Biology:Reference genome

A reference genome is a genome assembly that represents the complete genetic sequence of an organism as a continuous string of nucleotides (A, T, C, and G). For an assembly to serve as a reference genome, it is typically accompanied by annotations, produced through a process known as DNA or genome annotation. The annotations specify the genomic coordinates (start and end locations) of genes, exons, introns, and mRNA, and are often paired with corresponding transcript (mRNA) and protein sequences (algorithm predicted or experimentally validated).[1]
Reference genomes exist for a wide variety of species, including species of viruses, bacteria, fungi, plants and animals, and they differ in how they are constructed and represented. A reference may be derived from a single individual or from multiple individuals whose sequences are collapsed into one representative assembly - haplotype. Two main factors determine reference genome's assembly quality: the sequencing technology which affects sequence accuracy and the assembly level which indicates how complete the genome representation is.[2][3]
The ideal is a chromosome-level assembly, which is a complete DNA sequence for each chromosome with no unplaced segments. However, achieving this remains technically challenging, especially for large or repetitive genomes (dense in repetitive elements). Earlier sequencing technologies often produced assemblies at the contig (short contiguous sequences) or scaffold (ordered sets of contigs) level, with limited chromosomal context. The exact size of these fragments depends on the sequencing platform and bioinformatic methods available at the time.[4]
For assemblies that are not fully resolved, summary statistics such as N50 and L50 are commonly used to characterise contiguity and assembly fragmentation; these metrics are explained in the Contigs and Scaffolds section.
Reference genomes are central to omics research, particularly genomics. They provide a reference for "mapping" DNA sequence data from many individuals, enabling efficient identification of the genomic location of these sequences and the detection of polymorphisms (sequence differences among individuals) through a process known as variant calling.[5]
The limitations of this practice, such as reference bias and under-representation of population diversity, have led to the development of population-level reference sets and pangenomes.[6]
Reference genomes and their annotations are publicly accessible through online genome browsers and archives such as Ensembl,[7] the European Nucleotide Archive (ENA) at EMBL-EBI, the UCSC Genome Browser, and NCBI.
Properties of reference genomes
Measures of length
The length of a genome can be measured in multiple different ways.
A simple way to measure genome length is to count the number of base pairs in the assembly.[8]
The golden path is an alternative measure of length that omits redundant regions such as haplotypes and pseudo autosomal regions.[9][10] It is usually constructed by layering sequencing information over a physical map to combine scaffold information. It is a 'best estimate' of what the genome will look like and typically includes gaps, making it longer than the typical base pair assembly.[11]
Contigs and scaffolds

Reference genomes assembly requires reads overlapping, creating contigs, which are contiguous DNA regions of consensus sequences.[12] If there are gaps between contigs, these can be filled by scaffolding, either by contigs amplification with PCR and sequencing or by Bacterial Artificial Chromosome (BAC) cloning.[13][12] Filling these gaps is not always possible, in this case multiple scaffolds are created in a reference assembly.[14] Scaffolds are classified in 3 types: 1) Placed, whose chromosome, genomic coordinates and orientations are known; 2) Unlocalised, when only the chromosome is known but not the coordinates or orientation; 3) Unplaced, whose chromosome is not known.[15]
The number of contigs and scaffolds, as well as their average lengths are relevant parameters, among many others, for a reference genome assembly quality assessment since they provide information about the continuity of the final mapping from the original genome. The smaller the number of scaffolds per chromosome, until a single scaffold occupies an entire chromosome, the greater the continuity of the genome assembly.[16][17][18] Other related parameters are N50 and L50. N50 is the length of the contigs/scaffolds in which the 50% of the assembly is found in fragments of this length or greater, while L50 is the number of contigs/scaffolds whose length is N50. The higher the value of N50, the lower the value of L50, and vice versa, indicating high continuity in the assembly.[19][20][21]
Mammalian genomes
The human and mouse reference genomes are maintained and improved by the Genome Reference Consortium (GRC), a group of fewer than 20 scientists from a number of genome research institutes, including the European Bioinformatics Institute, the National Center for Biotechnology Information, the Sanger Institute and McDonnell Genome Institute at Washington University in St. Louis. GRC continues to improve reference genomes by building new alignments that contain fewer gaps, and fixing misrepresentations in the sequence.
Human reference genome
The original human reference genome was derived from thirteen anonymous volunteers from Buffalo, New York. Donors were recruited by advertisement in The Buffalo News, on Sunday, March 23, 1997. The first ten male and ten female volunteers were invited to make an appointment with the project's genetic counselors and donate blood from which DNA was extracted. As a result of how the DNA samples were processed, about 80 percent of the reference genome came from eight people and one male, designated RP11, accounts for 66 percent of the total. The ABO blood group system differs among humans, but the human reference genome contains only an O allele, although the others are annotated.[22][23][24][25][26]
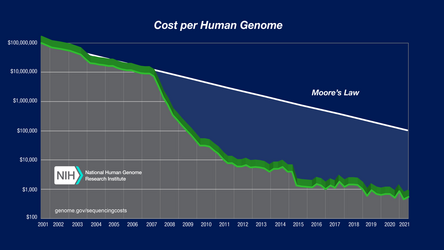
As the cost of DNA sequencing falls, and new full genome sequencing technologies emerge, more genome sequences continue to be generated. In several cases people such as James D. Watson had their genome assembled using massive parallel DNA sequencing.[27][28] Comparison between the reference (assembly NCBI36/hg18) and Watson's genome revealed 3.3 million single nucleotide polymorphism differences, while about 1.4 percent of his DNA could not be matched to the reference genome at all.[26][27] For regions where there is known to be large-scale variation, sets of alternate loci are assembled alongside the reference locus.

The latest human reference genome assembly, released by the Genome Reference Consortium, was GRCh38 in 2017.[30] Several patches were added to update it, the latest patch being GRCh38.p14, published on the 3rd of February 2022.[31][32] This build only has 349 gaps across the entire assembly, which implies a great improvement in comparison with the first version, which had roughly 150,000 gaps.[23] The gaps are mostly in areas such as telomeres, centromeres, and long repetitive sequences, with the biggest gap along the long arm of the Y chromosome, a region of ~30 Mb in length (~52% of the Y chromosome's length).[33] The number of genomic clone libraries contributing to the reference has increased steadily to >60 over the years, although individual RP11 still accounts for 70% of the reference genome.[34] Genomic analysis of this anonymous male suggests that he is of African-European ancestry.[34] According to the GRC website, their next assembly release for the human genome (version GRCh39) is currently "indefinitely postponed".[35]
In 2022, the Telomere-to-Telomere (T2T) Consortium,[36] an open, community-based effort, published the first completely assembled reference genome (version T2T-CHM13), without any gaps in the assembly. It did not contain a Y-chromosome until version 2.0.[37][38] This assembly allows for the examination of centromeric and pericentromeric sequence evolution. The consortium employed rigorous methods to assemble, clean, and validate complex repeat regions which are particularly difficult to sequence.[39] It used ultra-long–read (>100 kb) sequencing to accurately sequence segmental duplications.[40]
The T2T-CHM13 is sequenced from CHM13hTERT, a cell line from an essentially haploid hydatidiform mole. "CHM" stands for "Complete Hydatidiform Mole," and "13" is its line number. "hTERT" stands for "human Telomerase Reverse Transcriptase". The cell line has been transfected with the TERT gene, which is responsible for maintaining telomere length and thus contributes to the cell line's immortality.[41] A hydatidiform mole contains two copies of the same parental genome, and thus is essentially haploid. This eliminates allelic variation and allows better sequencing accuracy.[40]
Recent genome assemblies are as follows:[42]
| Release name | Date of release | Equivalent UCSC version |
|---|---|---|
| GRCh39 | Indefinitely postponed[35] | - |
| T2T-CHM13 | January 2022 | hs1 |
| GRCh38 | Dec 2013 | hg38 |
| GRCh37 | Feb 2009 | hg19 |
| NCBI Build 36.1 | Mar 2006 | hg18 |
| NCBI Build 35 | May 2004 | hg17 |
| NCBI Build 34 | Jul 2003 | hg16 |
Limitations
For much of a genome, the reference provides a good approximation of the DNA of any single individual. But in regions with high allelic diversity, such as the major histocompatibility complex in humans and the major urinary proteins of mice, the reference genome may differ significantly from other individuals.[43][44][45] Due to the fact that the reference genome is a "single" distinct sequence, which gives its utility as an index or locator of genomic features, there are limitations in terms of how faithfully it represents the human genome and its variability. Most of the initial samples used for reference genome sequencing came from people of European ancestry. In 2010, it was found that, by de novo assembling genomes from African and Asian populations with the NCBI reference genome (version NCBI36), these genomes had ~5Mb sequences that did not align against any region of the reference genome.[46]
Following projects to the Human Genome Project seek to address a deeper and more diverse characerization of the human genetic variability, which the reference genome is not able to represent. The HapMap Project, active during the period 2002 -2010, with the purpose of creating a haplotypes map and their most common variations among different human populations. Up to 11 populations of different ancestry were studied, such as individuals of the Han ethnic group from China, Gujaratis from India, the Yoruba people from Nigeria or Japanese people, among others.[47][48][49][50] The 1000 Genomes Project, carried out between 2008 and 2015, with the aim of creating a database that includes more than 95% of the variations present in the human genome and whose results can be used in studies of association with diseases (GWAS) such as diabetes, cardiovascular or autoimmune diseases. A total of 26 ethnic groups were studied in this project, expanding the scope of the HapMap project to new ethnic groups such as the Mende people of Sierra Leone, the Vietnamese people or the Bengali people.[51][52][53][54] The Human Pangenome Project, which started its initial phase in 2019 with the creation of the Human Pangenome Reference Consortium, seeks to create the largest map of human genetic variability taking the results of previous studies as a starting point.[55][56]
Mouse reference genome
Recent mouse genome assemblies are as follows:[42]
| Release name | Date of release | Equivalent UCSC version |
|---|---|---|
| GRCm39 | June 2020 | mm39 |
| GRCm38 | Dec 2011 | mm10 |
| NCBI Build 37 | Jul 2007 | mm9 |
| NCBI Build 36 | Feb 2006 | mm8 |
| NCBI Build 35 | Aug 2005 | mm7 |
| NCBI Build 34 | Mar 2005 | mm6 |
Other genomes
Since the Human Genome Project was finished, multiple international projects have started, focused on assembling reference genomes for many organisms. Model organisms (e.g., zebrafish (Danio rerio), chicken (Gallus gallus), Escherichia coli etc.) are of special interest to the scientific community, as well as, for example, endangered species (e.g., Asian arowana (Scleropages formosus) or the American bison (Bison bison)). As of August 2022, the NCBI database supports 71 886 partially or completely sequenced and assembled genomes from different species, such as 676 mammals, 590 birds and 865 fishes. Also noteworthy are the numbers of 1796 insects genomes, 3747 fungi, 1025 plants, 33 724 bacteria, 26 004 virus and 2040 archaea.[57] A lot of these species have annotation data associated with their reference genomes that can be publicly accessed and visualized in genome browsers such as Ensembl and UCSC Genome Browser.[58][59]
Some examples of these international projects are: the Chimpanzee Genome Project, carried out between 2005 and 2013 jointly by the Broad Institute and the McDonnell Genome Institute of Washington University in St. Louis, which generated the first reference genomes for 4 subspecies of Pan troglodytes;[60][61] the 100K Pathogen Genome Project, which started in 2012 with the main goal of creating a database of reference genomes for 100 000 pathogen microorganisms to use in public health, outbreaks detection, agriculture and environment;[62] the Earth BioGenome Project, which started in 2018 and aims to sequence and catalog the genomes of all the eukaryotic organisms on Earth to promote biodiversity conservation projects. Inside this big-science project there are up to 50 smaller-scale affiliated projects such as the Africa BioGenome Project or the 1000 Fungal Genomes Project.[63][64][65]
See also
References
- ↑ Ejigu, Girum Fitihamlak; Jung, Jaehee (2020-09-18). "Review on the Computational Genome Annotation of Sequences Obtained by Next-Generation Sequencing" (in en). Biology 9 (9): 295. doi:10.3390/biology9090295. ISSN 2079-7737. PMID 32962098.
- ↑ Giani, Alice Maria; Gallo, Guido Roberto; Gianfranceschi, Luca; Formenti, Giulio (2020-01-01). "Long walk to genomics: History and current approaches to genome sequencing and assembly" (in English). Computational and Structural Biotechnology Journal 18: 9–19. doi:10.1016/j.csbj.2019.11.002. ISSN 2001-0370. PMID 31890139. PMC 6926122. https://www.csbj.org/article/S2001-0370(19)30327-7/fulltext.
- ↑ Ballouz, Sara; Dobin, Alexander; Gillis, Jesse A. (2019-08-09). "Is it time to change the reference genome?". Genome Biology 20 (1): 159. doi:10.1186/s13059-019-1774-4. ISSN 1474-760X. PMID 31399121.
- ↑ Nurk, Sergey; Koren, Sergey; Rhie, Arang; Rautiainen, Mikko; Bzikadze, Andrey V.; Mikheenko, Alla; Vollger, Mitchell R.; Altemose, Nicolas et al. (April 2022). "The complete sequence of a human genome" (in en). Science 376 (6588): 44–53. doi:10.1126/science.abj6987. ISSN 0036-8075. PMID 35357919. Bibcode: 2022Sci...376...44N.
- ↑ Aganezov, Sergey; Yan, Stephanie M.; Soto, Daniela C.; Kirsche, Melanie; Zarate, Samantha; Avdeyev, Pavel; Taylor, Dylan J.; Shafin, Kishwar et al. (April 2022). "A complete reference genome improves analysis of human genetic variation" (in en). Science 376 (6588). doi:10.1126/science.abl3533. ISSN 0036-8075. PMID 35357935.
- ↑ Miga, Karen H.; Wang, Ting (2021-08-31). "The Need for a Human Pangenome Reference Sequence" (in en). Annual Review of Genomics and Human Genetics 22 (1): 81–102. doi:10.1146/annurev-genom-120120-081921. ISSN 1527-8204. PMID 33929893.
- ↑ Flicek, P.; Aken, B. L.; Beal, K.; Ballester, B.; Caccamo, M.; Chen, Y.; Clarke, L.; Coates, G. et al. (2007-12-23). "Ensembl 2008" (in en). Nucleic Acids Research 36 (Database): D707–D714. doi:10.1093/nar/gkm988. ISSN 0305-1048. PMID 18000006.
- ↑ "Help - Glossary - Homo sapiens - Ensembl genome browser 87". http://www.ensembl.org/Help/Glossary?id=230.
- ↑ "Golden path length | VectorBase". https://vectorbase.org/glossary/golden-path-length.
- ↑ "Help - Glossary - Homo sapiens - Ensembl genome browser 87". http://www.ensembl.org/Help/Glossary?id=229.
- ↑ "Whole assembly vs Golden path length in Ensembl? - SEQanswers". 31 July 2014. http://seqanswers.com/forums/showthread.php?t=45443.
- ↑ 12.0 12.1 Gibson, Greg; Muse, Spencer V. (2009). A Primer of Genome Science (3rd ed.). Sinauer Associates. p. 84. ISBN 978-0-878-93236-8.
- ↑ "Help - Glossary - Homo_sapiens - Ensembl genome browser 107". http://www.ensembl.org/Help/Glossary.
- ↑ Luo, Junwei; Wei, Yawei; Lyu, Mengna; Wu, Zhengjiang; Liu, Xiaoyan; Luo, Huimin; Yan, Chaokun (2021-09-02). "A comprehensive review of scaffolding methods in genome assembly". Briefings in Bioinformatics 22 (5). doi:10.1093/bib/bbab033. ISSN 1477-4054. PMID 33634311.
- ↑ "Chromosomes, scaffolds and contigs". http://www.ensembl.org/info/genome/genebuild/chromosomes_scaffolds_contigs.html.
- ↑ Meader, Stephen; Hillier, LaDeana W.; Locke, Devin; Ponting, Chris P.; Lunter, Gerton (May 2010). "Genome assembly quality: Assessment and improvement using the neutral indel model". Genome Research 20 (5): 675–684. doi:10.1101/gr.096966.109. ISSN 1088-9051. PMID 20305016.
- ↑ Rice, Edward S.; Green, Richard E. (2019-02-15). "New Approaches for Genome Assembly and Scaffolding" (in en). Annual Review of Animal Biosciences 7 (1): 17–40. doi:10.1146/annurev-animal-020518-115344. ISSN 2165-8102. PMID 30485757. https://www.annualreviews.org/doi/10.1146/annurev-animal-020518-115344.
- ↑ Cao, Minh Duc; Nguyen, Son Hoang; Ganesamoorthy, Devika; Elliott, Alysha G.; Cooper, Matthew A.; Coin, Lachlan J. M. (2017-02-20). "Scaffolding and completing genome assemblies in real-time with nanopore sequencing" (in en). Nature Communications 8 (1). doi:10.1038/ncomms14515. ISSN 2041-1723. PMID 28218240. Bibcode: 2017NatCo...814515C.
- ↑ Mende, Daniel R.; Waller, Alison S.; Sunagawa, Shinichi; Järvelin, Aino I.; Chan, Michelle M.; Arumugam, Manimozhiyan; Raes, Jeroen; Bork, Peer (2012-02-23). "Assessment of Metagenomic Assembly Using Simulated Next Generation Sequencing Data". PLOS ONE 7 (2). doi:10.1371/journal.pone.0031386. ISSN 1932-6203. PMID 22384016. Bibcode: 2012PLoSO...731386M.
- ↑ Alhakami, Hind; Mirebrahim, Hamid; Lonardi, Stefano (2017-05-18). "A comparative evaluation of genome assembly reconciliation tools". Genome Biology 18 (1): 93. doi:10.1186/s13059-017-1213-3. ISSN 1474-7596. PMID 28521789.
- ↑ Castro, Christina J.; Ng, Terry Fei Fan (2017-11-01). "U50: A New Metric for Measuring Assembly Output Based on Non-Overlapping, Target-Specific Contigs". Journal of Computational Biology 24 (11): 1071–1080. doi:10.1089/cmb.2017.0013. PMID 28418726.
- ↑ A short guide to the human genome. CSHL Press. 2008. p. 135. ISBN 978-0-87969-791-4.
- ↑ 23.0 23.1 "E pluribus unum". Nature Methods 7 (5): 331. May 2010. doi:10.1038/nmeth0510-331. PMID 20440876.
- ↑ "Is it time to change the reference genome?". Genome Biology 20 (1). August 2019. doi:10.1186/s13059-019-1774-4. PMID 31399121.
- ↑ "Limitations of the human reference genome for personalized genomics". PLOS ONE 7 (7). 11 July 2012. doi:10.1371/journal.pone.0040294. PMID 22811759. Bibcode: 2012PLoSO...740294R.
- ↑ 26.0 26.1 "Genome of DNA Pioneer Is Deciphered". New York Times. May 31, 2007. https://www.nytimes.com/2007/05/31/science/31cnd-gene.html.
- ↑ 27.0 27.1 "The complete genome of an individual by massively parallel DNA sequencing". Nature 452 (7189): 872–876. April 2008. doi:10.1038/nature06884. PMID 18421352. Bibcode: 2008Natur.452..872W.
- ↑ The exception to this is J. Craig Venter whose DNA was sequenced and assembled using shotgun sequencing methods.
- ↑ "Genome Data Viewer - NCBI". https://www.ncbi.nlm.nih.gov/gdv/browser/genome/?id=GCF_000001405.40.
- ↑ "Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly". Genome Research 27 (5): 849–864. May 2017. doi:10.1101/gr.213611.116. PMID 28396521.
- ↑ "GRCh38.p14 - hg38 - Genome - Assembly - NCBI". https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.40/.
- ↑ Genome Reference Consortium (2022-05-09). "GenomeRef: GRCh38.p14 is now released!". https://genomeref.blogspot.com/2022/05/grch38p14-is-now-released.html.
- ↑ "GRCh38.p14 - hg38 - Genome - Assembly - NCBI - Statistics Report". https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.40/.
- ↑ 34.0 34.1 "How many individuals were sequenced for the human reference genome assembly?". https://www.ncbi.nlm.nih.gov/grc/help/faq/#human-reference-genome-individuals.
- ↑ 35.0 35.1 "Genome Reference Consortium". https://www.ncbi.nlm.nih.gov/grc.
- ↑ "Telomere-to-Telomere" (in en). https://www.genome.gov/about-genomics/telomere-to-telomere.
- ↑ "The complete sequence of a human genome". Science 376 (6588): 44–53. April 2022. doi:10.1126/science.abj6987. PMID 35357919. Bibcode: 2022Sci...376...44N.
- ↑ "T2T-CHM13v2.0 - Genome - Assembly - NCBI". https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_009914755.1/.
- ↑ Altemose, Nicolas; Logsdon, Glennis A.; Bzikadze, Andrey V.; Sidhwani, Pragya; Langley, Sasha A.; Caldas, Gina V.; Hoyt, Savannah J.; Uralsky, Lev et al. (April 2022). "Complete genomic and epigenetic maps of human centromeres" (in en). Science 376 (6588). doi:10.1126/science.abl4178. ISSN 0036-8075. PMID 35357911.
- ↑ 40.0 40.1 Church, Deanna M. (April 2022). "A next-generation human genome sequence" (in en). Science 376 (6588): 34–35. doi:10.1126/science.abo5367. ISSN 0036-8075. PMID 35357937. Bibcode: 2022Sci...376...34C. https://www.science.org/doi/10.1126/science.abo5367.
- ↑ Steinberg, Karyn Meltz; Schneider, Valerie A.; Graves-Lindsay, Tina A.; Fulton, Robert S.; Agarwala, Richa; Huddleston, John; Shiryev, Sergey A.; Morgulis, Aleksandr et al. (December 2014). "Single haplotype assembly of the human genome from a hydatidiform mole". Genome Research 24 (12): 2066–2076. doi:10.1101/gr.180893.114. ISSN 1088-9051. PMID 25373144.
- ↑ 42.0 42.1 "UCSC Genome Bioinformatics: FAQ". https://genome.ucsc.edu/FAQ/FAQreleases.html#release1.
- ↑ MHC Sequencing Consortium (October 1999). "Complete sequence and gene map of a human major histocompatibility complex. The MHC sequencing consortium". Nature 401 (6756): 921–923. doi:10.1038/44853. PMID 10553908. Bibcode: 1999Natur.401..921T.
- ↑ "Species specificity in major urinary proteins by parallel evolution". PLOS ONE 3 (9). September 2008. doi:10.1371/journal.pone.0003280. PMID 18815613. Bibcode: 2008PLoSO...3.3280L.
- ↑ Urinary Lipocalins in Rodenta:is there a Generic Model?. Chemical Signals in Vertebrates 11. Springer New York. October 2007. ISBN 978-0-387-73944-1.
- ↑ "Building the sequence map of the human pan-genome". Nature Biotechnology 28 (1): 57–63. January 2010. doi:10.1038/nbt.1596. PMID 19997067.
- ↑ The International HapMap Consortium (October 2005). "A haplotype map of the human genome". Nature 437 (7063): 1299–1320. doi:10.1038/nature04226. PMID 16255080. Bibcode: 2005Natur.437.1299T.
- ↑ "A second generation human haplotype map of over 3.1 million SNPs". Nature 449 (7164): 851–861. October 2007. doi:10.1038/nature06258. PMID 17943122. Bibcode: 2007Natur.449..851F.
- ↑ "Integrating common and rare genetic variation in diverse human populations". Nature 467 (7311): 52–58. September 2010. doi:10.1038/nature09298. PMID 20811451. Bibcode: 2010Natur.467...52T.
- ↑ "International HapMap Project" (in en). https://www.genome.gov/10001688/international-hapmap-project.
- ↑ "A map of human genome variation from population-scale sequencing". Nature 467 (7319): 1061–1073. October 2010. doi:10.1038/nature09534. PMID 20981092. Bibcode: 2010Natur.467.1061T.
- ↑ "An integrated map of genetic variation from 1,092 human genomes". Nature 491 (7422): 56–65. November 2012. doi:10.1038/nature11632. PMID 23128226. Bibcode: 2012Natur.491...56T.
- ↑ "A global reference for human genetic variation". Nature 526 (7571): 68–74. October 2015. doi:10.1038/nature15393. PMID 26432245. Bibcode: 2015Natur.526...68T.
- ↑ "An integrated map of structural variation in 2,504 human genomes". Nature 526 (7571): 75–81. October 2015. doi:10.1038/nature15394. PMID 26432246. Bibcode: 2015Natur.526...75..
- ↑ "The Need for a Human Pangenome Reference Sequence". Annual Review of Genomics and Human Genetics 22 (1): 81–102. August 2021. doi:10.1146/annurev-genom-120120-081921. PMID 33929893.
- ↑ "The Human Pangenome Project: a global resource to map genomic diversity". Nature 604 (7906): 437–446. April 2022. doi:10.1038/s41586-022-04601-8. PMID 35444317. Bibcode: 2022Natur.604..437W.
- ↑ "Genome List - Genome - NCBI". https://www.ncbi.nlm.nih.gov/genome/browse#!/overview/.
- ↑ "Species List". https://uswest.ensembl.org/info/about/species.html.
- ↑ "GenArk: UCSC Genome Archive". https://hgdownload.soe.ucsc.edu/hubs/.
- ↑ "Chimpanzee Genome Project" (in en). BCM-HGSC. 2016-03-04. https://www.hgsc.bcm.edu/non-human-primates/chimpanzee-genome-project.
- ↑ "Great ape genetic diversity and population history". Nature 499 (7459): 471–475. July 2013. doi:10.1038/nature12228. PMID 23823723. Bibcode: 2013Natur.499..471P.
- ↑ "100K Pathogen Genome Project – Genomes for Public Health & Food Safety" (in en-US). https://100kgenomes.org/.
- ↑ "Earth BioGenome Project: Sequencing life for the future of life". Proceedings of the National Academy of Sciences of the United States of America 115 (17): 4325–4333. April 2018. doi:10.1073/pnas.1720115115. PMID 29686065. Bibcode: 2018PNAS..115.4325L.
- ↑ "African BioGenome Project – Genomics in the service of conservation and improvement of African biological diversity" (in en-US). https://africanbiogenome.org/.
- ↑ "1000 Fungal Genomes Project". https://mycocosm.jgi.doe.gov/mycocosm/home/1000-fungal-genomes.
External links
 |
