Biology:M13 bacteriophage
This article has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
(Learn how and when to remove this template message)
|
| Escherichia virus M13 | |
|---|---|
| |
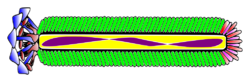
| Blue: Coat Protein pIII; Brown: Coat Protein pVI; Red: Coat Protein pVII; Limegreen: Coat Protein pVIII; Fuchsia: Coat Protein pIX; Purple: Single Stranded DNA | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Monodnaviria |
| Kingdom: | Loebvirae |
| Phylum: | Hofneiviricota |
| Class: | Faserviricetes |
| Order: | Tubulavirales |
| Family: | Inoviridae |
| Genus: | Inovirus |
| Species: | Escherichia virus M13
|
M13 is one of the Ff phages (fd and f1 are others), a member of the family filamentous bacteriophage (inovirus). Ff phages are composed of circular single-stranded DNA (ssDNA), which in the case of the m13 phage is 6407 nucleotides long and is encapsulated in approximately 2700 copies of the major coat protein p8, and capped with about 5 copies each of four different minor coat proteins (p3 and p6 at one end and p7 and p9 at the other end).[1][2][3] The minor coat protein p3 attaches to the receptor at the tip of the F pilus of the host Escherichia coli. The life cycle is relatively short, with the early phage progeny exiting the cell ten minutes after infection. Ff phages are chronic phage, releasing their progeny without killing the host cells. The infection causes turbid plaques in E. coli lawns, of intermediate opacity in comparison to regular lysis plaques. However, a decrease in the rate of cell growth is seen in the infected cells. The replicative form of M13 is circular double-stranded DNA similar to plasmids that are used for many recombinant DNA processes, and the virus has also been used for phage display, directed evolution, nanostructures and nanotechnology applications.[4][5][6]
Phage particles
The phage coat is primarily assembled from a 50 amino acid protein called p8, which is encoded by gene 8 in the phage genome. For a wild type M13 particle, it takes approximately 2700 copies of p8 to make the coat about 900 nm long. The coat's dimensions are flexible because the number of p8 copies adjusts to accommodate the size of the single stranded genome it packages.[7] The phage appear to be limited to approximately twice the natural DNA content. However, deletion of a phage protein (p3) prevents full escape from the host E. coli, and phage that are 10-20X the normal length with several copies of the phage genome can be seen shedding from the E. coli host.
At one end of the filament are up to five copies of the surface exposed protein (p9) and a more buried companion protein (p7). If p8 forms the shaft of the phage, p9 and p7 form the "blunt" end that is seen in micrographs. These proteins are very small, containing only 33 and 32 amino acids respectively, though some additional residues can be added to the N-terminal portion of each which are then presented on the outside of the coat. At the other end of the phage particle are five copies of the surface exposed (p3) and its less exposed accessory protein (p6). These form the rounded tip of the phage and are the first proteins to interact with the E. coli host during infection. Protein p3 is also the last point of contact with the host as a new phage buds from the bacterial surface.[8][9][10] The production of phage particles causes a host cell to grow and divide, but it does not lead to lysis of the cell.[8]
Replication in E. coli
Entry of the virus into a host cell is mediated by the p3 protein, specifically the N domains, binding to the primary and secondary receptors of the host cell.[11] After the positive single strand DNA has entered the cell, it is duplicated to form the double stranded DNA that is then used to transcribe the mRNA that will build the proteins.[8]
Below are steps involved with replication of M13 in E. coli.
- Viral (+) strand DNA enters cytoplasm
- Complementary (-) strand is synthesized by bacterial enzymes
- DNA Gyrase, a type II topoisomerase, acts on double-stranded DNA and catalyzes formation of negative supercoils in double-stranded DNA
- Final product is parental replicative form (RF) DNA
- Transcription and translation of the viral genome begins with p2.
- The phage protein, p2, nicks the (+) strand in the RF
- 3'-hydroxyl acts as a primer in the creation of new viral strand
- p2 circularizes displaced viral (+) strand DNA
- A pool of progeny double-stranded RF molecules is produced
- Negative strand of RF is template of transcription
- mRNAs are translated into the phage proteins
Phage proteins in the cytoplasm are p2, p10 and p5, and they are part of the replication process of DNA. The other phage proteins are synthesized and inserted into the cytoplasmic or outer membranes.
- p5 dimers bind newly synthesized single-stranded DNA and prevent conversion to RF DNA. The timing and attenuation of p5 translation is essential.
- RF DNA synthesis continues and amount of p5 reaches critical concentration
- DNA replication switches to synthesis of single-stranded (+) viral DNA
- p5-DNA structures from about 800 nm long and 8 nm in diameter
- p5-DNA complex is substrate in phage assembly reaction
Unusually, the major coat protein can insert post-translation into membranes, even those lacking translocation structures, and even into liposomes with no protein content.[12]
Research
George Smith, among others, showed that fragments of EcoRI endonuclease could be fused in the unique Bam site of f1 filamentous phage and thereby expressed in gene 3 whose protein p3 was externally accessible. M13 does not have this unique Bam site in gene 3. M13 had to be engineered to have accessible insertion sites, making it limited in its flexibility in handling different sized inserts. Because the M13 phage display system allows great flexibility in the location and number of recombinant proteins on the phage, it is a popular tool to construct or serve as a scaffold for nanostructures.[13] For example, the phage can be engineered to have a different protein on each end and along its length. This can be used to assemble structures like gold or cobalt oxide nano-wires for batteries[14] or to pack carbon nanotubes into straight bundles for use in photovoltaics.[15] The M13 capsid is also the first intact virus structure to ever be solved entirely by solid state NMR.[16]
See also
References
- ↑ "Simulation of the M13 life cycle I: Assembly of a genetically-structured deterministic chemical kinetic simulation". Virology 500: 259–274. January 2017. doi:10.1016/j.virol.2016.08.017. PMID 27644585.
- ↑ "Editorial: Filamentous Bacteriophage in Bio/Nano/Technology, Bacterial Pathogenesis and Ecology". Frontiers in Microbiology 7: 2109. 2016. doi:10.3389/fmicb.2016.02109. PMID 28066406.
- ↑ "Cryptic inoviruses revealed as pervasive in bacteria and archaea across Earth's biomes". Nature Microbiology 4 (11): 1895–1906. November 2019. doi:10.1038/s41564-019-0510-x. PMID 31332386.
- ↑ "Single M13 bacteriophage tethering and stretching". Proceedings of the National Academy of Sciences of the United States of America 104 (12): 4892–7. March 2007. doi:10.1073/pnas.0605727104. PMID 17360403.
- ↑ "M13 bacteriophage-polymer nanoassemblies as drug delivery vehicles.". Nano Research 4 (5): 483–93. May 2011. doi:10.1007/s12274-011-0104-2.
- ↑ "A system for the continuous directed evolution of biomolecules". Nature 472 (7344): 499–503. April 2011. doi:10.1038/nature09929. PMID 21478873. Bibcode: 2011Natur.472..499E.
- ↑ "Ff-nano, short functionalized nanorods derived from Ff (f1, fd, or M13) filamentous bacteriophage". Frontiers in Microbiology 6: 316. 2015. doi:10.3389/fmicb.2015.00316. PMID 25941520.
- ↑ 8.0 8.1 8.2 "Simulation of the M13 life cycle I: Assembly of a genetically-structured deterministic chemical kinetic simulation". Virology 500: 259–274. January 2017. doi:10.1016/j.virol.2016.08.017. PMID 27644585.
- ↑ Moon; Choi; Jeong; Sohn; Han; Oh (2019-10-11). "Research Progress of M13 Bacteriophage-Based Biosensors". Nanomaterials 9 (10): 1448. doi:10.3390/nano9101448. ISSN 2079-4991. PMID 31614669.
- ↑ Haase, Maximilian; Tessmer, Lutz; Köhnlechner, Lilian; Kuhn, Andreas (2022-05-27). "The M13 Phage Assembly Machine Has a Membrane-Spanning Oligomeric Ring Structure". Viruses 14 (6): 1163. doi:10.3390/v14061163. ISSN 1999-4915. PMID 35746635.
- ↑ Bennett, Nicholas J.; Gagic, Dragana; Sutherland-Smith, Andrew J.; Rakonjac, Jasna (2011). "Characterization of a Dual-Function Domain That Mediates Membrane Insertion and Excision of Ff Filamentous Bacteriophage". Journal of Molecular Biology 411 (5): 972–985. doi:10.1016/j.jmb.2011.07.002. ISSN 0022-2836. PMID 21763316. http://dx.doi.org/10.1016/j.jmb.2011.07.002.
- ↑ "Protein transport across the eukaryotic endoplasmic reticulum and bacterial inner membranes". Annual Review of Biochemistry (Annual Reviews) 65 (1): 271–303. 1996. doi:10.1146/annurev.bi.65.070196.001415. PMID 8811181.
- ↑ "Programmable assembly of nanoarchitectures using genetically engineered viruses". Nano Letters 5 (7): 1429–34. July 2005. doi:10.1021/nl050795d. PMID 16178252. Bibcode: 2005NanoL...5.1429H.
- ↑ "Virus-enabled synthesis and assembly of nanowires for lithium ion battery electrodes". Science 312 (5775): 885–8. May 2006. doi:10.1126/science.1122716. PMID 16601154. Bibcode: 2006Sci...312..885N.
- ↑ "Virus-templated self-assembled single-walled carbon nanotubes for highly efficient electron collection in photovoltaic devices". Nature Nanotechnology 6 (6): 377–84. April 2011. doi:10.1038/nnano.2011.50. PMID 21516089. Bibcode: 2011NatNa...6..377D.
- ↑ "The NMR–Rosetta capsid model of M13 bacteriophage reveals a quadrupled hydrophobic packing epitope". Proceedings of the National Academy of Sciences 112 (4): 971–976. January 2015. doi:10.1073/pnas.1415393112. PMID 25587134. Bibcode: 2015PNAS..112..971M.
Further reading
- Phage Display: A Laboratory Manual (1st ed.). Cold Spring Harbor Laboratory Press. October 2004. ISBN 978-0-87969-740-2.
- Griffin H.G., ed (1993). "M13 Cloning Vehicles". DNA Sequencing Protocols. Methods in Molecular Biology™. 23. Humana Press. pp. 9–22. doi:10.1385/0-89603-248-5:9. ISBN 0-89603-248-5. http://www.blogsua.com/pdf/dna-sequencing-protocols.pdf.
Wikidata ☰ Q625347 entry
 |
