Biology:Edman degradation
Edman degradation, developed by Pehr Edman, is a method of sequencing amino acids in a peptide.[1] In this method, the amino-terminal residue is labeled and cleaved from the peptide without disrupting the peptide bonds between other amino acid residues.
Mechanism
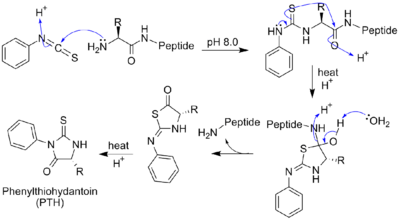
Phenyl isothiocyanate is reacted with an uncharged N-terminal amino group, under mildly alkaline conditions, to form a cyclical phenylthiocarbamoyl derivative. Then, under acidic conditions, this derivative of the terminal amino acid is cleaved as a thiazolinone derivative. The thiazolinone amino acid is then selectively extracted into an organic solvent and treated with acid to form the more stable phenylthiohydantoin (PTH)- amino acid derivative that can be identified by using chromatography or electrophoresis. This procedure can then be repeated again to identify the next amino acid. A major drawback to this technique is that the peptides being sequenced in this manner cannot have more than 50 to 60 residues (and in practice, under 30). The peptide length is limited due to the cyclical derivatization not always going to completion. The derivatization problem can be resolved by cleaving large peptides into smaller peptides before proceeding with the reaction. It is able to accurately sequence up to 30 amino acids with modern machines capable of over 99% efficiency per amino acid. An advantage of the Edman degradation is that it only uses 10 - 100 pico-moles of peptide for the sequencing process. The Edman degradation reaction was automated in 1967 by Edman and Beggs to speed up the process[2] and 100 automated devices were in use worldwide by 1973.[3]
Limitations
Because the Edman degradation proceeds from the N-terminus of the protein, it will not work if the N-terminus has been chemically modified (e.g. by acetylation or formation of pyroglutamic acid). Sequencing will stop if a non-α-amino acid is encountered (e.g. isoaspartic acid), since the favored five-membered ring intermediate is unable to be formed. Edman degradation is generally not useful to determine the positions of disulfide bridges. Protein sequencing of attomole level of Edman degraded sequences are obtainable but require accelerator mass spectrometery which requires large, complex, and expensive equipment as well as 30 hours of bench time to analyze a single run.[4]
Coupled analysis
Following 2D SDS PAGE the proteins can be transferred to a polyvinylidene difluoride (PVDF) blotting membrane for further analysis. Edman degradations can be performed directly from a PVDF membrane. N-terminal residue sequencing resulting in five to ten amino acid may be sufficient to identify a Protein of Interest (POI).[5]
See also
References
- ↑ Edman, P.; Högfeldt, Erik; Sillén, Lars Gunnar; Kinell, Per-Olof (1950). "Method for determination of the amino acid sequence in peptides". Acta Chem. Scand. 4: 283–293. doi:10.3891/acta.chem.scand.04-0283..
- ↑ Edman P, Begg G (March 1967). "A protein sequenator". Eur. J. Biochem. 1 (1): 80–91. doi:10.1111/j.1432-1033.1967.tb00047.x. PMID 6059350.
- ↑ Niall HD (1973). "Automated edman degradation: The protein sequenator". Part D: Enzyme Structure. Methods in Enzymology. 27. pp. 942–1010. doi:10.1016/S0076-6879(73)27039-8. ISBN 978-0-12-181890-6.
- ↑ Miyashita, Masahiro; Presley, Jack M.; Buchholz, Bruce A.; Lam, Kit S.; Lee, Young Moo; Vogel, John S.; Hammock, Bruce D. (April 10, 2001). "Attomole level protein sequencing by Edman degradation coupled with accelerator mass spectrometry". Proceedings of the National Academy of Sciences of the United States of America 98 (8): 4403–4408. doi:10.1073/pnas.071047998. PMID 11287636. Bibcode: 2001PNAS...98.4403M.
- ↑ Link, AJ; Robison, K; Church, GM (1997). "Comparing the predicted and observed properties of proteins encoded in the genome of Escherichia coli. Electrophoresis". Electrophoresis 18 (8): 1259-1313. PMID 9298646.
 |
