Biology:Pseudomonas fluorescens
| Pseudomonas fluorescens | |
|---|---|
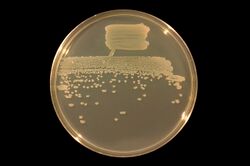
| |
| Pseudomonas fluorescens under white light | |

| |
| The same plate under UV light | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Pseudomonadales |
| Family: | Pseudomonadaceae |
| Genus: | Pseudomonas |
| Species: | P. fluorescens
|
| Binomial name | |
| Pseudomonas fluorescens (Flügge 1886)
Migula, 1895 | |
| Type strain | |
| ATCC 13525 CCUG 1253 | |
| Synonyms | |
|
Bacillus fluorescens liquefaciens Flügge 1886 | |
Pseudomonas fluorescens is a common Gram-negative, rod-shaped bacterium.[1] It belongs to the Pseudomonas genus; 16S rRNA analysis as well as phylogenomic analysis has placed P. fluorescens in the P. fluorescens group within the genus,[2][3] to which it lends its name.
General characteristics
Pseudomonas fluorescens has multiple flagella. It has an extremely versatile metabolism, and can be found in the soil and in water. It is an obligate aerobe, but certain strains are capable of using nitrate instead of oxygen as a final electron acceptor during cellular respiration.
Optimal temperatures for growth of P. fluorescens are 25–30°C. It tests positive for the oxidase test. It is also a nonsaccharolytic bacterial species.
Heat-stable lipases and proteases are produced by P. fluorescens and other similar pseudomonads.[4] These enzymes cause milk to spoil, by causing bitterness, casein breakdown, and ropiness due to production of slime and coagulation of proteins.[5][6]
The name
The word Pseudomonas means false unit, being derived from the Greek words pseudēs (Greek: ψευδής – false) and monas (Latin: monas, from Greek: μονάς – a single unit). The word was used early in the history of microbiology to refer to germs. The specific name fluorescens refers to the microbe's secretion of a soluble fluorescent pigment called pyoverdin, which is a type of siderophore.[7]
Genomics
Notable P. fluorescens strains SBW25,[8] Pf-5[9] and PfO-1[10] have been sequenced, among others.
A comparative genomic study (in 2020) analyzed 494 complete genomes from the entire Pseudomonas genus, with 25 of them being annotated as P. fluorescens.[3] The phylogenomic analysis clearly showed that the 25 strains annotated as P. fluorescens did not form a monophyletic group.[3] In addition, their Average Nucleotide Identities did not fulfil the criteria of a species, since they were very diverse. It was concluded that P. fluorescens is not a species in the strict sense, but should be considered as a wider evolutionary group, or a species complex, that includes within it other species too.[3] This finding is in accordance with previous analyses of 107 Pseudomonas species, using four core 'housekeeping' genes, that consider P. fluorescens as a relaxed species complex.[11]
The P. fluorescens relaxed evolutionary group that was defined in,[3] on the basis of the genus phylogenomic tree, comprised 96 genomes and displayed high levels of phylogenetic heterogeneity. It comprised many species, such as Pseudomonas corrugata, Pseudomonas brassicacearum, Pseudomonas frederiksbergensis, Pseudomonas mandelii, Pseudomonas kribbensis, Pseudomonas koreensis, Pseudomonas mucidolens, Pseudomonas veronii, Pseudomonas antarctica, Pseudomonas azotoformans, Pseudomonas trivialis, Pseudomonas lurida, Pseudomonas azotoformans, Pseudomonas poae, Pseudomonas libanensis, Pseudomonas synxantha, and Pseudomonas orientalis. The core proteome of the P. fluorescens group comprised 1396 proteins. The protein count and GC content of the strains of the P. fluorescens group ranged between 4152 and 6678 (average: 5603) and between 58.7–62% (average: 60.3%), respectively. Another comparative genomic analysis of 71 P. fluorescens genomes identified eight major subgroups and developed a set of nine genes as markers for classification within this lineage.[12]
Interactions with Dictyostelium
There are two strains of Pseudomonas fluorescens associated with Dictyostelium discoideum. One strain serves as a food source and the other strain does not. The main genetic difference between these two strains is a mutation of the global activator gene called gacA. This gene plays a key role in gene regulation; when this gene is mutated in the nonfood bacterial strain, it is transformed into a food bacterial strain.[13]
Biocontrol properties
Some P. fluorescens strains (CHA0 or Pf-5, for example) present biocontrol properties, protecting the roots of some plant species against parasitic fungi such as Fusarium or the oomycete Pythium, as well as some phytophagous nematodes.[14]
It is not clear exactly how the plant growth-promoting properties of P. fluorescens are achieved; theories include:
- The bacteria might induce systemic resistance in the host plant, so it can better resist attack by a true pathogen.
- The bacteria might outcompete other (pathogenic) soil microbes, e.g., by siderophores, giving a competitive advantage at scavenging for iron.
- The bacteria might produce compounds antagonistic to other soil microbes, such as phenazine-type antibiotics or hydrogen cyanide.
To be specific, certain P. fluorescens isolates produce the secondary metabolite 2,4-diacetylphloroglucinol (2,4-DAPG), the compound found to be responsible for antiphytopathogenic and biocontrol properties in these strains.[15] The phl gene cluster encodes factors for 2,4-DAPG biosynthesis, regulation, export, and degradation. Eight genes, phlHGFACBDE, are annotated in this cluster and conserved organizationally in 2,4-DAPG-producing strains of P. fluorescens. Of these genes, phlD encodes a type III polyketide synthase, representing the key biosynthetic factor for 2,4-DAPG production. PhlD shows similarity to plant chalcone synthases and has been theorized to originate from horizontal gene transfer.[15] Phylogenetic and genomic analysis, though, has revealed that the entire phl gene cluster is ancestral to P. fluorescens, many strains have lost the capacity, and it exists on different genomic regions among strains.[16]
Some experimental evidence supports all of these theories, in certain conditions; a good review of the topic is written by Haas and Defago.[17]
Several strains of P. fluorescens, such as Pf-5 and JL3985, have developed a natural resistance to ampicillin and streptomycin.[18] These antibiotics are regularly used in biological research as a selective pressure tool to promote plasmid expression.
The strain referred to as Pf-CL145A has proved itself a promising solution for the control of invasive zebra mussels and quagga mussels (Dreissena). This bacterial strain is an environmental isolate capable of killing >90% of these mussels by intoxication (i.e., not infection), as a result of natural product(s) associated with their cell walls, and with dead Pf-145A cells killing the mussels equally as well as live cells.[19] Following ingestion of the bacterial cells mussel death occurs following lysis and necrosis of the digestive gland and sloughing of stomach epithelium.[20] Research to date indicates very high specificity to zebra and quagga mussels, with low risk of nontarget impact.[21] Pf-CL145A has now been commercialized under the product name Zequanox, with dead bacterial cells as its active ingredient.
Recent results showed the production of the phytohormone cytokinin by P. fluorescens strain G20-18 to be critical for its biocontrol activity by activating plant resistance.[22]
Medical properties
By culturing P. fluorescens, mupirocin (an antibiotic) can be produced, which has been found to be useful in treating skin, ear, and eye disorders.[23] Mupirocin free acid and its salts and esters are agents currently used in creams, ointments, and sprays as a treatment of methicillin-resistant Staphylococcus aureus infection.
Pseudomonas fluorescens demonstrates hemolytic activity, and as a result, has been known to infect blood transfusions.[24]
Pseudomonas fluorescens produces the antibiotic Obafluorin.[25][26]
Disease
Pseudomonas fluorescens is an unusual cause of disease in humans, and usually affects patients with compromised immune systems (e.g., patients on cancer treatment). From 2004 to 2006, an outbreak of P. fluorescens in the United States involved 80 patients in six states. The source of the infection was contaminated heparinized saline flushes being used with cancer patients.[27]
Pseudomonas fluorescens is also a known cause of fin rot in fish.
Metabolism
Pseudomonas fluorescens produces phenazine, phenazine carboxylic acid,[28] 2,4-diacetylphloroglucinol[29] and the MRSA-active antibiotic mupirocin.[30]
Biodegradation capacities
4-Hydroxyacetophenone monooxygenase is an enzyme found in P. fluorescens that transforms piceol, NADPH, H+, and O2 into 4-hydroxyphenyl acetate, NADP+, and H2O.
References
- ↑ Palleroni, N.J. (1984) Pseudomonadaceae. Bergey's Manual of Systematic Bacteriology. Krieg, N. R. and Holt J. G. (editors) Baltimore: The Williams and Wilkins Co., pg. 141 – 199
- ↑ Anzai et al. (Jul 2000). "Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence". Int J Syst Evol Microbiol 50 (4): 1563–89. doi:10.1099/00207713-50-4-1563. PMID 10939664.
- ↑ 3.0 3.1 3.2 3.3 3.4 Nikolaidis, Marios; Mossialos, Dimitris; Oliver, Stephen G.; Amoutzias, Grigorios D. (2020-07-24). "Comparative Analysis of the Core Proteomes among the Pseudomonas Major Evolutionary Groups Reveals Species-Specific Adaptations for Pseudomonas aeruginosa and Pseudomonas chlororaphis" (in en). Diversity 12 (8): 289. doi:10.3390/d12080289. ISSN 1424-2818.
 Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Frank, J.F. 1997. Milk and dairy products. In Food Microbiology, Fundamentals and Frontiers, ed. M.P. Doyle, L.R. Beuchat, T.J. Montville, ASM Press, Washington, p. 101.
- ↑ Jay, J.M. 2000. Taxonomy, role, and significance of microorganisms in food. In Modern Food Microbiology, Aspen Publishers, Gaithersburg MD, p. 13.
- ↑ Ray, B. 1996. Spoilage of Specific food groups. In Fundamental Food Microbiology, CRC Press, Boca Raton FL, p. 220. I
- ↑ C D Cox and P Adams (1985) Infection and Immunity 48(1): 130–138
- ↑ Pseudomonas fluorescens
- ↑ "Pseudomonas fluorescens Pf-5 Genome Page". http://cmr.jcvi.org/tigr-scripts/CMR/GenomePage.cgi?org=gpf.
- ↑ "Pseudomonas fluorescens PfO-1 Genome Page". http://cmr.jcvi.org/tigr-scripts/CMR/GenomePage.cgi?org=ntpf02.
- ↑ Mulet, Magdalena; Lalucat, Jorge; García-Valdés, Elena (March 2010). "DNA sequence-based analysis of the Pseudomonas species" (in en). Environmental Microbiology 12 (6): 1513–1530. doi:10.1111/j.1462-2920.2010.02181.x. PMID 20192968. http://doi.wiley.com/10.1111/j.1462-2920.2010.02181.x.
- ↑ Garrido-Sanz, Daniel; Arrebola, Eva; Martínez-Granero, Francisco; García-Méndez, Sonia; Muriel, Candela; Blanco-Romero, Esther; Martín, Marta; Rivilla, Rafael et al. (2017-03-15). "Classification of Isolates from the Pseudomonas fluorescens Complex into Phylogenomic Groups Based in Group-Specific Markers". Frontiers in Microbiology 8: 413. doi:10.3389/fmicb.2017.00413. ISSN 1664-302X. PMID 28360897.
- ↑ Stallforth, Pierre; Brock, Debra A.; Cantley, Alexandra M.; Tian, Xiangjun; Queller, David C.; Strassmann, Joan E.; Clardy, Jon (2013-09-03). "A bacterial symbiont is converted from an inedible producer of beneficial molecules into food by a single mutation in the gacA gene". Proceedings of the National Academy of Sciences of the United States of America 110 (36): 14528–14533. doi:10.1073/pnas.1308199110. ISSN 0027-8424. PMID 23898207. Bibcode: 2013PNAS..11014528S.
- ↑ Haas, D.; Keel, C. (2003). "Regulation of antibiotic production in root-colonizing Pseudomonas spp. and relevance for biological control of plant disease". Annual Review of Phytopathology 41: 117–153. doi:10.1146/annurev.phyto.41.052002.095656. PMID 12730389.
- ↑ 15.0 15.1 Bangera M. G.; Thomashow L. S. (1999). "Identification and characterization of a gene cluster for synthesis of the polyketide antibiotic 2,4-diacetylphloroglucinol from pseudomonas fluorescens q2-87". Journal of Bacteriology 181 (10): 3155–3163. doi:10.1128/JB.181.10.3155-3163.1999. PMID 10322017.
- ↑ Moynihan J. A.; Morrissey J. P.; Coppoolse E. R.; Stiekema W. J.; O'Gara F.; Boyd E. F. (2009). "Evolutionary history of the phl gene cluster in the plant-associated bacterium pseudomonas fluorescens". Applied and Environmental Microbiology 75 (7): 2122–2131. doi:10.1128/aem.02052-08. PMID 19181839. Bibcode: 2009ApEnM..75.2122M.
- ↑ Haas, D; Defago, G (2005). "Biological control of soil-borne pathogens by fluorescent pseudomonads". Nature Reviews Microbiology 3 (4): 307–19. doi:10.1038/nrmicro1129. PMID 15759041.
- ↑ Alain Sarniguet (1995). "The sigma factor σs affects antibiotic production and biological control activity of Pseudomonas fluorescens Pf-5". Proc. Natl. Acad. Sci. U.S.A. 92 (26): 12255–12259. doi:10.1073/pnas.92.26.12255. PMID 8618880. Bibcode: 1995PNAS...9212255S.
- ↑ Molloy, D. P., Mayer, D. A., Gaylo, M. J., Morse, J. T., Presti, K. T., Sawyko, P. M., Karatayev, A. Y., Burlakova, L. E., Laruelle, F., Nishikawa, K. C., Griffin, B. H. 2013. Pseudomonas fluorescens strain CL145A – A biopesticide for the control of zebra and quagga mussels (Bivalvia: Dreissenidae). J. Invertebr. Pathol. 113(1):104–114.
- ↑ Molloy, D. P., Mayer, D. A., Giamberini, L., and Gaylo, M. J. 2013. Mode of action of Pseudomonas fluorescens strain CL145A, a lethal control agent of dreissenid mussels (Bivalvia: Dreissenidae). J. Invertebr. Pathol. 113(1):115–121.
- ↑ Molloy, D. P.; Mayer, D. A.; Gaylo, M. J.; Burlakova, L. E.; Karatayev, A. Y.; Presti, K. T.; Sawyko, P. M.; Morse, J. T. et al. (2013). "Non-target trials with Pseudomonas fluorescens strain CL145A, a lethal control agent of dreissenid mussels (Bivalvia: Dreissenidae)". Manag. Biol. Invasions 4 (1): 71–79. doi:10.3391/mbi.2013.4.1.09.
- ↑ "Cytokinin production by Pseudomonas fluorescens G20-18 determines biocontrol activity against Pseudomonas syringae in Arabidopsis". Scientific Reports 6: 23310. 2016. doi:10.1038/srep23310. PMID 26984671. Bibcode: 2016NatSR...623310G.
- ↑ Bactroban
- ↑ "Rate of growth of Pseudomonas fluorescens in donated blood". Journal of Clinical Pathology 48 (8): 717–8. 1995. doi:10.1136/jcp.48.8.717. PMID 7560196.
- ↑ Wells, J. Scott; Trejo, William H.; Principe, Pacifico A.; Sykes, Richard B. (1984). "Obafluorin, a novel .BETA.-lactone produced by Pseudomonas fluorescens. Taxonomy, fermentation and biological properties.". The Journal of Antibiotics 37 (7): 802–803. doi:10.7164/antibiotics.37.802. PMID 6432765.
- ↑ Tymiak, Adrienne A.; Culver, Catherine A.; Malley, Mary F.; Gougoutas, Jack Z. (December 1985). "Structure of obafluorin: an antibacterial .beta.-lactone from Pseudomonas fluorescens". The Journal of Organic Chemistry 50 (26): 5491–5495. doi:10.1021/jo00350a010.
- ↑ "Multistate outbreak of Pseudomonas fluorescens bloodstream infection after exposure to contaminated heparinized saline flush prepared by a compounding pharmacy". Clin Infect Dis 47 (11): 1372–1379. 2008. doi:10.1086/592968. PMID 18937575.
- ↑ Mavrodi, D.V.; Ksenzenko, V. N.; Bonsall, R. F.; Cook, R. J.; Boronin, A. M.; Thomashow, L. S. (1998). "A seven-gene locus for synthesis of phenazine-1-carboxylic acid by Pseudomonas fluorescens 2–79". J. Bacteriol. 180 (9): 2541–2548. doi:10.1128/JB.180.9.2541-2548.1998. PMID 9573209.
- ↑ Achkar, Jihane; Xian, Mo; Zhao, Huimin; Frost, J. W. (2005). "Biosynthesis of Phloroglucinol". J. Am. Chem. Soc. 127 (15): 5332–5333. doi:10.1021/ja042340g. PMID 15826166.
- ↑ Fuller, AT; Mellows, G; Woolford, M; Banks, GT; Barrow, KD; Chain, EB (1971). "Pseudomonic acid: an antibiotic produced by Pseudomonas fluorescens". Nature 234 (5329): 416–417. doi:10.1038/234416a0. PMID 5003547. Bibcode: 1971Natur.234..416F.
Further reading
Appanna, Varun P.; Auger, Christopher; Thomas, Sean C.; Omri, Abdelwahab (13 June 2014). "Fumarate metabolism and ATP production in Pseudomonas fluorescens exposed to nitrosative stress". Antonie van Leeuwenhoek 106 (3): 431–438. doi:10.1007/s10482-014-0211-7. PMID 24923559.
Cabrefiga, J.; Frances, J.; Montesinos, E.; Bonaterra, A. (1 October 2014). "Improvement of a dry formulation of Pseudomonas fluorescens EPS62e for fire blight disease biocontrol by combination of culture osmoadaptation with a freeze-drying lyoprotectant". Journal of Applied Microbiology 117 (4): 1122–1131. doi:10.1111/jam.12582. PMID 24947806.
External links
- The Pseudomonas Genome Database
- Type strain of Pseudomonas fluorescens at BacDive – the Bacterial Diversity Metadatabase
Wikidata ☰ Q135292 entry
 |

