen is crossed by strands of fibrin.[18] Schistocytes are commonly seen in patients that have hemolytic anemia. They are also a feature of advanced iron deficiency anemia, but in this case the observed fragmentation is most likely a result of the fragility of the cells produced under these conditions.
- 2. Sperm cell DNA fragmentation
- In an average male, less than 4% of his sperm cells will contain fragmented DNA. However, partaking in behaviors such as smoking can significantly increase DNA fragmentation in sperm cells. There is a negative correlation between the percentage of DNA fragmentation and the motility, morphology, and concentration of sperm. There is also a negative association between the percentage of sperm that contain fragmented DNA and the fertilization rate and embryo cleavage rate.[19]
References
- ↑ Lodish, Harvey, Arnold Berk, Chris A. Kaiser, Monty Kriger, Anthony Bretscher, Hidde Ploegh, Angelika Amon, and Matthew P. Scott. Molecular Cell Biology. 7th ed. New York: W.H. Freeman and, 2013. Print.
- ↑ Rao, Desirazu N., Swati Saha, and Vinita Krishnamurthy. "ATP-Dependent Restriction Enzymes." Progress in Nucleic Acid Research and Molecular Biology 64 (2000): 1-63. Print.
- ↑ Rao, Desirazu N., Swati Saha, and Vinita Krishnamurthy. "ATP-Dependent Restriction Enzymes." Progress in Nucleic Acid Research and Molecular Biology 64 (2000): 1-63. Print.
- ↑ Lodish, Harvey, Arnold Berk, Chris A. Kaiser, Monty Kriger, Anthony Bretscher, Hidde Ploegh, Angelika Amon, and Matthew P. Scott. Molecular Cell Biology. 7th ed. New York: W.H. Freeman and, 2013. Print.
- ↑ Tomkinson, Alan E., and Zachary B. Mackey. "Structure and Function of Mammalian DNA Ligases." Mutation Research/DNA Repair 407.1 (1998): 1-9. Print.
- ↑ Hung, Mien-Chie, and Pieter C. Wensink. "Different Restriction Enzyme-generated Sticky DNA Ends Can Be Joined in Vitro." Nucleic Acids Research 12.4 (1984): 1863-874. Print.
- ↑ "Ch 20." Avonapbio /. N.p., n.d. Web. 20 Nov. 2012. <http://avonapbio.pbworks.com/w/page/9429274/Ch%2020>.
- ↑ Lodish, Harvey, Arnold Berk, Chris A. Kaiser, Monty Kriger, Anthony Bretscher, Hidde Ploegh, Angelika Amon, and Matthew P. Scott. Molecular Cell Biology. 7th ed. New York: W.H. Freeman and, 2013. Print.
- ↑ Smith, Aaron; Parkes, Michael AF; Atkin-Smith, Georgia K; Tixeira, Rochelle; Poon, Ivan KH (2017). "Cell disassembly during apoptosis" (in en). WikiJournal of Medicine 4 (1). doi:10.15347/wjm/2017.008. https://en.wikiversity.org/wiki/WikiJournal_of_Medicine/Cell_disassembly_during_apoptosis.
- ↑ Lodish, Harvey, Arnold Berk, Chris A. Kaiser, Monty Kriger, Anthony Bretscher, Hidde Ploegh, Angelika Amon, and Matthew P. Scott. Molecular Cell Biology. 7th ed. New York: W.H. Freeman and, 2013. Print.
- ↑ Hua, Xhang J., and Ming Xu. "DNA Fragmentation in Apoptosis." Cell Research 10 (2000): 205-11. Nature. 17 July 2000. Web. 19 Nov. 2012.
- ↑ Lodish, Harvey, Arnold Berk, Chris A. Kaiser, Monty Kriger, Anthony Bretscher, Hidde Ploegh, Angelika Amon, and Matthew P. Scott. Molecular Cell Biology. 7th ed. New York: W.H. Freeman and, 2013. Print.
- ↑ Bortner, Carl D., Nicklas B.E. Oldenburg, and John A. Cidlowski. "The Role of DNA Fragmentation in Apoptosis." Trends in Cell Biology 5.1 (1995): 21-26. Print.
- ↑ Jog, Neelakshi R., Lorenza Frisoni, Qin Shi, Marc Monestier, Sairy Hernandez, Joe Craft, Eline T. Luning Prak, and Roberto Caricchio. "Caspase-activated DNase Is Required for Maintenance of Tolerance to Lupus Nuclear Autoantigens." Arthritis and Rheumatism 64.4 (2012): 1247-256. Print.
- ↑ Kutscher, Daniel, Alfred Pingoud, Albert Jeltsch, and Gregor Meiss. "Identification of ICAD-derived Peptides Capable of Inhibiting Caspase-activated DNase." FEBS Journal 279.16 (2012): 2917-928. Print.
- ↑ Bortner, Carl D., Nicklas B.E. Oldenburg, and John A. Cidlowski. "The Role of DNA Fragmentation in Apoptosis." Trends in Cell Biology 5.1 (1995): 21-26. Print.
- ↑ Bessman, JD. "Red Blood Cell Fragmentation. Improved Detection and Identification of Causes." American Journal of Clinical Pathology 90.3 (1988): 268-73. Print.
- ↑ "Schistocytes." Schistocytes. N.p., n.d. Web. 20 Nov. 2012. <http://ahdc.vet.cornell.edu/clinpath/modules/rbcmorph/schisto.htm>.
- ↑ Sun, J. G., A. Jurisicova, and R. F. Casper. "Detection of Deoxyribonucleic Acid Fragmentation in Human Sperm: Correlation with Fertilization in Vitro." Biology of Reproduction 56.3 (1997): 602-07. Print.
 |