Biology:Archaea
Archaea (/ɑːrˈkiːə/ (![]() listen) ar-KEE-ə) is a domain of organisms. Traditionally, Archaea included only its prokaryotic members, but has since been found to be paraphyletic, as eukaryotes are known to have evolved from archaea. Even though the domain Archaea cladistically includes eukaryotes, the term archaea (Template:Singular abbr /ɑːrˈkiːɒn/ ar-KEE-on; from grc ἀρχαῖον arkhaîon 'ancient') in English still generally refers specifically to prokaryotic members of Archaea.
listen) ar-KEE-ə) is a domain of organisms. Traditionally, Archaea included only its prokaryotic members, but has since been found to be paraphyletic, as eukaryotes are known to have evolved from archaea. Even though the domain Archaea cladistically includes eukaryotes, the term archaea (Template:Singular abbr /ɑːrˈkiːɒn/ ar-KEE-on; from grc ἀρχαῖον arkhaîon 'ancient') in English still generally refers specifically to prokaryotic members of Archaea.
Archaea were initially classified as bacteria, receiving the name archaebacteria (/ˌɑːrkibækˈtɪəriə/, in the Archaebacteria kingdom), but this term has fallen out of use.[1] Archaeal cells have unique properties distinguishing them from Bacteria and Eukaryota, including: cell membranes made of ether-linked lipids; metabolisms such as methanogenesis; and a unique motility structure known as an archaellum.[2] Archaea are further divided into multiple recognized phyla. Classification is difficult because most have not been isolated in a laboratory and have been identified only by their gene sequences in environmental samples. It is unknown whether they can produce endospores.
Archaea are often similar to bacteria in size and shape, although a few have very different shapes, such as the flat, square cells of Haloquadratum walsbyi.[3] Despite this, archaea possess genes and several metabolic pathways that are more closely related to those of eukaryotes, notably for the enzymes involved in transcription and translation. Other aspects of archaeal biochemistry are unique, such as their reliance on ether lipids in their cell membranes,[4] including archaeols. Archaea use more diverse energy sources than eukaryotes, ranging from organic compounds such as sugars, to ammonia, metal ions or even hydrogen gas. The salt-tolerant Halobacteria use sunlight as an energy source, and other species of archaea fix carbon (autotrophy), but unlike cyanobacteria, no known species of archaea does both. Archaea reproduce asexually by binary fission, fragmentation, or budding; unlike bacteria, no known species of Archaea form endospores. The first observed archaea were extremophiles, living in extreme environments such as hot springs and salt lakes with no other organisms. Improved molecular detection tools led to the discovery of archaea in almost every habitat, including soil,[5] oceans, and marshlands. Archaea are particularly numerous in the oceans, and the archaea in plankton may be one of the most abundant groups of organisms on the planet.
Archaea are a major part of Earth's life. They are part of the microbiota of all organisms. In the human microbiome, they are important in the gut, mouth, and on the skin.[6] Their morphological, metabolic, and geographical diversity permits them to play multiple ecological roles: carbon fixation; nitrogen cycling; organic compound turnover; and maintaining microbial symbiotic and syntrophic communities, for example.[5][7] As of 2024, only one species of non-eukaryotic archaea has been found to be parasitic;[8] many are mutualists or commensals, such as the methanogens (methane-producers) that inhabit the gastrointestinal tract in humans and ruminants, where their vast numbers facilitate digestion. Methanogens are used in biogas production and sewage treatment, while biotechnology exploits enzymes from extremophile archaea that can endure high temperatures and organic solvents.
Discovery and classification
Early concept

For much of the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their biochemistry, morphology and metabolism. Microbiologists tried to classify microorganisms based on the structures of their cell walls, their shapes, and the substances they consume.[9] In 1965, Emile Zuckerkandl and Linus Pauling[10] instead proposed using the sequences of the genes in different prokaryotes to work out how they are related to each other. This phylogenetic approach is the main method used today.[11]
Archaea were first classified separately from bacteria in 1977 by Carl Woese and George E. Fox, based on their ribosomal RNA (rRNA) genes.[12] (At that time only the methanogens were known). They called these groups the Urkingdoms of Archaebacteria and Eubacteria, though other researchers treated them as kingdoms or subkingdoms. Woese and Fox gave the first evidence for Archaebacteria as a separate "line of descent": 1. lack of peptidoglycan in their cell walls, 2. two unusual coenzymes, 3. results of 16S ribosomal RNA gene sequencing. To emphasize this difference, Woese, Otto Kandler and Mark Wheelis later proposed reclassifying organisms into three then thought to be natural domains known as the three-domain system: the Eukarya, the Bacteria and the Archaea,[13] in what is now known as the Woesian Revolution.[14]
The word archaea comes from the Ancient Greek ἀρχαῖα, meaning "ancient things",[15] as the first representatives of the domain Archaea were methanogens and it was assumed that their metabolism reflected Earth's primitive atmosphere and the organisms' antiquity, but as new habitats were studied, more organisms were discovered. Extreme halophilic[16] and hyperthermophilic microbes[17] were also included in Archaea. For a long time, archaea were seen as extremophiles that exist only in extreme habitats such as hot springs and salt lakes, but by the end of the 20th century, archaea had been identified in non-extreme environments as well. Today, they are known to be a large and diverse group of organisms abundantly distributed throughout nature.[18] This new appreciation of the importance and ubiquity of archaea came from using polymerase chain reaction (PCR) to detect prokaryotes from environmental samples (such as water or soil) by multiplying their ribosomal genes. This allows the detection and identification of organisms that have not been cultured in the laboratory.[19][20]
Classification

The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors.[21] These classifications rely heavily on the use of the sequence of ribosomal RNA genes to reveal relationships among organisms (molecular phylogenetics).[22] Most of the culturable and well-investigated species of archaea are members of two main kingdoms, the Methanobacteriati and the Thermoproteati (formerly TACK). Other groups have been tentatively created, such as the peculiar species Nanoarchaeum equitans – discovered in 2003 and assigned its own phylum, the Nanoarchaeota (reassigned to Nanobdellota in 2023[23]).[24] A new phylum "Korarchaeota" (now Thermoproteota)[25] has also been proposed, containing a small group of unusual thermophilic species sharing features of both the main phyla.[26][27] Other detected species of archaea are only distantly related to any of these groups, such as the Archaeal Richmond Mine acidophilic nanoorganisms (ARMAN, comprising Micrarchaeota and Parvarchaeota), which were discovered in 2006[28] and are some of the smallest organisms known.[29]
A superphylum – "TACK" (now kingdom Thermoproteati) – which includes the Thaumarchaeota (now Nitrososphaerota), "Augarchaeota", Crenarchaeota (now Thermoproteota), and "Korarchaeota" (now Thermoproteota)[25] was proposed in 2011 to be related to the origin of eukaryotes.[30] In 2017, the newly discovered and newly named "Asgard" (now kingdom Promethearchaeati) superphylum was proposed to be more closely related to the original eukaryote and a sister group to Thermoproteati / "TACK".[31]
In 2013, the superphylum "DPANN" (now kingdom Nanobdellati) was proposed to group "Nanoarchaeota", "Nanohaloarchaeota", Archaeal Richmond Mine acidophilic nanoorganisms (ARMAN, comprising "Micrarchaeota" and "Parvarchaeota"), and other similar archaea. This archaeal superphylum encompasses at least 10 different lineages and includes organisms with extremely small cell and genome sizes and limited metabolic capabilities. Therefore, Nanobdellati/"DPANN" may include members obligately dependent on symbiotic interactions, and may even include novel parasites. However, other phylogenetic analyses found that Nanobdellati/"DPANN" does not form a monophyletic group, and that the apparent grouping is caused by long branch attraction (LBA), suggesting that all these lineages belong to Methanobacteriati.[32][33]
Phylogeny
According to Tom A. Williams et al. 2017,[34] Castelle & Banfield (2018)[35] and GTDB release 10-RS226 (16th April 2025).[36][37][38]
| Tom A. Williams et al. 2017[34] and Castelle & Banfield 2018[35] | 10-RS226 (16th April 2025)[36][37][38] | ||
|---|---|---|---|
|
Lua error: Internal error: The interpreter exited with status 1. |
|
Concept of species
The classification of archaea into species is also controversial. Ernst Mayr's species definition – a reproductively isolated group of interbreeding organisms - does not apply, as archaea reproduce only asexually.[39]
Archaea show high levels of horizontal gene transfer between lineages. Some researchers suggest that individuals can be grouped into species-like populations given highly similar genomes and infrequent gene transfer to/from cells with less-related genomes, as in the genus Ferroplasma.[40] On the other hand, studies in Halorubrum found significant genetic transfer to/from less-related populations, limiting the criterion's applicability.[41] Some researchers question whether such species designations have practical meaning.[42]
Current knowledge on genetic diversity in archaeans is fragmentary, so the total number of species cannot be estimated with any accuracy.[22] Estimates of the number of phyla range from 18 to 23, of which only 8 have representatives that have been cultured and studied directly. Many of these hypothesized groups are known from a single rRNA sequence, so the level of diversity remains obscure.[43] This situation is also seen in the Bacteria; many uncultured microbes present similar issues with characterization.[44]
Prokaryotic phyla
Valid phyla
The following phyla have been validly published according to the Prokaryotic Code; belonging to the four kingdoms of archaea:[45][46][25]
- Methanobacteriota
- Microcaldota
- Nanobdellota
- Promethearchaeota
- Thermoproteota
Candidate phyla
The following phyla have been proposed, but have not been validly published according to the Prokaryotic Code; phyla that do not belong to any kingdom are shown in bold:[47][25]
- "Aenigmatarchaeota"
- "Altarchaeota"
- "Augarchaeota"
- "Geoarchaeota"
- "Hadarchaeota"
- "Hadesarchaeota"
- "Huberarchaeota"
- "Hydrothermarchaeota"
- "Iainarchaeota"
- "Micrarchaeota"
- "Nanohalarchaeota"
- "Nezhaarchaeota"
- "Parvarchaeota"
- "Poseidoniota"
- "Undinarchaeota"
Origin and evolution
The age of the Earth is about 4.54 billion years.[48][49][50] Scientific evidence suggests that life began on Earth at least 3.5 billion years ago.[51][52] The earliest evidence for life on Earth is graphite found to be biogenic in 3.7-billion-year-old metasedimentary rocks discovered in Western Greenland[53] and microbial mat fossils found in 3.48-billion-year-old sandstone discovered in Western Australia.[54][55] In 2015, possible remains of biotic matter were found in 4.1-billion-year-old rocks in Western Australia.[56][57]
Although probable prokaryotic cell fossils date to almost 3.5 billion years ago, most prokaryotes do not have distinctive morphologies, and fossil shapes cannot be used to identify them as archaea.[58] Instead, chemical fossils of unique lipids are more informative because such compounds do not occur in other organisms.[59] Some publications suggest that archaeal or eukaryotic lipid remains are present in shales dating from 2.7 billion years ago,[60] though such data have since been questioned.[61] These lipids have also been detected in even older rocks from west Greenland. The oldest such traces come from the Isua district, which includes Earth's oldest known sediments, formed 3.8 billion years ago.[62] The archaeal lineage may be the most ancient that exists on Earth.[63]
Woese argued that the bacteria, archaea, and eukaryotes represent separate lines of descent that diverged early on from an ancestral colony of organisms.[64][65] One possibility[65][66] is that this occurred before the evolution of cells, when the lack of a typical cell membrane allowed unrestricted lateral gene transfer, and that the common ancestors of the three domains arose by fixation of specific subsets of genes.[65][66] It is possible that the last common ancestor of bacteria and archaea was a thermophile, which raises the possibility that lower temperatures are "extreme environments" for archaea, and organisms that live in cooler environments appeared only later.[67] Since archaea and bacteria are no more related to each other than they are to eukaryotes, the term prokaryote may suggest a false similarity between them.[68] However, structural and functional similarities between lineages often occur because of shared ancestral traits or evolutionary convergence. These similarities are known as a grade, and prokaryotes are best thought of as a grade of life, characterized by such features as an absence of membrane-bound organelles.
Comparison with other domains
The following table compares some major characteristics of the three domains, to illustrate their similarities and differences.[69]
| Property | Archaea | Bacteria | Eukaryota |
|---|---|---|---|
| Cell membrane | Ether-linked lipids | Ester-linked lipids | Ester-linked lipids |
| Cell wall | Glycoprotein, or S-layer; rarely pseudopeptidoglycan | Peptidoglycan, S-layer, or no cell wall | Various structures |
| Gene structure | Circular chromosomes, similar translation and transcription to Eukaryota | Circular chromosomes, unique translation and transcription | Multiple, linear chromosomes, but translation and transcription similar to Archaea |
| Internal cell structure | No membrane-bound organelles (?[70]) or nucleus | No membrane-bound organelles or nucleus | Membrane-bound organelles and nucleus |
| Metabolism[71] | Various, including diazotrophy, with methanogenesis unique to Archaea | Various, including photosynthesis, aerobic and anaerobic respiration, fermentation, diazotrophy, and autotrophy | Photosynthesis, cellular respiration, and fermentation; no diazotrophy |
| Reproduction | Asexual reproduction, horizontal gene transfer | Asexual reproduction, horizontal gene transfer | Sexual and asexual reproduction |
| Protein synthesis initiation | Methionine | Formylmethionine | Methionine |
| RNA polymerase | One | One | Many |
| EF-2/EF-G | Sensitive to diphtheria toxin | Resistant to diphtheria toxin | Sensitive to diphtheria toxin |
Archaea were split off as a third domain because of the large differences in their ribosomal RNA structure. The particular molecule 16S rRNA is key to the production of proteins in all organisms. Because this function is so central to life, organisms with mutations in their 16S rRNA are unlikely to survive, leading to great (but not absolute) stability in the structure of this polynucleotide over generations. 16S rRNA is large enough to show organism-specific variations, but still small enough to be compared quickly. In 1977, Carl Woese, a microbiologist studying the genetic sequences of organisms, developed a new comparison method that involved splitting the RNA into fragments that could be sorted and compared with other fragments from other organisms.[12] The more similar the patterns between species, the more closely they are related.[72]
Woese used his new rRNA comparison method to categorize and contrast different organisms. He compared a variety of species and happened upon a group of methanogens with rRNA vastly different from any known prokaryotes or eukaryotes.[12] These methanogens were much more similar to each other than to other organisms, leading Woese to propose the new domain of Archaea.[12] His experiments showed that the archaea were genetically more similar to eukaryotes than prokaryotes, even though they were more similar to prokaryotes in structure.[73] This led to the conclusion that Archaea and Eukarya shared a common ancestor more recent than Eukarya and Bacteria.[73] The development of the nucleus occurred after the split between Bacteria and this common ancestor.[73][13]
One property unique to archaea is the abundant use of ether-linked lipids in their cell membranes. Ether linkages are more chemically stable than the ester linkages found in bacteria and eukarya, which may be a contributing factor to the ability of many archaea to survive in extreme environments that place heavy stress on cell membranes, such as extreme heat and salinity. Comparative analysis of archaeal genomes has also identified several molecular conserved signature indels and signature proteins uniquely present in either all archaea or different main groups within archaea.[74][75][76] Another unique feature of archaea, found in no other organisms, is methanogenesis (the metabolic production of methane). Methanogenic archaea play a pivotal role in ecosystems with organisms that derive energy from oxidation of methane, many of which are bacteria, as they are often a major source of methane in such environments and can play a role as primary producers. Methanogens also play a critical role in the carbon cycle, breaking down organic carbon into methane, which is also a major greenhouse gas.[77]
This difference in the biochemical structure of Bacteria and Archaea has been explained by researchers through evolutionary processes. It is theorized that both domains originated at deep sea alkaline hydrothermal vents. At least twice, microbes evolved lipid biosynthesis and cell wall biochemistry. It has been suggested that the last universal common ancestor was a non-free-living organism.[78] It may have had a permeable membrane composed of bacterial simple chain amphiphiles (fatty acids), including archaeal simple chain amphiphiles (isoprenoids). These stabilize fatty acid membranes in seawater; this property may have driven the divergence of bacterial and archaeal membranes, "with the later biosynthesis of phospholipids giving rise to the unique G1P and G3P headgroups of archaea and bacteria respectively. If so, the properties conferred by membrane isoprenoids place the lipid divide as early as the origin of life".[79]
Relationship to bacteria
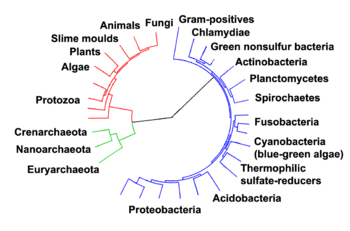
The relationships among the three domains are of central importance for understanding the origin of life. Most of the metabolic pathways, which are the object of the majority of an organism's genes, are common between Archaea and Bacteria, while most genes involved in gene expression are common between Archaea and Eukarya.[81] Within prokaryotes, archaeal cell structure is most similar to that of Gram-positive bacteria, largely because both have a single lipid bilayer[82] and usually contain a thick sacculus (exoskeleton) of varying chemical composition.[83] In some phylogenetic trees based upon different gene / protein sequences of prokaryotic homologs, the archaeal homologs are more closely related to those of gram-positive bacteria.[82] Archaea and gram-positive bacteria also share conserved indels in a number of important proteins, such as Hsp70 and glutamine synthetase I;[82][84] but the phylogeny of these genes was interpreted to reveal inter-domain gene transfer,[85][86] and might not reflect the organismal relationship(s).[87]
It has been proposed that the archaea evolved from Gram-positive bacteria in response to antibiotic selection pressure.[82][84][88] This is suggested by the observation that archaea are resistant to a wide variety of antibiotics that are produced primarily by Gram-positive bacteria,[82][84] and that these antibiotics act primarily on the genes that distinguish archaea from bacteria. The proposal is that the selective pressure towards resistance generated by the gram-positive antibiotics was eventually sufficient to cause extensive changes in many of the antibiotics' target genes, and that these strains represented the common ancestors of present-day Archaea.[88] The evolution of Archaea in response to antibiotic selection, or any other competitive selective pressure, could also explain their adaptation to extreme environments (such as high temperature or acidity) as the result of a search for unoccupied niches to escape from antibiotic-producing organisms;[88][89] Cavalier-Smith has made a similar suggestion, the Neomura hypothesis.[90] This proposal is also supported by other work investigating protein structural relationships[91] and studies that suggest that gram-positive bacteria may constitute the earliest branching lineages within the prokaryotes.[92]
Relation to eukaryotes
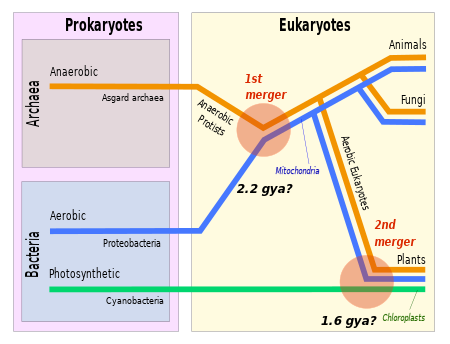
The evolutionary relationship between archaea and eukaryotes remains unclear. Aside from the similarities in cell structure and function that are discussed below, many genetic trees group the two.[94]
Complicating factors include claims that the relationship between eukaryotes and the archaeal phylum Thermoproteota is closer than the relationship between the Methanobacteriati and the phylum Thermoproteota[95] and the presence of archaea-like genes in certain bacteria, such as Thermotoga maritima, from horizontal gene transfer.[96] The standard hypothesis states that the ancestor of the eukaryotes diverged early from the Archaea,[97][98] and that eukaryotes arose through symbiogenesis, the fusion of an archaean and a eubacterium, which formed the mitochondria; this hypothesis explains the genetic similarities between the groups.[93] The eocyte hypothesis instead posits that Eukaryota emerged relatively late from the Archaea.[99]
A lineage of archaea discovered in 2015, Lokiarchaeum (of the proposed new phylum "Lokiarchaeota"), named for a hydrothermal vent called Loki's Castle in the Arctic Ocean, was found to be the most closely related to eukaryotes known at that time. It has been called a transitional organism between prokaryotes and eukaryotes.[100][101]
Several sister phyla of "Lokiarchaeota" have since been found ("Thorarchaeota", "Odinarchaeota", "Heimdallarchaeota"), all together comprising a newly proposed supergroup "Asgard".[31][102][103]
Details of the relation of Promethearchaeati / "Asgard" members and eukaryotes are still under consideration,[104] although, in January 2020, scientists reported that Promethearchaeum syntrophicum, a type of Promethearchaeati / "Asgard" archaea, may be a possible link between simple prokaryotic and complex eukaryotic microorganisms about two billion years ago.[105][106][107]
Morphology
Individual archaea range from 0.1 micrometers (μm) to over 15 μm in diameter, and occur in various shapes, commonly as spheres, rods, spirals or plates.[108] Other morphologies in the Thermoproteota include irregularly shaped lobed cells in Sulfolobus, needle-like filaments that are less than half a micrometer in diameter in Thermofilum, and almost perfectly rectangular rods in Thermoproteus and Pyrobaculum.[109] Archaea in the genus Haloquadratum such as Haloquadratum walsbyi are flat, square specimens that live in hypersaline pools.[110] These unusual shapes are probably maintained by both their cell walls and a prokaryotic cytoskeleton. Proteins related to the cytoskeleton components of other organisms exist in archaea,[111] and filaments form within their cells,[112] but in contrast with other organisms, these cellular structures are poorly understood.[113] In Thermoplasma and Ferroplasma the lack of a cell wall means that the cells have irregular shapes, and can resemble amoebae.[114]
Some species form aggregates or filaments of cells up to 200 μm long.[108] These organisms can be prominent in biofilms.[115] Notably, aggregates of Thermococcus coalescens cells fuse together in culture, forming single giant cells.[116] Archaea in the genus Pyrodictium produce an elaborate multicell colony involving arrays of long, thin hollow tubes called cannulae that stick out from the cells' surfaces and connect them into a dense bush-like agglomeration.[117] The function of these cannulae is not settled, but they may allow communication or nutrient exchange with neighbors.[118] Multi-species colonies exist, such as the "string-of-pearls" community that was discovered in 2001 in a German swamp. Round whitish colonies of a novel Methanobacteriati species are spaced along thin filaments that can range up to Lua error: Internal error: The interpreter exited with status 1. long; these filaments are made of a particular bacteria species.[119]
Structure, composition development, and operation
Archaea and bacteria have generally similar cell structure, but cell composition and organization set the archaea apart. Like bacteria, archaea lack interior membranes and organelles.[68] Like bacteria, the cell membranes of archaea are usually bounded by a cell wall and they swim using one or more flagella.[120] Structurally, archaea are most similar to gram-positive bacteria. Most have a single plasma membrane and cell wall, and lack a periplasmic space; the exception to this general rule is Ignicoccus, which possess a particularly large periplasm that contains membrane-bound vesicles and is enclosed by an outer membrane.[121]
Cell wall and archaella
Most archaea (but not Thermoplasma and Ferroplasma) possess a cell wall.[114] In most archaea, the wall is assembled from surface-layer proteins, which form an S-layer.[122] An S-layer is a rigid array of protein molecules that cover the outside of the cell (like chain mail).[123] This layer provides both chemical and physical protection, and can prevent macromolecules from contacting the cell membrane.[124] Unlike bacteria, archaea lack peptidoglycan in their cell walls.[125] Methanobacteriales do have cell walls containing pseudopeptidoglycan, which resembles eubacterial peptidoglycan in morphology, function, and physical structure, but pseudopeptidoglycan is distinct in chemical structure; it lacks D-amino acids and N-acetylmuramic acid, substituting the latter with N-Acetyltalosaminuronic acid.[124]
Archaeal flagella are known as archaella, that operate like bacterial flagella – their long stalks are driven by rotatory motors at the base. These motors are powered by a proton gradient across the membrane, but archaella are notably different in composition and development.[120] The two types of flagella evolved from different ancestors. The bacterial flagellum shares a common ancestor with the type III secretion system,[126][127] while archaeal flagella appear to have evolved from bacterial type IV pili.[128] In contrast with the bacterial flagellum, which is hollow and assembled by subunits moving up the central pore to the tip of the flagella, archaeal flagella are synthesized by adding subunits at the base.[129]
Membranes

Archaeal membranes are made of molecules that are distinctly different from those in all other life forms, showing that archaea are related only distantly to bacteria and eukaryotes.[130] In all organisms, cell membranes are made of molecules known as phospholipids. These molecules possess both a polar part that dissolves in water (the phosphate "head"), and a "greasy" non-polar part that does not (the lipid tail). These dissimilar parts are connected by a glycerol moiety. In water, phospholipids cluster, with the heads facing the water and the tails facing away from it. The major structure in cell membranes is a double layer of these phospholipids, which is called a lipid bilayer.[131]
The phospholipids of archaea are unusual in four ways:
- They have membranes composed of glycerol-ether lipids, whereas bacteria and eukaryotes have membranes composed mainly of glycerol-ester lipids.[132] The difference is the type of bond that joins the lipids to the glycerol moiety; the two types are shown in yellow in the figure at the right. In ester lipids, this is an ester bond, whereas in ether lipids this is an ether bond.[133]
- The stereochemistry of the archaeal glycerol moiety is the mirror image of that found in other organisms. The glycerol moiety can occur in two forms that are mirror images of one another, called enantiomers. Just as a right hand does not fit easily into a left-handed glove, enantiomers of one type generally cannot be used or made by enzymes adapted for the other. The archaeal phospholipids are built on a backbone of sn-glycerol-1-phosphate, which is an enantiomer of sn-glycerol-3-phosphate, the phospholipid backbone found in bacteria and eukaryotes. This suggests that archaea use entirely different enzymes for synthesizing phospholipids as compared to bacteria and eukaryotes. Such enzymes developed very early in life's history, indicating an early split from the other two domains.[130]
- Archaeal lipid tails differ from those of other organisms in that they are based upon long isoprenoid chains with multiple side-branches, sometimes with cyclopropane or cyclohexane rings.[134] By contrast, the fatty acids in the membranes of other organisms have straight chains without side branches or rings. Although isoprenoids play an important role in the biochemistry of many organisms, only the archaea use them to make phospholipids. These branched chains may help prevent archaeal membranes from leaking at high temperatures.[135]
- In some archaea, the lipid bilayer is replaced by a monolayer. In effect, the archaea fuse the tails of two phospholipid molecules into a single molecule with two polar heads (a bolaamphiphile); this fusion may make their membranes more rigid and better able to resist harsh environments.[136] For example, the lipids in Ferroplasma are of this type, which is thought to aid this organism's survival in its highly acidic habitat.[137]
Metabolism
Archaea exhibit a great variety of chemical reactions in their metabolism and use many sources of energy. These reactions are classified into nutritional groups, depending on energy and carbon sources. Some archaea obtain energy from inorganic compounds such as sulfur or ammonia (they are chemotrophs). These include nitrifiers, methanogens and anaerobic methane oxidisers.[138] In these reactions, one compound passes electrons to another (in a redox reaction), releasing energy to fuel the cell's activities. One compound acts as an electron donor and one as an electron acceptor. The energy released is used to generate adenosine triphosphate (ATP) through chemiosmosis, the same basic process that happens in the mitochondrion of eukaryotic cells.[139]
Other groups of archaea use sunlight as a source of energy (they are phototrophs), but oxygen–generating photosynthesis does not occur in any of these organisms.[139] Many basic metabolic pathways are shared among all forms of life; for example, archaea use a modified form of glycolysis (the Entner–Doudoroff pathway) and either a complete or partial citric acid cycle.[140] These similarities to other organisms probably reflect both early origins in the history of life and their high level of efficiency.[141]
| Nutritional type | Source of energy | Source of carbon | Examples |
|---|---|---|---|
| Phototrophs | Sunlight | Organic compounds | Halobacterium |
| Lithotrophs | Inorganic compounds | Organic compounds or carbon fixation | Ferroglobus, Methanobacteria or Pyrolobus |
| Organotrophs | Organic compounds | Organic compounds or carbon fixation | Pyrococcus, Sulfolobus or Methanosarcinales |
Some Methanobacteriati are methanogens (archaea that produce methane as a result of metabolism) living in anaerobic environments, such as swamps. This form of metabolism evolved early, and it is even possible that the first free-living organism was a methanogen.[142] A common reaction involves the use of carbon dioxide as an electron acceptor to oxidize hydrogen. Methanogenesis involves a range of coenzymes that are unique to these archaea, such as coenzyme M and methanofuran.[143] Other organic compounds such as alcohols, acetic acid or formic acid are used as alternative electron acceptors by methanogens. These reactions are common in gut-dwelling archaea. Acetic acid is also broken down into methane and carbon dioxide directly, by acetotrophic archaea. These acetotrophs are archaea in the order Methanosarcinales, and are a major part of the communities of microorganisms that produce biogas.[144]

Other archaea use CO2 in the atmosphere as a source of carbon, in a process called carbon fixation (they are autotrophs). This process involves either a highly modified form of the Calvin cycle[146] or another metabolic pathway called the 3-hydroxypropionate / 4-hydroxybutyrate cycle.[147] The Thermoproteota also use the reverse Krebs cycle while the Methanobacteriati also use the reductive acetyl-CoA pathway.[148] Carbon fixation is powered by inorganic energy sources. No known archaea carry out photosynthesis.[149] Archaeal energy sources are extremely diverse, and range from the oxidation of ammonia by the Nitrosopumilales[150][151] to the oxidation of hydrogen sulfide or elemental sulfur by species of Sulfolobus, using either oxygen or metal ions as electron acceptors.[139]
Phototrophic archaea use light to produce chemical energy in the form of ATP. In the Halobacteria, light-activated ion pumps like bacteriorhodopsin and halorhodopsin generate ion gradients by pumping ions out of and into the cell across the plasma membrane. The energy stored in these electrochemical gradients is then converted into ATP by ATP synthase.[108] This process is a form of photophosphorylation. The ability of these light-driven pumps to move ions across membranes depends on light-driven changes in the structure of a retinol cofactor buried in the center of the protein.[152]
Genetics
Archaea usually have a single circular chromosome,[153] but many euryarchaea have been shown to bear multiple copies of this chromosome.[154] The largest known archaeal genome as of 2002 was 5,751,492 base pairs in Methanosarcina acetivorans.[155] The tiny 490,885 base-pair genome of Nanoarchaeum equitans is one-tenth of this size and the smallest archaeal genome known; it is estimated to contain only 537 protein-encoding genes.[156] Smaller independent pieces of DNA, called plasmids, are also found in archaea. Plasmids may be transferred between cells by physical contact, in a process that may be similar to bacterial conjugation.[157][158]

Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the domain, although most of these unique genes have no known function.[160] Of the remainder of the unique proteins that have an identified function, most belong to the Methanobacteriati and are involved in methanogenesis. The proteins that archaea, bacteria and eukaryotes share form a common core of cell function, relating mostly to transcription, translation, and nucleotide metabolism.[161] Other characteristic archaeal features are the organization of genes of related function – such as enzymes that catalyze steps in the same metabolic pathway into novel operons, and large differences in tRNA genes and their aminoacyl tRNA synthetases.[161]
Transcription in archaea more closely resembles eukaryotic than bacterial transcription, with the archaeal RNA polymerase being very close to its equivalent in eukaryotes,[153] while archaeal translation shows signs of both bacterial and eukaryotic equivalents.[162] Although archaea have only one type of RNA polymerase, its structure and function in transcription seems to be close to that of the eukaryotic RNA polymerase II, with similar protein assemblies (the general transcription factors) directing the binding of the RNA polymerase to a gene's promoter,[163] but other archaeal transcription factors are closer to those found in bacteria.[164] Post-transcriptional modification is simpler than in eukaryotes, since most archaeal genes lack introns, although there are many introns in their transfer RNA and ribosomal RNA genes,[165] and introns may occur in a few protein-encoding genes.[166][167]
Horizontal gene transfer and genetic exchange
Haloferax volcanii, an extreme halophilic archaeon, forms cytoplasmic bridges between cells that appear to be used for transfer of DNA from one cell to another in either direction.[168]
When the hyperthermophilic archaea Sulfolobus solfataricus[169] and Sulfolobus acidocaldarius[170] are exposed to DNA-damaging UV irradiation or to the agents bleomycin or mitomycin C, species-specific cellular aggregation is induced. Aggregation in S. solfataricus could not be induced by other physical stressors, such as pH or temperature shift,[169] suggesting that aggregation is induced specifically by DNA damage. Ajon et al.[170] showed that UV-induced cellular aggregation mediates chromosomal marker exchange with high frequency in S. acidocaldarius. Recombination rates exceeded those of uninduced cultures by up to three orders of magnitude. Frols et al.[169][171] and Ajon et al.[170] hypothesized that cellular aggregation enhances species-specific DNA transfer between Sulfolobus cells in order to provide increased repair of damaged DNA by means of homologous recombination. This response may be a primitive form of sexual interaction similar to the more well-studied bacterial transformation systems that are also associated with species-specific DNA transfer between cells leading to homologous recombinational repair of DNA damage.[172]
Archaeal viruses
Archaea are the target of a number of viruses in a diverse virosphere distinct from bacterial and eukaryotic viruses. They have been organized into 15–18 DNA-based families so far, but multiple species remain un-isolated and await classification.[173][174][175] These families can be informally divided into two groups: archaea-specific and cosmopolitan. Archaeal-specific viruses target only archaean species and currently include 12 families. Numerous unique, previously unidentified viral structures have been observed in this group, including: bottle-shaped, spindle-shaped, coil-shaped, and droplet-shaped viruses.[174] While the reproductive cycles and genomic mechanisms of archaea-specific species may be similar to other viruses, they bear unique characteristics that were specifically developed due to the morphology of host cells they infect.[173] Their virus release mechanisms differ from that of other phages. Bacteriophages generally undergo either lytic pathways, lysogenic pathways, or (rarely) a mix of the two.[176] Most archaea-specific viral strains maintain a stable, somewhat lysogenic, relationship with their hosts – appearing as a chronic infection. This involves the gradual, and continuous, production and release of virions without killing the host cell.[177] Prangishyili (2013) noted that it has been hypothesized that tailed archaeal phages originated from bacteriophages capable of infecting haloarchaeal species. If the hypothesis is correct, it can be concluded that other double-stranded DNA viruses that make up the rest of the archaea-specific group are their own unique group in the global viral community. Krupovic et al. (2018) states that the high levels of horizontal gene transfer, rapid mutation rates in viral genomes, and lack of universal gene sequences have led researchers to perceive the evolutionary pathway of archaeal viruses as a network. The lack of similarities among phylogenetic markers in this network and the global virosphere, as well as external linkages to non-viral elements, may suggest that some species of archaea specific viruses evolved from non-viral mobile genetic elements (MGE).[174]
These viruses have been studied in most detail in thermophilics, particularly the orders Sulfolobales and Thermoproteales.[178] Two groups of single-stranded DNA viruses that infect archaea have been recently isolated. One group is exemplified by the Halorubrum pleomorphic virus 1 (Pleolipoviridae) infecting halophilic archaea,[179] and the other one by the Aeropyrum coil-shaped virus (Spiraviridae) infecting a hyperthermophilic (optimal growth at 90–95 °C) host.[180] Notably, the latter virus has the largest currently reported ssDNA genome. Defenses against these viruses may involve RNA interference from repetitive DNA sequences that are related to the genes of the viruses.[181][182]
Reproduction
Archaea reproduce asexually by binary or multiple fission, fragmentation, or budding; mitosis and meiosis do not occur, so if a species of archaea exists in more than one form, all have the same genetic material.[108] Cell division is controlled in a cell cycle; after the cell's chromosome is replicated and the two daughter chromosomes separate, the cell divides.[183] In the genus Sulfolobus, the cycle has characteristics that are similar to both bacterial and eukaryotic systems. The chromosomes replicate from multiple starting points (origins of replication) using DNA polymerases that resemble the equivalent eukaryotic enzymes.[184]
In Methanobacteriati the cell division protein FtsZ, which forms a contracting ring around the cell, and the components of the septum that is constructed across the center of the cell, are similar to their bacterial equivalents.[183] In cren-[185][186] and thaumarchaea,[187] the cell division machinery Cdv fulfills a similar role. This machinery is related to the eukaryotic ESCRT-III machinery which, while best known for its role in cell sorting, also has been seen to fulfill a role in separation between divided cell, suggesting an ancestral role in cell division.[188]
Both bacteria and eukaryotes, but not archaea, make spores.[189] Some species of Haloarchaea undergo phenotypic switching and grow as several different cell types, including thick-walled structures that are resistant to osmotic shock and allow the archaea to survive in water at low salt concentrations, but these are not reproductive structures and may instead help them reach new habitats.[190]
Behavior
Communication
Quorum sensing was originally thought to not exist in Archaea, but recent studies have shown evidence of some species being able to perform cross-talk through quorum sensing. Other studies have shown syntrophic interactions between archaea and bacteria during biofilm growth. Although research is limited in archaeal quorum sensing, some studies have uncovered LuxR proteins in archaeal species, displaying similarities with bacteria LuxR, and ultimately allowing for the detection of small molecules that are used in high density communication. Similarly to bacteria, Archaea LuxR solos have shown to bind to AHLs (lactones) and non-AHLs ligans, which is a large part in performing intraspecies, interspecies, and interkingdom communication through quorum sensing.[191]
Biofilms
Archaea are known to form biofilms, a common strategy among microorganisms.[192] Quorum sensing is thought to play a role in Archaeal biofilm formation, but less is known about Archaeal quorum sensing than Bacterial quorum sensing. Some Archaea have been observed to form biofilms when the pH is a specific value, without necessarily relying on quorum sensing for instance. These biofilms are sessile communities of micro-organisms (they can contain multiple different species) that produce extracellular polymeric substances, which are used to create a matrix, within which the micro-organisms can grow. Biofilms are beneficial as they: protect organisms from abiotic stresses; facilitate horizontal gene transfer; and enable syntropy to take place.
Biofilm formation has a number of stages: attachment; micro-colony formation; maturation; and dispersal. During the attachment phase the Archaea are reversibly attached to a surface by type-4 pili and archaella. Some Archaea possess other structures involved in attachment such as hami; fimbriae; or cannulae. During the micro colony phase, the extracellular polymeric substance matrix is produced, and numerous archaea have been observed to produce pills and nanowires between cells. During the maturation phase the matrix takes on a sophisticated architecture, such as including pathways for waste to exit the biofilms. Different archaea have been observed to produce biofilms with different architectures. Finally, dispersal can take place, where cells begin to leave the biofilm, this process has been observed in Archaea but the mechanisms behind it are not understood.[192]
Ecology
Habitats

Archaea exist in a broad range of habitats, are recognized as a major part of global ecosystems,[18] and may represent about 20% of microbial cells in the oceans.[193] However, the first-discovered archaeans were extremophiles.[138] Indeed, some archaea survive high temperatures, often above Lua error: Internal error: The interpreter exited with status 1., as found in geysers, black smokers, and oil wells. Other common habitats include very cold habitats and highly saline, acidic, or alkaline water, but archaea include mesophiles that grow in mild conditions, in swamps and marshland, sewage, the oceans, the intestinal tract of animals, and soils.[5][18] Similar to PGPR, Archaea are considered a source of plant growth promotion as well.[5]
Extremophile archaea are members of four main physiological groups. These are the halophiles, thermophiles, alkaliphiles, and acidophiles.[194] These groups are not comprehensive or phylum-specific, nor are they mutually exclusive, since some archaea belong to several groups. Nonetheless, they are a useful starting point for classification.[195]
Halophiles, including the genus Halobacterium, live in extremely saline environments such as salt lakes and outnumber their bacterial counterparts at salinities greater than 20–25%.[138] Thermophiles grow best at temperatures above Lua error: Internal error: The interpreter exited with status 1., in places such as hot springs; hyperthermophilic archaea grow optimally at temperatures greater than Lua error: Internal error: The interpreter exited with status 1..[196] The archaeal Methanopyrus kandleri Strain 116 can even reproduce at Lua error: Internal error: The interpreter exited with status 1., the highest recorded temperature of any organism.[197]
Other archaea exist in very acidic or alkaline conditions.[194] For example, one of the most extreme archaean acidophiles is Picrophilus torridus, which grows at pH 0, which is equivalent to thriving in 1.2 molar sulfuric acid.[198]
This resistance to extreme environments has made archaea the focus of speculation about the possible properties of extraterrestrial life.[199] Some extremophile habitats are not dissimilar to those on Mars,[200] leading to the suggestion that viable microbes could be transferred between planets in meteorites.[201]
Recently, several studies have shown that archaea exist not only in mesophilic and thermophilic environments but are also present, sometimes in high numbers, at low temperatures as well. For example, archaea are common in cold oceanic environments such as polar seas.[202] Even more significant are the large numbers of archaea found throughout the world's oceans in non-extreme habitats among the plankton community (as part of the picoplankton).[203] Although these archaea can be present in extremely high numbers (up to 40% of the microbial biomass), almost none of these species have been isolated and studied in pure culture.[204] Consequently, our understanding of the role of archaea in ocean ecology is rudimentary, so their full influence on global biogeochemical cycles remains largely unexplored.[205] Some marine Thermoproteota are capable of nitrification, suggesting these organisms may affect the oceanic nitrogen cycle,[150] although these oceanic Thermoproteota may also use other sources of energy.[206]
Vast numbers of archaea are also found in the sediments that cover the sea floor, with these organisms making up the majority of living cells at depths over 1 meter below the ocean bottom.[207][208] It has been demonstrated that in all oceanic surface sediments (from 1,000- to 10,000-m water depth), the impact of viral infection is higher on archaea than on bacteria and virus-induced lysis of archaea accounts for up to one-third of the total microbial biomass killed, resulting in the release of ~0.3 to 0.5 gigatons of carbon per year globally.[209]
Role in chemical cycling
Archaea recycle elements such as carbon, nitrogen, and sulfur through their various habitats.[210] Archaea carry out many steps in the nitrogen cycle. This includes both reactions that remove nitrogen from ecosystems (such as nitrate-based respiration and denitrification) as well as processes that introduce nitrogen (such as nitrate assimilation and nitrogen fixation).[211][212]
Researchers recently discovered archaeal involvement in ammonia oxidation reactions. These reactions are particularly important in the oceans.[151][213] The archaea also appear crucial for ammonia oxidation in soils. They produce nitrite, which other microbes then oxidize to nitrate. Plants and other organisms consume the latter.[214]
In the sulfur cycle, archaea that grow by oxidizing sulfur compounds release this element from rocks, making it available to other organisms, but the archaea that do this, such as Sulfolobus, produce sulfuric acid as a waste product, and the growth of these organisms in abandoned mines can contribute to acid mine drainage and other environmental damage.[215]
In the carbon cycle, methanogen archaea remove hydrogen and play an important role in the decay of organic matter by the populations of microorganisms that act as decomposers in anaerobic ecosystems, such as sediments, marshes, and sewage-treatment works.[216]
Interactions with other organisms

The well-characterized interactions between archaea and other organisms are either mutual or commensal. There are no clear examples of known archaeal pathogens or parasites as of 2003,[217][218] but some species of methanogens have been suggested to be involved in infections in the mouth,[219][220] and Nanoarchaeum equitans may be a parasite of another species of archaea, since it only survives and reproduces within the cells of the Crenarchaeon Ignicoccus hospitalis,[156] and appears to offer no benefit to its host.[221]
Mutualism
Mutualism is an interaction between individuals of different species that results in positive (beneficial) effects on per capita reproduction and/or survival of the interacting populations. One well-understood example of mutualism is the interaction between protozoa and methanogenic archaea in the digestive tracts of animals that digest cellulose, such as ruminants and termites.[222] In these anaerobic environments, protozoa break down plant cellulose to obtain energy. This process releases hydrogen as a waste product, but high levels of hydrogen reduce energy production. When methanogens convert hydrogen to methane, protozoa benefit from more energy.[223]
In anaerobic protozoa, such as Plagiopyla frontata, Trimyema, Heterometopus and Metopus contortus, archaea reside inside the protozoa and consume hydrogen produced in their hydrogenosomes.[224][225][226][227][228] Archaea associate with larger organisms, too. For example, the marine archaean Cenarchaeum symbiosum is an endosymbiont of the sponge Axinella mexicana.[229]
Commensalism
Some archaea are commensals, benefiting from an association without helping or harming the other organism. For example, the methanogen Methanobrevibacter smithii is by far the most common archaean in the human flora, making up about one in ten of the prokaryotes in the human gut.[230] In termites and in humans, these methanogens may in fact be mutualists, interacting with other microbes in the gut to aid digestion.[231] Archaean communities associate with a range of other organisms, such as on the surface of corals,[232] and in the region of soil that surrounds plant roots (the rhizosphere).[233][234]
Parasitism
Although Archaea do not have a historical reputation of being pathogens, Archaea are often found with similar genomes to more common pathogens like E. coli,[235] showing metabolic links and evolutionary history with today's pathogens. Archaea have been inconsistently detected in clinical studies because of the lack of categorization of Archaea into more specific species.[236] The reduced genome of Candidatus Sukunaarchaeum mirabile suggests it is a specialized parasite highly dependant on its dinoflagellate host.[237]
Significance in technology and industry
Extremophile archaea, particularly those resistant either to heat or to extremes of acidity and alkalinity, are a source of enzymes that function under these harsh conditions.[238][239] These enzymes have found many uses. For example, thermostable DNA polymerases, such as the Pfu DNA polymerase from Pyrococcus furiosus, revolutionized molecular biology by allowing the polymerase chain reaction to be used in research as a simple and rapid technique for cloning DNA. In industry, amylases, galactosidases and pullulanases in other species of Pyrococcus that function at over Lua error: Internal error: The interpreter exited with status 1. allow food processing at high temperatures, such as the production of low lactose milk and whey.[240] Enzymes from these thermophilic archaea also tend to be very stable in organic solvents, allowing their use in environmentally friendly processes in green chemistry that synthesize organic compounds.[239] This stability makes them easier to use in structural biology. Consequently, the counterparts of bacterial or eukaryotic enzymes from extremophile archaea are often used in structural studies.[241]
In contrast with the range of applications of archaean enzymes, the use of the organisms themselves in biotechnology is less developed. Methanogenic archaea are a vital part of sewage treatment, since they are part of the community of microorganisms that carry out anaerobic digestion and produce biogas.[242] In mineral processing, acidophilic archaea display promise for the extraction of metals from ores, including gold, cobalt and copper.[243]
Archaea host a new class of potentially useful antibiotics. A few of these archaeocins have been characterized, but hundreds more are believed to exist, especially within Halobacteria and Sulfolobus. These compounds differ in structure from bacterial antibiotics, so they may have novel modes of action. In addition, they may allow the creation of new selectable markers for use in archaeal molecular biology.[244]
See also
- Aerobic methane production
- Earliest known life forms
- List of Archaea genera
- List of sequenced archaeal genomes
- Nuclear localization sequence
- Stirrup protein domain
- The Surprising Archaea (book)
- Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya
- Unique properties of hyperthermophilic archaea
- Branching order of bacterial phyla (Genome Taxonomy Database, 2018)
References
- ↑ "Time for a change". Nature 441 (7091): 289. May 2006. doi:10.1038/441289a. PMID 16710401. Bibcode: 2006Natur.441..289P.
- ↑ van Wolferen, Marleen; Pulschen, Andre Arashiro; Baum, Buzz; Gribaldo, Simonetta; Albers, Sonja-Verena (November 2022). "The cell biology of archaea" (in en). Nature Microbiology 7 (11): 1744–1755. doi:10.1038/s41564-022-01215-8. ISSN 2058-5276. PMID 36253512.
- ↑ "Walsby's square bacterium: fine structure of an orthogonal procaryote". Journal of Bacteriology 148 (1): 352–60. October 1981. doi:10.1128/JB.148.1.352-360.1981. PMID 7287626.
- ↑ "Archaea Basic Biology". March 2018. https://basicbiology.net/micro/microorganisms/archaea.
- ↑ 5.0 5.1 5.2 5.3 "An Archaic Approach to a Modern Issue: Endophytic Archaea for Sustainable Agriculture". Current Microbiology 79 (11). September 2022. doi:10.1007/s00284-022-03016-y. PMID 36125558.
- ↑ "Archaea associated with human surfaces: not to be underestimated". FEMS Microbiology Reviews 39 (5): 631–48. September 2015. doi:10.1093/femsre/fuv010. PMID 25907112.
- ↑ "Archaea Are Interactive Components of Complex Microbiomes". Trends in Microbiology 26 (1): 70–85. January 2018. doi:10.1016/j.tim.2017.07.004. PMID 28826642. Bibcode: 2018TrMic..26...70M.
- ↑ "The parasitic lifestyle of an archaeal symbiont". Nature Communications 15 (1). 31 July 2024. doi:10.1038/s41467-024-49962-y. PMID 39085207. Bibcode: 2024NatCo..15.6449H.
- ↑ "The bacterial species dilemma and the genomic-phylogenetic species concept". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 361 (1475): 1899–909. November 2006. doi:10.1098/rstb.2006.1914. PMID 17062409.
- ↑ "Molecules as documents of evolutionary history". Journal of Theoretical Biology 8 (2): 357–66. March 1965. doi:10.1016/0022-5193(65)90083-4. PMID 5876245. Bibcode: 1965JThBi...8..357Z.
- ↑ "A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life". Nature Biotechnology 36 (10): 996–1004. November 2018. doi:10.1038/nbt.4229. PMID 30148503.
- ↑ 12.0 12.1 12.2 12.3 "Phylogenetic structure of the prokaryotic domain: the primary kingdoms". Proceedings of the National Academy of Sciences of the United States of America 74 (11): 5088–90. November 1977. doi:10.1073/pnas.74.11.5088. PMID 270744. Bibcode: 1977PNAS...74.5088W.
- ↑ 13.0 13.1 "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proceedings of the National Academy of Sciences of the United States of America 87 (12): 4576–9. June 1990. doi:10.1073/pnas.87.12.4576. PMID 2112744. Bibcode: 1990PNAS...87.4576W.
- ↑ The new foundations of evolution: on the tree of life. New York: Oxford University Press. 2009. ISBN 978-0-19-973438-2. https://books.google.com/books?id=d7zOviXnbSYC&q=Jan+Sapp.
- ↑ "Archaea". Merriam-Webster Online Dictionary. 2008. http://www.merriam-webster.com/dictionary/archaea.
- ↑ "Are extreme halophiles actually "bacteria"?". Journal of Molecular Evolution 11 (1): 1–8. May 1978. doi:10.1007/bf01768019. PMID 660662. Bibcode: 1978JMolE..11....1M.
- ↑ "Hyperthermophiles in the history of life". Ciba Foundation Symposium 202: 1–10; discussion 11–8. 1996. PMID 9243007.
- ↑ 18.0 18.1 18.2 "Everything in moderation: archaea as 'non-extremophiles'". Current Opinion in Genetics & Development 8 (6): 649–54. December 1998. doi:10.1016/S0959-437X(98)80032-4. PMID 9914204.
- ↑ "Molecular techniques for determining microbial diversity and community structure in natural environments". Critical Reviews in Microbiology 26 (1): 37–57. 2000. doi:10.1080/10408410091154174. PMID 10782339.
- ↑ "The maturing of microbial ecology". International Microbiology 9 (3): 217–23. September 2006. PMID 17061212. http://www.im.microbios.org/0903/0903217.pdf.
- ↑ "Stepping stones towards a new prokaryotic taxonomy". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 361 (1475): 1911–16. November 2006. doi:10.1098/rstb.2006.1915. PMID 17062410.
- ↑ 22.0 22.1 "Phylogenetic diversity and ecology of environmental Archaea". Current Opinion in Microbiology 8 (6): 638–642. December 2005. doi:10.1016/j.mib.2005.10.003. PMID 16236543.
- ↑ Göker, Markus; Oren, Aharon (2023). "Valid publication of four additional phylum names". International Journal of Systematic and Evolutionary Microbiology 73 (9): 006024. doi:10.1099/ijsem.0.006024. ISSN 1466-5034. PMID 37695645.
- ↑ "A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont". Nature 417 (6884): 63–67. May 2002. doi:10.1038/417063a. PMID 11986665. Bibcode: 2002Natur.417...63H.
- ↑ 25.0 25.1 25.2 25.3 Parte, Aidan C.; Sardà Carbasse, Joaquim; Meier-Kolthoff, Jan P.; Reimer, Lorenz C.; Göker, Markus (2020-11-01). "List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ" (in en). International Journal of Systematic and Evolutionary Microbiology 70 (11): 5607–5612. doi:10.1099/ijsem.0.004332. ISSN 1466-5026. PMID 32701423.
- ↑ "Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences". Proceedings of the National Academy of Sciences of the United States of America 93 (17): 9188–93. August 1996. doi:10.1073/pnas.93.17.9188. PMID 8799176. Bibcode: 1996PNAS...93.9188B.
- ↑ "A korarchaeal genome reveals insights into the evolution of the Archaea". Proceedings of the National Academy of Sciences of the United States of America 105 (23): 8102–07. June 2008. doi:10.1073/pnas.0801980105. PMID 18535141. Bibcode: 2008PNAS..105.8102E.
- ↑ "Lineages of acidophilic archaea revealed by community genomic analysis". Science 314 (5807): 1933–35. December 2006. doi:10.1126/science.1132690. PMID 17185602. Bibcode: 2006Sci...314.1933B.
- ↑ "Enigmatic, ultrasmall, uncultivated Archaea". Proceedings of the National Academy of Sciences of the United States of America 107 (19): 8806–11. May 2010. doi:10.1073/pnas.0914470107. PMID 20421484. Bibcode: 2010PNAS..107.8806B.
- ↑ "The archaeal 'TACK' superphylum and the origin of eukaryotes". Trends in Microbiology 19 (12): 580–87. December 2011. doi:10.1016/j.tim.2011.09.002. PMID 22018741. Bibcode: 2011TrMic..19..580G.
- ↑ 31.0 31.1 "Asgard archaea illuminate the origin of eukaryotic cellular complexity". Nature 541 (7637): 353–58. January 2017. doi:10.1038/nature21031. PMID 28077874. Bibcode: 2017Natur.541..353Z. https://escholarship.org/content/qt0qh5400s/qt0qh5400s.pdf?t=pgp8bj.
- ↑ Nina Dombrowski, Jun-Hoe Lee, Tom A Williams, Pierre Offre, Anja Spang (2019). Genomic diversity, lifestyles and evolutionary origins of DPANN archaea. Nature.
- ↑ "Rooting the domain archaea by phylogenomic analysis supports the foundation of the new kingdom Proteoarchaeota". Genome Biology and Evolution 7 (1): 191–204. December 2014. doi:10.1093/gbe/evu274. PMID 25527841.
- ↑ 34.0 34.1 "Integrative modeling of gene and genome evolution roots the archaeal tree of life". Proceedings of the National Academy of Sciences of the United States of America 114 (23): E4602–E4611. June 2017. doi:10.1073/pnas.1618463114. PMID 28533395. Bibcode: 2017PNAS..114E4602W.
- ↑ 35.0 35.1 "Major New Microbial Groups Expand Diversity and Alter our Understanding of the Tree of Life". Cell 172 (6): 1181–1197. 2018. doi:10.1016/j.cell.2018.02.016. PMID 29522741.
- ↑ 36.0 36.1 "GTDB release 10-RS226". https://gtdb.ecogenomic.org/about#4%7C.
- ↑ 37.0 37.1 "ar53_r226.sp_label". https://data.gtdb.ecogenomic.org/releases/release226/226.0/auxillary_files/ar53_r226.sp_labels.tree.
- ↑ 38.0 38.1 "Taxon History". https://gtdb.ecogenomic.org/taxon_history/.
- ↑ "Ernst Mayr and the modern concept of species". Proceedings of the National Academy of Sciences of the United States of America 102 (Supplement 1): 6600–6007. May 2005. doi:10.1073/pnas.0502030102. PMID 15851674. Bibcode: 2005PNAS..102.6600D.
- ↑ "Genetic exchange across a species boundary in the archaeal genus ferroplasma". Genetics 177 (1): 407–16. September 2007. doi:10.1534/genetics.107.072892. PMID 17603112. Bibcode: 2007Genet.177..407E.
- ↑ "Searching for species in haloarchaea". Proceedings of the National Academy of Sciences of the United States of America 104 (35): 14092–97. August 2007. doi:10.1073/pnas.0706358104. PMID 17715057. Bibcode: 2007PNAS..10414092P.
- ↑ "The net of life: reconstructing the microbial phylogenetic network". Genome Research 15 (7): 954–59. July 2005. doi:10.1101/gr.3666505. PMID 15965028.
- ↑ "Exploring prokaryotic diversity in the genomic era". Genome Biology 3 (2). 2002. doi:10.1186/gb-2002-3-2-reviews0003. PMID 11864374.
- ↑ "The uncultured microbial majority". Annual Review of Microbiology 57: 369–94. 2003. doi:10.1146/annurev.micro.57.030502.090759. PMID 14527284. http://pdfs.semanticscholar.org/b762/aceb7d94a73637fb761f9168fc8dc3601bf6.pdf.
- ↑ "Valid publication of the names of forty-two phyla of prokaryotes". Int J Syst Evol Microbiol 71 (10): 5056. 2021. doi:10.1099/ijsem.0.005056. PMID 34694987.
- ↑ Göker, Markus; Oren, Aharon (11 September 2023). "Valid publication of four additional phylum names". International Journal of Systematic and Evolutionary Microbiology 73 (9). doi:10.1099/ijsem.0.006024. PMID 37695645.
- ↑ Valentin-Alvarado, Luis E.; Appler, Kathryn E.; De Anda, Valerie; Schoelmerich, Marie C.; West-Roberts, Jacob; Kivenson, Veronika; Crits-Christoph, Alexander; Ly, Lynn et al. (2024-07-31). "Asgard archaea modulate potential methanogenesis substrates in wetland soil" (in en). Nature Communications 15 (1): 6384. doi:10.1038/s41467-024-49872-z. ISSN 2041-1723. PMID 39085194. Bibcode: 2024NatCo..15.6384V.
- ↑ "Age of the Earth". U.S. Geological Survey. 1997. https://pubs.usgs.gov/gip/geotime/age.html.
- ↑ "The age of the Earth in the twentieth century: a problem (mostly) solved". Special Publications, Geological Society of London 190 (1): 205–21. 2001. doi:10.1144/GSL.SP.2001.190.01.14. Bibcode: 2001GSLSP.190..205D.
- ↑ "Lead isotope study of basic-ultrabasic layered complexes: Speculations about the age of the earth and primitive mantle characteristics". Earth and Planetary Science Letters 47 (3): 370–82. 1980. doi:10.1016/0012-821X(80)90024-2. Bibcode: 1980E&PSL..47..370M.
- ↑ "The Beginnings of Life on Earth". American Scientist. October 1995. http://www.americanscientist.org/issues/pub/the-beginnings-of-life-on-earth/1. Retrieved 15 January 2014.
- ↑ "3.5 billion year old organic deposits show signs of life". Ars Technica. 4 September 2012. https://arstechnica.com/science/2012/09/3-5-billion-year-old-organic-deposts-show-signs-of-life/.
- ↑ "Evidence for biogenic graphite in early Archaean Isua metasedimentary rocks". Nature Geoscience 7 (1): 25. 8 December 2013. doi:10.1038/ngeo2025. Bibcode: 2014NatGe...7...25O.
- ↑ "Oldest fossil found: Meet your microbial mom". Associated Press. 13 November 2013. http://apnews.excite.com/article/20131113/DAA1VSC01.html.
- ↑ "Microbially induced sedimentary structures recording an ancient ecosystem in the ca. 3.48 billion-year-old Dresser Formation, Pilbara, Western Australia". Astrobiology 13 (12): 1103–24. December 2013. doi:10.1089/ast.2013.1030. PMID 24205812. Bibcode: 2013AsBio..13.1103N.
- ↑ "Hints of life on what was thought to be desolate early Earth". Excite. Associated Press (Yonkers, NY: Mindspark Interactive Network). 19 October 2015. http://apnews.excite.com/article/20151019/us-sci--earliest_life-a400435d0d.html.
- ↑ "Potentially biogenic carbon preserved in a 4.1 billion-year-old zircon". Proceedings of the National Academy of Sciences of the United States of America (National Academy of Sciences) 112 (47): 14518–21. November 2015. doi:10.1073/pnas.1517557112. PMID 26483481. PMC 4664351. Bibcode: 2015PNAS..11214518B. http://www.pnas.org/content/early/2015/10/14/1517557112.full.pdf.
- ↑ "Fossil evidence of Archaean life". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 361 (1470): 869–85. June 2006. doi:10.1098/rstb.2006.1834. PMID 16754604.
- ↑ "Polar lipids of archaebacteria in sediments and petroleums". Science 217 (4554): 65–66. July 1982. doi:10.1126/science.217.4554.65. PMID 17739984. Bibcode: 1982Sci...217...65C.
- ↑ "Archean molecular fossils and the early rise of eukaryotes". Science 285 (5430): 1033–36. August 1999. doi:10.1126/science.285.5430.1033. PMID 10446042. Bibcode: 1999Sci...285.1033B.
- ↑ "Reassessing the first appearance of eukaryotes and cyanobacteria". Nature 455 (7216): 1101–4. October 2008. doi:10.1038/nature07381. PMID 18948954. Bibcode: 2008Natur.455.1101R.
- ↑ "Traces of Archaebacteria in ancient sediments". System Applied Microbiology 7 (Archaebacteria '85 Proceedings): 178–83. 1986. doi:10.1016/S0723-2020(86)80002-9. Bibcode: 1986SyApM...7..178H.
- ↑ "Reductive evolution of architectural repertoires in proteomes and the birth of the tripartite world". Genome Research 17 (11): 1572–85. November 2007. doi:10.1101/gr.6454307. PMID 17908824.
- ↑ "Are archaebacteria merely derived 'prokaryotes'?". Nature 289 (5793): 95–96. January 1981. doi:10.1038/289095a0. PMID 6161309. Bibcode: 1981Natur.289...95W.
- ↑ 65.0 65.1 65.2 "The universal ancestor". Proceedings of the National Academy of Sciences of the United States of America 95 (12): 6854–59. June 1998. doi:10.1073/pnas.95.12.6854. PMID 9618502. Bibcode: 1998PNAS...95.6854W.
- ↑ 66.0 66.1 "The early diversification of life and the origin of the three domains: a proposal.". Thermophiles: the keys to molecular evolution and the origin of life.. Athens: Taylor and Francis. August 1998. pp. 19–31. ISBN 978-1-4822-7304-5. https://books.google.com/books?id=c0ZZDwAAQBAJ&pg=PA19.
- ↑ "The origin and evolution of Archaea: a state of the art". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 361 (1470): 1007–22. June 2006. doi:10.1098/rstb.2006.1841. PMID 16754611.
- ↑ 68.0 68.1 "There must be a prokaryote somewhere: microbiology's search for itself". Microbiological Reviews 58 (1): 1–9. March 1994. doi:10.1128/MMBR.58.1.1-9.1994. PMID 8177167.
- ↑ Information is from Willey JM, Sherwood LM, Woolverton CJ. Microbiology 7th ed. (2008), Ch. 19 pp. 474–475, except where noted.
- ↑ "A Complex Endomembrane System in the Archaeon Ignicoccus hospitalis Tapped by Nanoarchaeum equitans". Frontiers in Microbiology 8. 13 June 2017. doi:10.3389/fmicb.2017.01072. PMID 28659892.
- ↑ "Bacterial Metabolism". Medical Microbiology (4th ed.). Galveston (TX): University of Texas Medical Branch at Galveston. 1996. ISBN 978-0-9631172-1-2. https://www.ncbi.nlm.nih.gov/books/NBK7919/.
- ↑ The Surprising Archaea: Discovering Another Domain of Life. Oxford: Oxford University Press. 2000. pp. 25–30. ISBN 978-0-19-511183-5. https://books.google.com/books?id=cWae0vGkX4QC&pg=PA25.
- ↑ 73.0 73.1 73.2 "Archaea – timeline of the third domain". Nature Reviews. Microbiology 9 (1): 51–61. January 2011. doi:10.1038/nrmicro2482. PMID 21132019.
- ↑ "Molecular signatures for the Crenarchaeota and the Thaumarchaeota". Antonie van Leeuwenhoek 99 (2): 133–57. February 2011. doi:10.1007/s10482-010-9488-3. PMID 20711675.
- ↑ "Phylogenomic analysis of proteins that are distinctive of Archaea and its main subgroups and the origin of methanogenesis". BMC Genomics 8 (1): 86. March 2007. doi:10.1186/1471-2164-8-86. PMID 17394648. Bibcode: 2007BMCG....8...86G.
- ↑ "Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbales ord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov". International Journal of Systematic and Evolutionary Microbiology 65 (Pt 3): 1050–69. March 2015. doi:10.1099/ijs.0.070136-0. PMID 25428416. Bibcode: 2015IJSEM..65.1050G.
- ↑ The unique biochemistry of methanogenesis. Progress in Nucleic Acid Research and Molecular Biology. 71. 2002. pp. 223–83. doi:10.1016/s0079-6603(02)71045-3. ISBN 978-0-12-540071-8.
- ↑ "On the origins of cells: a hypothesis for the evolutionary transitions from abiotic geochemistry to chemoautotrophic prokaryotes, and from prokaryotes to nucleated cells". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 358 (1429): 59–85. January 2003. doi:10.1098/rstb.2002.1183. PMID 12594918.
- ↑ "Isoprenoids enhance the stability of fatty acid membranes at the emergence of life potentially leading to an early lipid divide". Interface Focus 9 (6). December 2019. doi:10.1098/rsfs.2019.0067. PMID 31641436.
- ↑ "Toward automatic reconstruction of a highly resolved tree of life". Science 311 (5765): 1283–1287. March 2006. doi:10.1126/science.1123061. PMID 16513982. Bibcode: 2006Sci...311.1283C.
- ↑ "Comparison of archaeal and bacterial genomes: Computer analysis of protein sequences predicts novel functions and suggests a chimeric origin for the archaea". Molecular Microbiology 25 (4): 619–637. August 1997. doi:10.1046/j.1365-2958.1997.4821861.x. PMID 9379893.
- ↑ 82.0 82.1 82.2 82.3 82.4 "Protein phylogenies and signature sequences: A reappraisal of evolutionary relationships among archaebacteria, eubacteria, and eukaryotes". Microbiology and Molecular Biology Reviews 62 (4): 1435–1491. December 1998. doi:10.1128/MMBR.62.4.1435-1491.1998. PMID 9841678.
- ↑ "Were Gram-positive rods the first bacteria?". Trends in Microbiology 11 (4): 166–170. April 2003. doi:10.1016/S0966-842X(03)00063-5. PMID 12706994.
- ↑ 84.0 84.1 84.2 "What are archaebacteria: Life's third domain or monoderm prokaryotes related to gram-positive bacteria? A new proposal for the classification of prokaryotic organisms". Molecular Microbiology 29 (3): 695–707. August 1998. doi:10.1046/j.1365-2958.1998.00978.x. PMID 9723910.
- ↑ "Which is the most conserved group of proteins? Homology-orthology, paralogy, xenology, and the fusion of independent lineages". Journal of Molecular Evolution 39 (5): 541–543. November 1994. doi:10.1007/bf00173425. PMID 7807544. Bibcode: 1994JMolE..39..541G.
- ↑ "Evolutionary relationships of bacterial and archaeal glutamine synthetase genes". Journal of Molecular Evolution 38 (6): 566–576. June 1994. doi:10.1007/BF00175876. PMID 7916055. Bibcode: 1994JMolE..38..566B.
- ↑ "Recent events dominate inter-domain lateral gene transfers between prokaryotes and eukaryotes and, with the exception of endosymbiotic gene transfers, few ancient transfer events persist". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 370 (1678). September 2015. doi:10.1098/rstb.2014.0324. PMID 26323756.
- ↑ 88.0 88.1 88.2 "The natural evolutionary relationships among prokaryotes". Critical Reviews in Microbiology 26 (2): 111–131. 2000. doi:10.1080/10408410091154219. PMID 10890353.
- ↑ "Molecular sequences and the early history of life". Microbial Phylogeny and Evolution: Concepts and controversies. New York, NY: Oxford University Press. 2005. pp. 160–183.
- ↑ "The neomuran origin of archaebacteria, the negibacterial root of the universal tree and bacterial megaclassification". International Journal of Systematic and Evolutionary Microbiology 52 (Pt 1): 7–76. January 2002. doi:10.1099/00207713-52-1-7. PMID 11837318. Bibcode: 2002IJSEM..52....7C.
- ↑ "The origin of a derived superkingdom: How a gram-positive bacterium crossed the desert to become an archaeon". Biology Direct 6: 16. February 2011. doi:10.1186/1745-6150-6-16. PMID 21356104.
- ↑ "Evidence that the root of the tree of life is not within the Archaea". Molecular Biology and Evolution 23 (9): 1648–1651. September 2006. doi:10.1093/molbev/msl046. PMID 16801395.
- ↑ 93.0 93.1 "The role of symbiosis in eukaryotic evolution". Origins and Evolution of Life: An astrobiological perspective. Cambridge: Cambridge University Press. 2011. pp. 326–339. ISBN 978-0-521-76131-4. https://books.google.com/books?id=m3oFebknu1cC&pg=PA326. Retrieved 27 August 2017.
- ↑ "Archaea and the origin of eukaryotes". Nature Reviews. Microbiology 15 (12): 711–723. November 2017. doi:10.1038/nrmicro.2017.133. PMID 29123225.
- ↑ "Origin of the eukaryotic nucleus determined by rate-invariant analysis of rRNA sequences". Nature 331 (6152): 184–86. January 1988. doi:10.1038/331184a0. PMID 3340165. Bibcode: 1988Natur.331..184L.
- ↑ "Evidence for lateral gene transfer between Archaea and bacteria from genome sequence of Thermotoga maritima". Nature 399 (6734): 323–29. May 1999. doi:10.1038/20601. PMID 10360571. Bibcode: 1999Natur.399..323N.
- ↑ "Phylogenetic analysis based on rRNA sequences supports the archaebacterial rather than the eocyte tree". Nature 339 (6220): 145–47. May 1989. doi:10.1038/339145a0. PMID 2497353. Bibcode: 1989Natur.339..145G.
- ↑ "The deep archaeal roots of eukaryotes". Molecular Biology and Evolution 25 (8): 1619–30. August 2008. doi:10.1093/molbev/msn108. PMID 18463089.
- ↑ "An archaeal origin of eukaryotes supports only two primary domains of life". Nature 504 (7479): 231–36. December 2013. doi:10.1038/nature12779. PMID 24336283. Bibcode: 2013Natur.504..231W. https://eprint.ncl.ac.uk/fulltext.aspx?url=199146/704E71B4-8D68-4F3C-93BA-E73C50449593.pdf&pub_id=199146.
- ↑ "Under the Sea, a Missing Link in the Evolution of Complex Cells". The New York Times. 6 May 2015. https://www.nytimes.com/2015/05/07/science/under-the-sea-a-missing-link-in-the-evolution-of-complex-cells.html.
- ↑ "Complex archaea that bridge the gap between prokaryotes and eukaryotes". Nature 521 (7551): 173–179. May 2015. doi:10.1038/nature14447. PMID 25945739. Bibcode: 2015Natur.521..173S.
- ↑ NCBI taxonomy page on Archaea, https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?webmode=Undef&id=2157&lvl=5&lin=f&keep=1&srchmode=1&unlock
- ↑ "Genomic reconstruction of a novel, deeply branched sediment archaeal phylum with pathways for acetogenesis and sulfur reduction". The ISME Journal 10 (7): 1696–705. July 2016. doi:10.1038/ismej.2015.233. PMID 26824177. Bibcode: 2016ISMEJ..10.1696S.
- ↑ "Asgard archaea: Diversity, function, and evolutionary implications in a range of microbiomes". AIMS Microbiology 5 (1): 48–61. 2019. doi:10.3934/microbiol.2019.1.48. PMID 31384702. Bibcode: 2019AIMSM...5...48M.
- ↑ "This Strange Microbe May Mark One of Life's Great Leaps – A organism living in ocean muck offers clues to the origins of the complex cells of all animals and plants.". The New York Times. 15 January 2020. https://www.nytimes.com/2020/01/15/science/cells-eukaryotes-archaea.html.
- ↑ "Isolation of an archaeon at the prokaryote-eukaryote interface". Nature 577 (7791): 519–525. January 2020. doi:10.1038/s41586-019-1916-6. PMID 31942073. Bibcode: 2020Natur.577..519I.
- ↑ "Ribosomal Protein Cluster Organization in Asgard Archaea". Archaea 2023: 1–16. September 2023. doi:10.1155/2023/5512414. PMID 38314098. Bibcode: 2023Archa202312414T.
- ↑ 108.0 108.1 108.2 108.3 Bergey's Manual of Systematic Bacteriology. US: Springer. 2005. pp. 21–26. ISBN 978-0-387-24143-2.
- ↑ "Crenarchaeota". The Tree of Life Web Project. 1997. http://tolweb.org/Crenarchaeota/9.
- ↑ "A square bacterium". Nature 283 (5742): 69–71. 1980. doi:10.1038/283069a0. Bibcode: 1980Natur.283...69W.
- ↑ "An actin homolog of the archaeon Thermoplasma acidophilum that retains the ancient characteristics of eukaryotic actin". Journal of Bacteriology 189 (5): 2039–45. March 2007. doi:10.1128/JB.01454-06. PMID 17189356.
- ↑ "Chaperonin filaments: the archaeal cytoskeleton?". Proceedings of the National Academy of Sciences of the United States of America 94 (10): 5383–88. May 1997. doi:10.1073/pnas.94.10.5383. PMID 9144246. Bibcode: 1997PNAS...94.5383T.
- ↑ "Cytoskeleton in the archaebacterium Thermoplasma acidophilum? Viscosity increase in soluble extracts". Bio Systems 29 (2–3): 151–60. 1993. doi:10.1016/0303-2647(93)90091-P. PMID 8374067. Bibcode: 1993BiSys..29..151H.
- ↑ 114.0 114.1 "Ferroplasma acidiphilum gen. nov., sp. nov., an acidophilic, autotrophic, ferrous-iron-oxidizing, cell-wall-lacking, mesophilic member of the Ferroplasmaceae fam. nov., comprising a distinct lineage of the Archaea". International Journal of Systematic and Evolutionary Microbiology 50 (3): 997–1006. May 2000. doi:10.1099/00207713-50-3-997. PMID 10843038.
- ↑ "Bacterial biofilms: from the natural environment to infectious diseases". Nature Reviews. Microbiology 2 (2): 95–108. February 2004. doi:10.1038/nrmicro821. PMID 15040259.
- ↑ "Thermococcus coalescens sp. nov., a cell-fusing hyperthermophilic archaeon from Suiyo Seamount". International Journal of Systematic and Evolutionary Microbiology 55 (Pt 6): 2507–14. November 2005. doi:10.1099/ijs.0.63432-0. PMID 16280518. Bibcode: 2005IJSEM..55.2507K.
- ↑ "Pyrodictium cannulae enter the periplasmic space but do not enter the cytoplasm, as revealed by cryo-electron tomography". Journal of Structural Biology 141 (1): 34–42. January 2003. doi:10.1016/S1047-8477(02)00581-6. PMID 12576018.
- ↑ "In vivo observation of cell division of anaerobic hyperthermophiles by using a high-intensity dark-field microscope". Journal of Bacteriology 181 (16): 5114–18. August 1999. doi:10.1128/JB.181.16.5114-5118.1999. PMID 10438790.
- ↑ "Natural communities of novel archaea and bacteria growing in cold sulfurous springs with a string-of-pearls-like morphology". Applied and Environmental Microbiology 67 (5): 2336–44. May 2001. doi:10.1128/AEM.67.5.2336-2344.2001. PMID 11319120. Bibcode: 2001ApEnM..67.2336R.
- ↑ 120.0 120.1 "The archaeal flagellum: a different kind of prokaryotic motility structure". FEMS Microbiology Reviews 25 (2): 147–74. April 2001. doi:10.1111/j.1574-6976.2001.tb00575.x. PMID 11250034. Bibcode: 2001FEMMR..25..147T.
- ↑ "The ultrastructure of Ignicoccus: evidence for a novel outer membrane and for intracellular vesicle budding in an archaeon". Archaea 1 (1): 9–18. March 2002. doi:10.1155/2002/307480. PMID 15803654.
- ↑ "S-Layer proteins". Journal of Bacteriology 182 (4): 859–68. February 2000. doi:10.1128/JB.182.4.859-868.2000. PMID 10648507.
- ↑ "Structural research on surface layers: a focus on stability, surface layer homology domains, and surface layer-cell wall interactions". Journal of Structural Biology 124 (2–3): 276–302. December 1998. doi:10.1006/jsbi.1998.4070. PMID 10049812.
- ↑ 124.0 124.1 "Cell wall polymers in Archaea (Archaebacteria)". Cellular and Molecular Life Sciences 54 (4): 305–08. April 1998. doi:10.1007/s000180050156. PMID 9614965.
- ↑ The Surprising Archaea: Discovering Another Domain of Life. Oxford: Oxford University Press. 2000. p. 32. ISBN 978-0-19-511183-5.
- ↑ "Bacterial type III secretion systems are ancient and evolved by multiple horizontal-transfer events". Gene 312: 151–63. July 2003. doi:10.1016/S0378-1119(03)00612-7. PMID 12909351.
- ↑ "Phylogenetic analyses of the constituents of Type III protein secretion systems". Journal of Molecular Microbiology and Biotechnology 2 (2): 125–44. April 2000. PMID 10939240.
- ↑ "Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications". Journal of Molecular Microbiology and Biotechnology 11 (3–5): 167–91. 2006. doi:10.1159/000094053. PMID 16983194.
- ↑ "Prokaryotic motility structures". Microbiology 149 (Pt 2): 295–304. February 2003. doi:10.1099/mic.0.25948-0. PMID 12624192.
- ↑ 130.0 130.1 "Biosynthesis of ether-type polar lipids in archaea and evolutionary considerations". Microbiology and Molecular Biology Reviews 71 (1): 97–120. March 2007. doi:10.1128/MMBR.00033-06. PMID 17347520.
- ↑ "Thermal conductivity and rectification in asymmetric archaeal lipid membranes". The Journal of Chemical Physics 148 (17). May 2018. doi:10.1063/1.5018589. PMID 29739208. Bibcode: 2018JChPh.148q4901Y.
- ↑ "Structure, biosynthesis, and physicochemical properties of archaebacterial lipids". Microbiological Reviews 50 (1): 70–80. March 1986. doi:10.1128/MMBR.50.1.70-80.1986. PMID 3083222.
- ↑ "Ether- versus ester-linked phospholipid bilayers containing either linear or branched apolar chains". Biophysical Journal 107 (6): 1364–74. September 2014. doi:10.1016/j.bpj.2014.07.036. PMID 25229144. Bibcode: 2014BpJ...107.1364B.
- ↑ "Crenarchaeol: The characteristic core glycerol dibiphytanyl glycerol tetraether membrane lipid of cosmopolitan pelagic crenarchaeota". Journal of Lipid Research 43 (10): 1641–51. October 2002. doi:10.1194/jlr.M200148-JLR200. PMID 12364548.
- ↑ "Recent advances in structural research on ether lipids from archaea including comparative and physiological aspects". Bioscience, Biotechnology, and Biochemistry 69 (11): 2019–34. November 2005. doi:10.1271/bbb.69.2019. PMID 16306681.
- ↑ "Archaeal tetraether lipids: unique structures and applications". Applied Biochemistry and Biotechnology 97 (1): 45–62. January 2002. doi:10.1385/ABAB:97:1:45. PMID 11900115.
- ↑ "Tetraether-linked membrane monolayers in Ferroplasma spp: a key to survival in acid". Extremophiles 8 (5): 411–19. October 2004. doi:10.1007/s00792-004-0404-5. PMID 15258835.
- ↑ 138.0 138.1 138.2 "Adaptations to energy stress dictate the ecology and evolution of the Archaea". Nature Reviews. Microbiology 5 (4): 316–23. April 2007. doi:10.1038/nrmicro1619. PMID 17334387.
- ↑ 139.0 139.1 139.2 "Bioenergetics of the Archaea". Microbiology and Molecular Biology Reviews 63 (3): 570–620. September 1999. doi:10.1128/MMBR.63.3.570-620.1999. PMID 10477309. Bibcode: 1999MMBR...63..570S.
- ↑ "Comparative biochemistry of Archaea and Bacteria". Current Opinion in Genetics & Development 1 (4): 544–51. December 1991. doi:10.1016/S0959-437X(05)80206-0. PMID 1822288.
- ↑ "Evolution of carbohydrate metabolic pathways". Research in Microbiology 147 (6–7): 448–55. 1996. doi:10.1016/0923-2508(96)83998-2. PMID 9084754.
- ↑ How did bacteria come to be?. Advances in Microbial Physiology. 40. 1998. pp. 353–99. doi:10.1016/S0065-2911(08)60135-6. ISBN 978-0-12-027740-7.
- ↑ "Unusual coenzymes of methanogenesis". Annual Review of Biochemistry 59: 355–94. 1990. doi:10.1146/annurev.bi.59.070190.002035. PMID 2115763.
- ↑ "Characterization of the methanogenic Archaea within two-phase biogas reactor systems operated with plant biomass". Systematic and Applied Microbiology 31 (3): 190–205. August 2008. doi:10.1016/j.syapm.2008.02.003. PMID 18501543. Bibcode: 2008SyApM..31..190K.
- ↑ Based on PDB 1FBB. Data published in "Molecular mechanism of vectorial proton translocation by bacteriorhodopsin". Nature 406 (6796): 653–57. August 2000. doi:10.1038/35020614. PMID 10949309. Bibcode: 2000Natur.406..653S.
- ↑ "New roads lead to Rubisco in archaebacteria". BioEssays 29 (8): 722–24. August 2007. doi:10.1002/bies.20616. PMID 17621634.
- ↑ "A 3-hydroxypropionate / 4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea". Science 318 (5857): 1782–86. December 2007. doi:10.1126/science.1149976. PMID 18079405. Bibcode: 2007Sci...318.1782B.
- ↑ "Microbiology. A fifth pathway of carbon fixation". Science 318 (5857): 1732–33. December 2007. doi:10.1126/science.1152209. PMID 18079388.
- ↑ "Prokaryotic photosynthesis and phototrophy illuminated". Trends in Microbiology 14 (11): 488–96. November 2006. doi:10.1016/j.tim.2006.09.001. PMID 16997562.
- ↑ 150.0 150.1 "Isolation of an autotrophic ammonia-oxidizing marine archaeon". Nature 437 (7058): 543–46. September 2005. doi:10.1038/nature03911. PMID 16177789. Bibcode: 2005Natur.437..543K.
- ↑ 151.0 151.1 "New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation". The ISME Journal 1 (1): 19–27. May 2007. doi:10.1038/ismej.2007.8. PMID 18043610. Bibcode: 2007ISMEJ...1...19F.
- ↑ "Bacteriorhodopsin". Annual Review of Physiology 66 (1): 665–88. 2004. doi:10.1146/annurev.physiol.66.032102.150049. PMID 14977418. Bibcode: 2004ARPhy..66..665L.
- ↑ 153.0 153.1 "Archaeal genetics – the third way". Nature Reviews Genetics 6 (1): 58–73. January 2005. doi:10.1038/nrg1504. PMID 15630422. http://eprints.nottingham.ac.uk/45896/1/Allers%20%26%20Mevarech%20%282005%29.pdf. Retrieved 12 September 2019.
- ↑ "Genome copy numbers and gene conversion in methanogenic archaea". Journal of Bacteriology 193 (3): 734–43. February 2011. doi:10.1128/JB.01016-10. PMID 21097629.
- ↑ "The genome of M. acetivorans reveals extensive metabolic and physiological diversity". Genome Research 12 (4): 532–42. April 2002. doi:10.1101/gr.223902. PMID 11932238.
- ↑ 156.0 156.1 "The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism". Proceedings of the National Academy of Sciences of the United States of America 100 (22): 12984–88. October 2003. doi:10.1073/pnas.1735403100. PMID 14566062. Bibcode: 2003PNAS..10012984W.
- ↑ "A multicopy plasmid of the extremely thermophilic archaeon Sulfolobus effects its transfer to recipients by mating". Journal of Bacteriology 177 (15): 4417–26. August 1995. doi:10.1128/jb.177.15.4417-4426.1995. PMID 7635827.
- ↑ "Horizontal Gene Transfer Mediated by Plasmids". Plasmids: Current Research and Future Trends. Caister Academic Press. 2008. ISBN 978-1-904455-35-6. http://www.horizonpress.com/pla.
- ↑ "Sulfolobus tengchongensis spindle-shaped virus STSV1: virus-host interactions and genomic features". Journal of Virology 79 (14): 8677–86. July 2005. doi:10.1128/JVI.79.14.8677-8686.2005. PMID 15994761. Bibcode: 2005JVir...79.8677X.
- ↑ "An archaeal genomic signature". Proceedings of the National Academy of Sciences of the United States of America 97 (7): 3304–08. March 2000. doi:10.1073/pnas.050564797. PMID 10716711. Bibcode: 2000PNAS...97.3304G.
- ↑ 161.0 161.1 "Archaeal genomics". Current Opinion in Microbiology 2 (5): 542–47. October 1999. doi:10.1016/S1369-5274(99)00014-4. PMID 10508726.
- ↑ "Ancient ciphers: translation in Archaea". Cell 89 (7): 1007–10. June 1997. doi:10.1016/S0092-8674(00)80288-3. PMID 9215623. Bibcode: 1997Cell...89.1007D.
- ↑ "Structure and function of archaeal RNA polymerases". Molecular Microbiology 65 (6): 1395–404. September 2007. doi:10.1111/j.1365-2958.2007.05876.x. PMID 17697097.
- ↑ "DNA-binding proteins and evolution of transcription regulation in the archaea". Nucleic Acids Research 27 (23): 4658–70. December 1999. doi:10.1093/nar/27.23.4658. PMID 10556324.
- ↑ "Archaeal introns: splicing, intercellular mobility and evolution". Trends in Biochemical Sciences 22 (9): 326–31. September 1997. doi:10.1016/S0968-0004(97)01113-4. PMID 9301331.
- ↑ "Introns in protein-coding genes in Archaea". FEBS Letters 510 (1–2): 27–30. January 2002. doi:10.1016/S0014-5793(01)03219-7. PMID 11755525. Bibcode: 2002FEBSL.510...27W.
- ↑ "Archaeal pre-mRNA splicing: a connection to hetero-oligomeric splicing endonuclease". Biochemical and Biophysical Research Communications 346 (3): 1024–32. August 2006. doi:10.1016/j.bbrc.2006.06.011. PMID 16781672. Bibcode: 2006BBRC..346.1024Y.
- ↑ "The mechanism of DNA transfer in the mating system of an archaebacterium". Science 245 (4924): 1387–89. September 1989. doi:10.1126/science.2818746. PMID 2818746. Bibcode: 1989Sci...245.1387R.
- ↑ 169.0 169.1 169.2 "UV-inducible cellular aggregation of the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by pili formation". Molecular Microbiology 70 (4): 938–52. November 2008. doi:10.1111/j.1365-2958.2008.06459.x. PMID 18990182. https://www.rug.nl/research/portal/en/publications/uvinducible-cellular-aggregation-of-the-hyperthermophilic-archaeon-sulfolobus-solfataricus-is-mediated-by-pili-formation(0dd2a8eb-0f4b-4382-805d-158a870be95e).html.
- ↑ 170.0 170.1 170.2 "UV-inducible DNA exchange in hyperthermophilic archaea mediated by type IV pili". Molecular Microbiology 82 (4): 807–17. November 2011. doi:10.1111/j.1365-2958.2011.07861.x. PMID 21999488. https://pure.rug.nl/ws/files/6771142/2011MolMicrobiolAjon.pdf.
- ↑ "Reactions to UV damage in the model archaeon Sulfolobus solfataricus". Biochemical Society Transactions 37 (Pt 1): 36–41. February 2009. doi:10.1042/BST0370036. PMID 19143598.
- ↑ "Sexual Communication in Archaea, the Precursor to Eukaryotic Meiosis". Biocommunication of Archaea. Springer Nature. 2017. pp. 301–117. doi:10.1007/978-3-319-65536-9_7. ISBN 978-3-319-65535-2.
- ↑ 173.0 173.1 "Archaea – An Introduction" (in en). Encyclopedia of Microbiology (Fourth ed.). Academic Press. January 2019. pp. 243–252. doi:10.1016/B978-0-12-809633-8.20884-4. ISBN 978-0-12-811737-8.
- ↑ 174.0 174.1 174.2 "Viruses of archaea: Structural, functional, environmental and evolutionary genomics". Virus Research 244: 181–193. January 2018. doi:10.1016/j.virusres.2017.11.025. PMID 29175107.
- ↑ "Archaeal viruses and bacteriophages: comparisons and contrasts". Trends in Microbiology 22 (6): 334–44. June 2014. doi:10.1016/j.tim.2014.02.007. PMID 24647075. Bibcode: 2014TrMic..22..334P. http://www.sciencedirect.com/science/article/pii/S0966842X14000419.
- ↑ "Advantages and Limitations of Bacteriophages for the Treatment of Bacterial Infections" (in en). Frontiers in Pharmacology 10. 2019. doi:10.3389/fphar.2019.00513. PMID 31139086.
- ↑ "Brenner's Encyclopedia of Genetics" (in en). Brenner's Encyclopedia of Genetics (Second Edition). Academic Press. 2013-01-01. pp. 295–298. doi:10.1016/B978-0-12-374984-0.01627-2. ISBN 978-0-08-096156-9. http://www.sciencedirect.com/science/article/pii/B9780123749840016272. Retrieved 2020-03-16.
- ↑ "Exceptionally diverse morphotypes and genomes of crenarchaeal hyperthermophilic viruses". Biochemical Society Transactions 32 (Pt 2): 204–08. April 2004. doi:10.1042/BST0320204. PMID 15046572. https://curis.ku.dk/ws/files/51497971/0320204.pdf.
- ↑ "An ssDNA virus infecting archaea: a new lineage of viruses with a membrane envelope". Molecular Microbiology 72 (2): 307–19. April 2009. doi:10.1111/j.1365-2958.2009.06642.x. PMID 19298373.
- ↑ "Archaeal virus with exceptional virion architecture and the largest single-stranded DNA genome". Proceedings of the National Academy of Sciences of the United States of America 109 (33): 13386–91. August 2012. doi:10.1073/pnas.1203668109. PMID 22826255. Bibcode: 2012PNAS..10913386M.
- ↑ "Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements". Journal of Molecular Evolution 60 (2): 174–82. February 2005. doi:10.1007/s00239-004-0046-3. PMID 15791728. Bibcode: 2005JMolE..60..174M.
- ↑ "A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action". Biology Direct 1. March 2006. doi:10.1186/1745-6150-1-7. PMID 16545108.
- ↑ 183.0 183.1 "Archaea and the cell cycle". Molecular Microbiology 29 (4): 955–61. August 1998. doi:10.1046/j.1365-2958.1998.00956.x. PMID 9767564.
- ↑ "Multiple origins of replication in archaea". Trends in Microbiology 12 (9): 399–401. September 2004. doi:10.1016/j.tim.2004.07.001. PMID 15337158.
- ↑ "A unique cell division machinery in the Archaea". Proceedings of the National Academy of Sciences of the United States of America 105 (48): 18942–46. December 2008. doi:10.1073/pnas.0809467105. PMID 18987308. Bibcode: 2008PNAS..10518942L.
- ↑ "A role for the ESCRT system in cell division in archaea". Science 322 (5908): 1710–13. December 2008. doi:10.1126/science.1165322. PMID 19008417. Bibcode: 2008Sci...322.1710S.
- ↑ "Cdv-based cell division and cell cycle organization in the thaumarchaeon Nitrosopumilus maritimus". Molecular Microbiology 82 (3): 555–66. November 2011. doi:10.1111/j.1365-2958.2011.07834.x. PMID 21923770.
- ↑ "Dividing the Archaeal Way: The Ancient Cdv Cell-Division Machinery". Frontiers in Microbiology 9. 2018. doi:10.3389/fmicb.2018.00174. PMID 29551994.
- ↑ "Sporulation genes in members of the low G+C Gram-type-positive phylogenetic branch ( Firmicutes)". Archives of Microbiology 182 (2–3): 182–92. October 2004. doi:10.1007/s00203-004-0696-y. PMID 15340788. Bibcode: 2004ArMic.182..182O.
- ↑ "Cytological peculiarities of some extremely halophilic soil archaeobacteria". Arch. Microbiol. 156 (5): 344–49. 1991. doi:10.1007/BF00248708. Bibcode: 1991ArMic.156..344K.
- ↑ "Computational Exploration of Putative LuxR Solos in Archaea and Their Functional Implications in Quorum Sensing" (in English). Frontiers in Microbiology 8. 2017. doi:10.3389/fmicb.2017.00798. PMID 28515720.
- ↑ 192.0 192.1 van Wolferen, Marleen; Orell, Alvaro; Albers, Sonja-Verena (November 2018). "Archaeal biofilm formation" (in en). Nature Reviews Microbiology 16 (11): 699–713. doi:10.1038/s41579-018-0058-4. ISSN 1740-1534. PMID 30097647.
- ↑ "Environmental diversity of bacteria and archaea". Systematic Biology 50 (4): 470–78. August 2001. doi:10.1080/106351501750435040. PMID 12116647.
- ↑ 194.0 194.1 "Microbial extremophiles at the limits of life". Critical Reviews in Microbiology 33 (3): 183–209. 2007. doi:10.1080/10408410701451948. PMID 17653987.
- ↑ "The growing tree of Archaea: new perspectives on their diversity, evolution and ecology". The ISME Journal 11 (11): 2407–2425. November 2017. doi:10.1038/ismej.2017.122. PMID 28777382. Bibcode: 2017ISMEJ..11.2407A.
- ↑ Brock Biology of Microorganisms (11th ed.). Pearson. 2006. p. 136. ISBN 978-0-13-196893-6.
- ↑ "Cell proliferation at 122 °C and isotopically heavy CH4 production by a hyperthermophilic methanogen under high-pressure cultivation". Proceedings of the National Academy of Sciences of the United States of America 105 (31): 10949–54. August 2008. doi:10.1073/pnas.0712334105. PMID 18664583. Bibcode: 2008PNAS..10510949T.
- ↑ "Another extreme genome: how to live at pH 0". Trends in Microbiology 13 (2): 49–51. February 2005. doi:10.1016/j.tim.2004.12.001. PMID 15680761.
- ↑ "Extreme life on Earth--past, present and possibly beyond". Research in Microbiology 157 (1): 37–48. 2006. doi:10.1016/j.resmic.2005.07.008. PMID 16376523.
- ↑ "Post-Viking microbiology: new approaches, new data, new insights". Origins of Life and Evolution of the Biosphere 29 (1): 73–93. January 1999. doi:10.1023/A:1006515817767. PMID 11536899. Bibcode: 1999OLEB...29...73N. http://www.kluweronline.com/art.pdf?issn=0169-6149&volume=29&page=73. Retrieved 24 June 2008.
- ↑ "The Transfer of Viable Microorganisms Between Planets". Ciba Foundation Symposium 202 – Evolution of Hydrothermal Ecosystems on Earth (And Mars?). Novartis Foundation Symposia. 202. 1996. pp. 304–14; discussion 314–17. doi:10.1002/9780470514986.ch16. ISBN 978-0-470-51498-6.
- ↑ "Diversity of free-living prokaryotes from a deep-sea site at the Antarctic Polar Front". FEMS Microbiology Ecology 36 (2–3): 193–202. July 2001. doi:10.1016/s0168-6496(01)00133-7. PMID 11451524. Bibcode: 2001FEMME..36..193L.
- ↑ "Archaeal dominance in the mesopelagic zone of the Pacific Ocean". Nature 409 (6819): 507–10. January 2001. doi:10.1038/35054051. PMID 11206545. Bibcode: 2001Natur.409..507K.
- ↑ "Molecular diversity and ecology of microbial plankton". Nature 437 (7057): 343–48. September 2005. doi:10.1038/nature04158. PMID 16163344. Bibcode: 2005Natur.437..343G.
- ↑ "Genomic perspectives in microbial oceanography". Nature 437 (7057): 336–42. September 2005. doi:10.1038/nature04157. PMID 16163343. Bibcode: 2005Natur.437..336D.
- ↑ "Major gradients in putatively nitrifying and non-nitrifying Archaea in the deep North Atlantic". Nature 456 (7223): 788–91. December 2008. doi:10.1038/nature07535. PMID 19037244. Bibcode: 2008Natur.456..788A. https://hal.archives-ouvertes.fr/hal-01086735/file/Nature%2C%20HAL.pdf.
- ↑ "Uncultured archaea in deep marine subsurface sediments: have we caught them all?". The ISME Journal 2 (1): 3–18. January 2008. doi:10.1038/ismej.2007.90. PMID 18180743. Bibcode: 2008ISMEJ...2....3T.
- ↑ "Significant contribution of Archaea to extant biomass in marine subsurface sediments". Nature 454 (7207): 991–94. August 2008. doi:10.1038/nature07174. PMID 18641632. Bibcode: 2008Natur.454..991L.
- ↑ "Virus-mediated archaeal hecatomb in the deep seafloor". Science Advances 2 (10). October 2016. doi:10.1126/sciadv.1600492. PMID 27757416. Bibcode: 2016SciA....2E0492D.
- ↑ "Diversity and distribution of Archaea in global estuarine ecosystems". The Science of the Total Environment 637–638: 349–358. October 2018. doi:10.1016/j.scitotenv.2018.05.016. PMID 29753224. Bibcode: 2018ScTEn.637..349L.
- ↑ "Nitrate reduction and the nitrogen cycle in archaea". Microbiology 150 (Pt 11): 3527–46. November 2004. doi:10.1099/mic.0.27303-0. PMID 15528644.
- ↑ "Nitrogen fixation at 92 °C by a hydrothermal vent archaeon". Science 314 (5806): 1783–86. December 2006. doi:10.1126/science.1134772. PMID 17170307. Bibcode: 2006Sci...314.1783M.
- ↑ "Putative ammonia-oxidizing Crenarchaeota in suboxic waters of the Black Sea: A basin-wide ecological study using 16S ribosomal and functional genes and membrane lipids". Environmental Microbiology 9 (4): 1001–16. April 2007. doi:10.1111/j.1462-2920.2006.01227.x. PMID 17359272. Bibcode: 2007EnvMi...9.1001C.
- ↑ "Archaea predominate among ammonia-oxidizing prokaryotes in soils". Nature 442 (7104): 806–09. August 2006. doi:10.1038/nature04983. PMID 16915287. Bibcode: 2006Natur.442..806L.
- ↑ "Microbial communities in acid mine drainage". FEMS Microbiology Ecology 44 (2): 139–52. May 2003. doi:10.1016/S0168-6496(03)00028-X. PMID 19719632. Bibcode: 2003FEMME..44..139B.
- ↑ "Playing scales in the methane cycle: from microbial ecology to the globe". Proceedings of the National Academy of Sciences of the United States of America 101 (34): 12400–01. August 2004. doi:10.1073/pnas.0405075101. PMID 15314221. Bibcode: 2004PNAS..10112400S.
- ↑ "Archaea and their potential role in human disease". Infection and Immunity 71 (2): 591–96. February 2003. doi:10.1128/IAI.71.2.591-596.2003. PMID 12540534.
- ↑ "Pathogenic archaea: do they exist?". BioEssays 25 (11): 1119–28. November 2003. doi:10.1002/bies.10354. PMID 14579252.
- ↑ "Methanogenic Archaea and human periodontal disease". Proceedings of the National Academy of Sciences of the United States of America 101 (16): 6176–81. April 2004. doi:10.1073/pnas.0308766101. PMID 15067114. Bibcode: 2004PNAS..101.6176L.
- ↑ "Identification and quantification of archaea involved in primary endodontic infections". Journal of Clinical Microbiology 44 (4): 1274–82. April 2006. doi:10.1128/JCM.44.4.1274-1282.2006. PMID 16597851.
- ↑ "Nanoarchaeum equitans and Ignicoccus hospitalis: new insights into a unique, intimate association of two archaea". Journal of Bacteriology 190 (5): 1743–50. March 2008. doi:10.1128/JB.01731-07. PMID 18165302. Bibcode: 2008JBact.190.1743J.
- ↑ "Archaeal habitats – from the extreme to the ordinary". Canadian Journal of Microbiology 52 (2): 73–116. February 2006. doi:10.1139/w05-147. PMID 16541146.
- ↑ "Energetics of syntrophic cooperation in methanogenic degradation". Microbiology and Molecular Biology Reviews 61 (2): 262–80. June 1997. doi:10.1128/mmbr.61.2.262-280.1997. PMID 9184013.
- ↑ "Morphology and Phylogeny of a New Species of Anaerobic Ciliate, Trimyema finlayi n. sp., with Endosymbiotic Methanogens". Frontiers in Microbiology 9. 2018. doi:10.3389/fmicb.2018.00140. PMID 29515525.
- ↑ Lua error: Internal error: The interpreter exited with status 1.
- ↑ "Archaea in symbioses". Archaea 2012. 2012. doi:10.1155/2012/596846. PMID 23326206.
- ↑ "Archaea in protozoa and metazoa". Applied Microbiology and Biotechnology 66 (5): 465–474. February 2005. doi:10.1007/s00253-004-1790-4. PMID 15630514.
- ↑ "Multiple acquisition of methanogenic archaeal symbionts by anaerobic ciliates". Molecular Biology and Evolution 17 (2): 251–258. February 2000. doi:10.1093/oxfordjournals.molbev.a026304. PMID 10677847.
- ↑ "A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov". Proceedings of the National Academy of Sciences of the United States of America 93 (13): 6241–6246. June 1996. doi:10.1073/pnas.93.13.6241. PMID 8692799. Bibcode: 1996PNAS...93.6241P.
- ↑ "Diversity of the human intestinal microbial flora". Science 308 (5728): 1635–38. June 2005. doi:10.1126/science.1110591. PMID 15831718. Bibcode: 2005Sci...308.1635E.
- ↑ "A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism". Proceedings of the National Academy of Sciences of the United States of America 103 (26): 10011–16. June 2006. doi:10.1073/pnas.0602187103. PMID 16782812. Bibcode: 2006PNAS..10310011S.
- ↑ "Coral-associated Archaea". Marine Ecology Progress Series 273: 89–96. 2004. doi:10.3354/meps273089. Bibcode: 2004MEPS..273...89W.
- ↑ "The Diversity of Archaea and Bacteria in Association with the Roots of Zea mays L". Microbial Ecology 41 (3): 252–263. April 2001. doi:10.1007/s002480000087. PMID 11391463. Bibcode: 2001MicEc..41..252C.
- ↑ "Crenarchaeota colonize terrestrial plant roots". Environmental Microbiology 2 (5): 495–505. October 2000. doi:10.1046/j.1462-2920.2000.00131.x. PMID 11233158. Bibcode: 2000EnvMi...2..495S.
- ↑ "Have archaeal genes contributed to bacterial virulence?". Trends in Microbiology 12 (5): 213–219. May 2004. doi:10.1016/j.tim.2004.03.002. PMID 15120140.
- ↑ "The search for archaeal pathogens". Reviews in Medical Microbiology 23 (3): 45–51. 2012. doi:10.1097/MRM.0b013e328353c7c9. ISSN 0954-139X.
- ↑ Harada, Ryo; Nishimura, Yuki; Nomura, Mami; Yabuki, Akinori; Shiba, Kogiku; Inaba, Kazuo; Inagaki, Yuji; Nakayama, Takuro (2025-05-02) (in en), A cellular entity retaining only its replicative core: Hidden archaeal lineage with an ultra-reduced genome, doi:10.1101/2025.05.02.651781, http://biorxiv.org/lookup/doi/10.1101/2025.05.02.651781, retrieved 2026-03-10
- ↑ "The hunt for living gold. The search for organisms in extreme environments yields useful enzymes for industry". EMBO Reports 2 (11): 968–71. November 2001. doi:10.1093/embo-reports/kve238. PMID 11713183.
- ↑ 239.0 239.1 "Industrial relevance of thermophilic Archaea". Current Opinion in Microbiology 8 (6): 649–55. December 2005. doi:10.1016/j.mib.2005.10.015. PMID 16257257. Bibcode: 2005COMb....8..649E.
- ↑ "Sources, properties and suitability of new thermostable enzymes in food processing". Critical Reviews in Food Science and Nutrition 46 (3): 197–205. 2006. doi:10.1080/10408690590957296. PMID 16527752.
- ↑ "The impact of extremophiles on structural genomics (and vice versa)". Extremophiles 12 (1): 39–50. January 2008. doi:10.1007/s00792-007-0087-9. PMID 17563834.
- ↑ "Perspectives on biotechnological applications of archaea". Archaea 1 (2): 75–86. September 2002. doi:10.1155/2002/436561. PMID 15803645.
- ↑ "Acidophiles in bioreactor mineral processing". Extremophiles 4 (2): 71–76. April 2000. doi:10.1007/s007920050139. PMID 10805560. Bibcode: 2000Extmo...4.0071N.
- ↑ "Archaeal Antimicrobials: An Undiscovered Country". Archaea: New Models for Prokaryotic Biology. Caister Academic Press. 2008. ISBN 978-1-904455-27-1.
Lua error: Internal error: The interpreter exited with status 1.
Further reading
- The Surprising Archaea: Discovering Another Domain of Life. Oxford University. 2000. ISBN 978-0-19-511183-5. https://books.google.com/books?id=25LQQRBJI8QC.
- Brock Biology of Microorganisms (11th ed.). Englewood Cliffs, N.J: Prentice Hall. 2005. ISBN 978-0-13-144329-7.
- Archaea: Evolution, Physiology and Molecular Biology. WileyBlackwell. 2005. ISBN 978-1-4051-4404-9.
- Archaea: Molecular and Cellular Biology. American Society for Microbiology. 2007. ISBN 978-1-55581-391-8.
- Archaea: New Models for Prokaryotic Biology. Caister Academic Press. 2008. ISBN 978-1-904455-27-1.
- "Archaeal Plasmids". Plasmids: Current Research and Future Trends. Caister Academic Press. 2008. ISBN 978-1-904455-35-6.
- The New Foundations of Evolution: On the Tree of Life. New York: Oxford University Press. 2009. ISBN 978-0-19-538850-3.
- Archaea (Overview) in The Desk Encyclopedia of Microbiology (2nd ed.). San Diego and London: Elsevier Academic Press. 2009. ISBN 978-0-12-374980-2.
External links
General
- Introduction to the Archaea, ecology, systematics and morphology
- Oceans of Archaea – E.F. DeLong, ASM News, 2003
Classification
- NCBI taxonomy page on Archaea
- Genera of the domain Archaea – list of Prokaryotic names with Standing in Nomenclature
- Shotgun sequencing finds nanoorganisms – discovery of the ARMAN group of archaea
Genomics
- Browse any completed archaeal genome at UCSC
- Comparative Analysis of Archaeal Genomes (at DOE's IMG system)
Lua error: Internal error: The interpreter exited with status 1. Lua error: Internal error: The interpreter exited with status 1. Lua error: Internal error: The interpreter exited with status 1. Lua error: Internal error: The interpreter exited with status 1. Lua error: Internal error: The interpreter exited with status 1. Wikidata ☰ Q10872 entry
Lua error: Internal error: The interpreter exited with status 1.