Biology:Actin assembly-inducing protein
| Actin assembly-inducing protein | |
|---|---|
 EVH1 domain-ActA peptide complex | |
| Identifiers | |
| Symbol | ActA |
| NCBI gene | 2798121 |
| UniProt | P33379 |
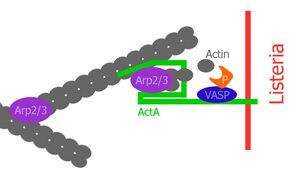
The Actin assembly-inducing protein (ActA) is a protein encoded and used by Listeria monocytogenes to propel itself through a mammalian host cell. ActA is a bacterial surface protein comprising a membrane-spanning region.[1] In a mammalian cell the bacterial ActA interacts with the Arp2/3 complex and actin monomers to induce actin polymerization on the bacterial surface generating an actin comet tail. The gene encoding ActA is named actA or prtB.[2]
Introduction
As soon as L. monocytogenes bacteria are ingested by humans, they get internalized into intestinal epithelium cells and rapidly try to escape their internalization vacuole.[3][4] In the cytosol they start to polymerize actin on their surface by the help of the ActA protein. It has been shown that ActA is not only necessary but also sufficient to induce motility of bacteria in the absence of other bacterial factors.[5]
Discovery
ActA was discovered by analysing lecithinase-negative Tn917-lac Listeria mutants because of the phenotype that they were unable to spread from cell to cell. These mutant bacteria still escaped from the phagosomes as efficiently as wild-type bacteria and multiplied within the infected cells but they were not surrounded by actin like wild-type bacteria. Further analysis showed, that Tn917-lac had inserted into actA, the second gene of an operon. The third gene of this operon, plcB, encodes the L. monocytogenes lecithinase. To determine whether actA itself, plcB or other co-transcribed downstream regions are involved in actin assembly, mutations in the appropriate genes were generated. All mutants except the actA mutants were similar to wild-type concerning association with F-actin and cell-cell spreading. Complementation with actA restored wild-type phenotype in the actA mutants.[1]
Function
ActA is a protein which acts as a mimic of Wiskott-Aldrich syndrome protein (WASP), a nucleation promoting factor (NPF) present in host cells. NPFs in the mammalian cell recruit and bind to the already existing actin-related-protein 2 and 3 complex (Arp2/3 complex) and induce an activating conformational change of the Arp2/3 complex.[6] Due to this conformational change, NPFs initiate polymerization of a new actin filament at a 70° angle, which leads to the characteristic Y-branched actin structures in the leading edge of motile cells. ActA localizes to the old pole of the bacterium and spans both the bacterial cell membrane and the cell wall, lateral diffusion is inhibited; thus ActA localizes in a polarized and anchored manner on the bacterial surface. Consequently, actin polymerization only starts in this region on the surface of the bacterium.[7] Expression of ActA is induced only after entering a mammalian host cell.[8]
Actin filament assembly generates the force that pushes the bacterium in the mammalian host cytoplasm forward. Continuous actin polymerization is sufficient for motility in the cytoplasm and even for infection of adjacent cells.[9]
Research
New data indicates that ActA plays a role also in vacuolar disruption. A deletion mutant of ActA was defective in permeabilizing the vacuole. An 11 amino acid stretch of the N-terminus of the acidic region (32-42) was shown to be important for disruption of the phagosome.[10]
Structure
The primary proteinous product of the actA gene consists of 639 amino acids and includes the signal peptide (first N-terminal 29 amino acids) and the ActA chain (C-terminal 610 amino acids). Therefore, the sequence of the mature ActA protein consist of 610 amino acids. ActA has a molecular weight of 70,349 Da and is a surface protein.[1][2]
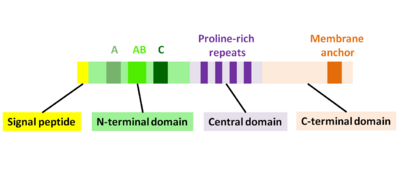
ActA is a natively unfolded protein which can be divided into three functional domains (Fig. 2):[1][11][12]
- N-terminal domain that is highly charged: amino acid residues 1-234
- central domain with proline-rich repeats: amino acid residues 235-394
- C-terminal domain with a transmembrane domain: amino acid residues 395-610
N-terminal domain
The first 156 amino acids of the N-terminal domain consist of three regions[10][13] (Fig. 2):
- A-region with a stretch of acidic residues: 32-45
- AB-region, an actin monomer-binding region: 59-102
- C-region, a cofilin homology sequence: 145-156
The N-terminal portion of ActA plays an important role in actin polymerization.[14] The domain displays consensus elements present in eukaryotic WASP family NPFs which include an actin monomer-binding region as well as an Arp2/3 binding C (central or cofilin homology) and A (acidic) region.[7] The actin monomer-binding region of ActA has functional properties like the WASP-Homology-2 (WH2) or V domain, but differs in the sequence.[15] Thus in WASP-family NPFs the order of the domains is WH2 followed by C, and then by A, which is not the case in ActA.
Central domain
The central proline-rich region of ActA is crucial for ensuring efficient bacterial motility. There are four proline-rich repeats containing either FPPPP or FPPIP motifs. These regions mimic those of the host cell cytoskeletal protein zyxin, vinculin and palladin, known to associate with focal adhesions or stress fibers.[16] The vasodilator-stimulated phosphoprotein (VASP) can bind through its Ena/VASP homology 1 domain (EVH1 domain) to the central proline-rich region and recruits profilin, an actin monomer binding protein, which itself promotes polymerization at barbed ends of actin filaments. Furthermore, VASP seems to interact with F-actin through its carboxy-terminal EVH2 domain, which provides a linkage of the bacterium to the tail.[17] This statement is supported by the fact that ActA can bind multiple Ena/VASP proteins simultaneously and has a high affinity between ActA and Ena/VASP. VASP has been shown to reduce the frequency actin-Y-branches in vitro and thus increases the proportion of filaments which are organized in a parallel alignment in comet tails.[18][19]
C-terminal domain
The C-terminal domain of ActA has a hydrophobic region which anchors the protein in the bacterial membrane.[20][21][22]
In summary, besides
- the absence of sequence homology in the actin-binding-region and
- an alteration in the sequence of ARP2/3 activating domains typical for WASP-family NPFs (V(WH2)-C-A),
- a major difference between ActA and host NPFs is that ActA does not have elements that bind to regulatory proteins such as Rho family GTPases. This structural difference between ActA and host NPFs can be advantageous for L. monocytogenes and its pathogenesis because the actin nucleation activity of L. monocytogenes is independent of host regulation.[7]
Analogues
WASP/N-WASP, which is functionally mimicked by ActA, is highly conserved in eukaryotes. It is an important actin-cytoskeleton organizer and is critical for processes such as endocytosis and cell motility. Activated by Cdc42, a Rho-family small GTPase, WASP/N-WASP activates the Arp2/3 complex, which leads to rapid actin polymerization.[23]
Actin-based motility of other pathogens
In Shigella the protein IcsA activates N-WASP, which in non-infected mammalian cells is activated by the GTPase Cdc42. Active N-WASP/WASP leads to actin polymerization by activating the Arp2/3 complex. In contrast, the Listeria ActA protein interacts with and activates directly the Arp2/3 complex.[7]
The Rickettsia RickA protein is also able to activate the Arp2/3 complex in a WASP-like manner. In contrast to Listeria, the actin filaments are organized in long, unbranched parallel bundles. The Arp2/3 complex is only localized near the bacterial surface and thus it is assumed that a more frequent Arp2/3 complex-independent elongation occurs.[16]
In Burkholderia pseudomallei BimA initiates actin polymerization in vitro. It is assumed that intracellular migration of this bacterium functions independently of the Arp2/3 complex.[16]
See also
References
- ↑ 1.0 1.1 1.2 1.3 "L. monocytogenes-induced actin assembly requires the actA gene product, a surface protein". Cell 68 (3): 521–31. February 1992. doi:10.1016/0092-8674(92)90188-I. PMID 1739966.
- ↑ 2.0 2.1 Uniprot P33379
- ↑ "Bacterial invasion: the paradigms of enteroinvasive pathogens". Science 304 (5668): 242–8. April 2004. doi:10.1126/science.1090124. PMID 15073367. Bibcode: 2004Sci...304..242C.
- ↑ "Invasion of mammalian cells by Listeria monocytogenes: functional mimicry to subvert cellular functions". Trends in Cell Biology 13 (1): 23–31. January 2003. doi:10.1016/S0962-8924(02)00006-5. PMID 12480337.
- ↑ Zigmond SH (February 2004). "Formin-induced nucleation of actin filaments". Current Opinion in Cell Biology 16 (1): 99–105. doi:10.1016/j.ceb.2003.10.019. PMID 15037312.
- ↑ "Critical conformational changes in the Arp2/3 complex are induced by nucleotide and nucleation promoting factor". Molecular Cell 16 (2): 269–79. October 2004. doi:10.1016/j.molcel.2004.09.018. PMID 15494313.
- ↑ 7.0 7.1 7.2 7.3 "Actin-based motility of intracellular pathogens". Current Opinion in Microbiology 8 (1): 35–45. February 2005. doi:10.1016/j.mib.2004.12.013. PMID 15694855.
- ↑ "Mechanism of polarization of Listeria monocytogenes surface protein ActA". Molecular Microbiology 59 (4): 1262–79. February 2006. doi:10.1111/j.1365-2958.2006.05025.x. PMID 16430699.
- ↑ Goldberg MB (December 2001). "Actin-Based Motility of Intracellular Microbial Pathogens". Microbiology and Molecular Biology Reviews 65 (4): 595–626. doi:10.1128/MMBR.65.4.595-626.2001. PMID 11729265.
- ↑ 10.0 10.1 "Evidence for involvement of ActA in maturation of the Listeria monocytogenes phagosome". Cell Research 20 (1): 109–12. January 2010. doi:10.1038/cr.2009.142. PMID 20029388.
- ↑ "Host-pathogen interactions during entry and actin-based movement of Listeria monocytogenes". Annual Review of Genetics 31: 113–38. 1997. doi:10.1146/annurev.genet.31.1.113. PMID 9442892.
- ↑ Footer, Matthew J.; Lyo, John K.; Theriot, Julie A. (2008-08-29). "Close packing of Listeria monocytogenes ActA, a natively unfolded protein, enhances F-actin assembly without dimerization". The Journal of Biological Chemistry 283 (35): 23852–23862. doi:10.1074/jbc.M803448200. ISSN 0021-9258. PMID 18577520.
- ↑ Welch, Matthew D. (2007). "Actin-Based Motility and Cell-to-Cell Spread of Listeria monocytogenes". in Goldfine, Howard; Shen, Hao. Listeria monocytogenes: Pathogenesis and Host Response. New York: Springer. pp. 197–223. doi:10.1007/978-0-387-49376-3_10. ISBN 978-0-387-49373-2.
- ↑ "Interaction of human Arp2/3 complex and the Listeria monocytogenes ActA protein in actin filament nucleation". Science 281 (5373): 105–8. July 1998. doi:10.1126/science.281.5373.105. PMID 9651243. Bibcode: 1998Sci...281..105W.
- ↑ "Activation of the Arp2/3 complex by the Listeria acta protein. Acta binds two actin monomers and three subunits of the Arp2/3 complex". The Journal of Biological Chemistry 276 (5): 3468–75. February 2001. doi:10.1074/jbc.M006407200. PMID 11029465.
- ↑ 16.0 16.1 16.2 "Listeria comet tails: the actin-based motility machinery at work". Trends in Cell Biology 18 (5): 220–7. May 2008. doi:10.1016/j.tcb.2008.03.001. PMID 18396046.
- ↑ Laurent V; Loisel TP; Harbeck B et al. (March 1999). "Role of Proteins of the Ena/VASP Family in Actin-based Motility of Listeria monocytogenes". The Journal of Cell Biology 144 (6): 1245–58. doi:10.1083/jcb.144.6.1245. PMID 10087267.
- ↑ "Pivotal role of VASP in Arp2/3 complex–mediated actin nucleation, actin branch-formation, and Listeria monocytogenes motility". The Journal of Cell Biology 155 (1): 89–100. October 2001. doi:10.1083/jcb.200106061. PMID 11581288.
- ↑ Bear J.E.; Svitkina T.M.; Krause M. et al. (May 2002). "Antagonism between Ena/VASP proteins and actin filament capping regulates fibroblast motility". Cell 109 (4): 509–21. doi:10.1016/S0092-8674(02)00731-6. PMID 12086607.
- ↑ Vazquez-Boland JA; Kocks C; Dramsi S et al. (January 1992). "Nucleotide sequence of the lecithinase operon of Listeria monocytogenes and possible role of lecithinase in cell-to-cell spread". Infection and Immunity 60 (1): 219–30. doi:10.1128/iai.60.1.219-230.1992. PMID 1309513.
- ↑ Domann E; Wehland J; Rohde M et al. (May 1992). "A novel bacterial virulence gene in Listeria monocytogenes required for host cell microfilament interaction with homology to the proline-rich region of vinculin". The EMBO Journal 11 (5): 1981–90. doi:10.1002/j.1460-2075.1992.tb05252.x. PMID 1582425.
- ↑ "Polarized distribution of Listeria monocytogenes surface protein ActA at the site of directional actin assembly". Journal of Cell Science 105 (3): 699–710. July 1993. doi:10.1242/jcs.105.3.699. PMID 8408297. http://jcs.biologists.org/cgi/pmidlookup?view=long&pmid=8408297.
- ↑ "The WASP and WAVE family proteins". Genome Biology 10 (6): 226. 2009. doi:10.1186/gb-2009-10-6-226. PMID 19589182.
External links
- YouTube video from Nature, Listeria monocytogenes [2:00–4:12]
 |