Biology:Single-cell transcriptomics
Single-cell transcriptomics examines the gene expression level of individual cells in a given population by simultaneously measuring the RNA concentration, typically messenger RNA (mRNA), of hundreds to thousands of genes.[1] Single-cell transcriptomics makes it possible to unravel heterogeneous cell populations, reconstruct cellular developmental pathways, and model transcriptional dynamics—all previously masked in bulk RNA sequencing.[2]
Background
The development of high-throughput RNA sequencing (RNA-seq) and microarrays has made gene expression analysis a routine. RNA analysis was previously limited to tracing individual transcripts by Northern blots or quantitative PCR. Higher throughput and speed allow researchers to frequently characterize the expression profiles of populations of thousands of cells. The data from bulk assays has led to identifying genes differentially expressed in distinct cell populations, and biomarker discovery.[3]

These studies are limited as they provide measurements for whole tissues and, as a result, show an average expression profile for all the constituent cells. This has a couple of drawbacks. Firstly, different cell types within the same tissue can have distinct roles in multicellular organisms. They often form subpopulations with unique transcriptional profiles. Correlations in the gene expression of the subpopulations can often be missed due to the lack of subpopulation identification.[1] Secondly, bulk assays fail to recognize whether a change in the expression profile is due to a change in regulation or composition — for example if one cell type arises to dominate the population. Lastly, when one's goal is to study cellular progression through differentiation, average expression profiles can only order cells by time rather than by developmental stage. Consequently, they cannot show trends in gene expression levels specific to certain stages.[4]
Recent advances in biotechnology allow the measurement of gene expression in hundreds to thousands of individual cells simultaneously. While these breakthroughs in transcriptomics technologies have enabled the generation of single-cell transcriptomic data, they also presented new computational and analytical challenges. Bioinformaticians can use techniques from bulk RNA-seq for single-cell data. Still, many new computational approaches have had to be designed for this data type to facilitate a complete and detailed study of single-cell expression profiles.[5]
Experimental steps
There is so far no standardized technique to generate single-cell data: all methods must include cell isolation from the population, lysate formation, amplification through reverse transcription, and quantification of expression levels. Common techniques for measuring expression are quantitative PCR or RNA-seq.[6]
Isolating single cells

Several methods are available to isolate and amplify cells for single-cell analysis, differing primarily in throughput and potential for cell selection. Low-throughput techniques, such as micropipetting, cytoplasmic aspiration,[7] and laser capture microdissection, typically isolate hundreds of cells but enable deliberate cell selection.
High-throughput methods allow for the rapid isolation of hundreds to tens of thousands of cells.[8] Common high-throughput approaches include Fluorescence Activated Cell Sorting (FACS) and the use of microfluidic devices. Microfluidic platforms often isolate single cells either by mechanical separation into microwells (e.g., BD Rhapsody, Takara ICELL8, Vycap Puncher Platform, CellMicrosystems CellRaft) or by encapsulation within droplets (e.g., 10x Genomics Chromium, Illumina Bio-Rad ddSEQ, 1CellBio InDrop, Dolomite Bio Nadia).[9] Furthermore, optimized protocols have been developed by integrating these isolation techniques directly with scRNA-seq workflows. For instance, combining FACS with scRNA-seq led to protocols like SORT-seq,[10] and a list of studies utilizing SORT-seq can be found here.[11] Similarly, the integration of microfluidic devices with scRNA-seq has been highly optimized in protocols such as those developed by 10x Genomics.[12]
Single cell RNA-seq techniques that rely on split-pool barcoding can uniquely label cells without requiring the isolation of individual cells, including sci-RNA-seq, SPLiT-seq, and microSPLiT.[13][14][15]
Quantitative PCR (qPCR)
To measure the level of expression of each transcript qPCR can be applied. Gene specific primers are used to amplify the corresponding gene as with regular PCR and as a result data is usually only obtained for sample sizes of less than 100 genes. The inclusion of housekeeping genes, whose expression should be constant under the conditions, is used for normalization. The most commonly used house keeping genes include GAPDH and α-actin, although the reliability of normalization through this process is questionable as there is evidence that the level of expression can vary significantly.[16] Fluorescent dyes are used as reporter molecules to detect the PCR product and monitor the progress of the amplification - the increase in fluorescence intensity is proportional to the amplicon concentration. A plot of fluorescence vs. cycle number is made and a threshold fluorescence level is used to find cycle number at which the plot reaches this value. The cycle number at this point is known as the threshold cycle (Ct) and is measured for each gene.[17]
Single-cell RNA-seq (scRNA-Seq)
The single-cell RNA-seq technique converts a population of RNAs to a library of cDNA fragments that can be sequenced. In droplet-based technologies such as 10x Genomics Chromium, single cells are isolated in droplets together with beads coated with barcoded oligonucleotides. Both cells and beads are supplied in limited amounts such that co-occupancy with multiple cells and beads is a very rare event. Cells are lysed within the droplets, and RNAs are reverse transcribed using the barcoded oligo-dT oligonucleotides as primers. After reverse transcription, the emulsion is broken, releasing the barcoded cDNA from all the droplets into a single solution. This pooled cDNA is then prepared for sequencing via the addition of sequencing adapters and PCR amplification.

These fragments are sequenced by high-throughput next generation sequencing techniques and the reads are mapped back to the reference genome, providing a count of the number of reads associated with each gene.[18] Transcripts from a particular cell are identified by each cell's unique barcode.[19][20]
Normalization of RNA-Seq data accounts for cell to cell variation in the efficiencies of the cDNA library formation and sequencing. One method relies on the use of extrinsic RNA spike-ins that are added in equal quantities to each cell lysate and used to normalize read count by the number of reads mapped to spike-in mRNA.[21] Another control uses unique molecular identifiers (UMIs)-short DNA sequences (6–10nt) that are added to each cDNA before amplification and act as a bar code for each cDNA molecule. Normalization is achieved by using the count number of unique UMIs associated with each gene to account for differences in amplification efficiency.[22]
A combination of both spike-ins, UMIs and other approaches have been combined to help identify artifacts during library preparation[23] and for more accurate normalization.
Applications
scRNA-Seq is becoming widely used across biological disciplines including Development, Neurology,[24] Oncology,[25][26][27] Autoimmune disease,[28] Infectious disease.,[29] brain disease,[30] and environmental virology.[31][32] Several scRNA-Seq protocols have been published: Tang et al.,[33] STRT,[34] SMART-seq,[35] CEL-seq,[36] RAGE-seq,[37] Quartz-seq[38] and C1-CAGE.[39] These protocols differ in terms of strategies for reverse transcription, cDNA synthesis and amplification, and the possibility to accommodate sequence-specific barcodes (i.e. UMIs) or the ability to process pooled samples.[40] In 2017, two approaches were introduced to simultaneously measure single-cell mRNA and protein expression through oligonucleotide-labeled antibodies known as REAP-seq,[41] and CITE-seq.[42] A 2025 review in Science reported that applying single-cell transcriptomics to microbial communities reveals functional heterogeneity within gut communities, characteristic antibiotic responses, and the dynamics of mobile genetic elements. Pountain, Andrew W.; Yanai, Itai (2025-09-04). "Dissecting microbial communities with single-cell transcriptome analysis". Science 389 (6764). doi:10.1126/science.adp6252. PMID 40906858.
scRNA-Seq has provided considerable insight into the development of embryos and organisms, including the worm Caenorhabditis elegans,[43] and the regenerative planarian Schmidtea mediterranea.[44][45] The first vertebrate animals to be mapped in this way were Zebrafish[46][47] and Xenopus laevis.[48] In each case multiple stages of the embryo were studied, allowing the entire process of development to be mapped on a cell-by-cell basis.[49] Science recognized these advances as the 2018 Breakthrough of the Year.[50]
Considerations
A problem associated with single-cell data occurs in the form of zero inflated gene expression distributions, known as technical dropouts, that are common due to low mRNA concentrations of less-expressed genes that are not captured in the reverse transcription process. The percentage of mRNA molecules in the cell lysate that are detected is often only 10-20%.[51]
When using RNA spike-ins for normalization the assumption is made that the amplification and sequencing efficiencies for the endogenous and spike-in RNA are the same. Evidence suggests that this is not the case given fundamental differences in size and features, such as the lack of a polyadenylated tail in spike-ins and therefore shorter length.[52] Additionally, normalization using UMIs assumes the cDNA library is sequenced to saturation, which is not always the case.[22]
In the amplification step, either PCR or in vitro transcription (IVT) is currently used to amplify cDNA. One of the advantages of PCR-based methods is the ability to generate full-length cDNA. However, different PCR efficiency on particular sequences (for instance, GC content and snapback structure) may also be exponentially amplified, producing libraries with uneven coverage. On the other hand, while libraries generated by IVT can avoid PCR-induced sequence bias, specific sequences may be transcribed inefficiently, thus causing sequence drop-out or generating incomplete sequences.[53][54]
Challenges for scRNA-Seq include preserving the initial relative abundance of mRNA in a cell and identifying rare transcripts.[55] The reverse transcription step is critical as the efficiency of the RT reaction determines how much of the cell's RNA population will be eventually analyzed by the sequencer. The processivity of reverse transcriptases and the priming strategies used may affect full-length cDNA production and the generation of libraries biased toward the 3' or 5' end of genes.
A further consideration when sequencing large, branched cell types, such as neurons, comes from the removal of distal processes containing local pools of RNA during the single-cell isolation process. In these cells, scRNA-seq datasets only capture transcript in the central cell body, omitting transcripts from RNA pools localized to cellular processes that can be involved in local translation or other RNA-mediated subcellular mechanisms. In the brain it has been estimated that over 40% of total RNA is not sequenced by scRNA-seq due to the prevalence of local transcriptomes in cellular processes such as axons, dendrites, myelin, and endfeet.[56]
Data analysis
Insights based on single-cell data analysis assume that the input is a matrix of normalized gene expression counts, generated by the approaches outlined above, and can provide opportunities that are not obtainable by bulk.
Three main insights provided:[57]
- Identification and characterization of cell types and their spatial organisation in time
- Inference of gene regulatory networks and their strength across individual cells
- Classification of the stochastic component of transcription
The techniques outlined have been designed to help visualise and explore patterns in the data in order to facilitate the revelation of these three features.
Clustering

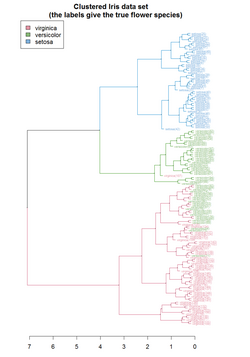
Clustering allows for the formation of subgroups in the cell population. Cells can be clustered by their transcriptomic profile in order to analyse the sub-population structure and identify rare cell types or cell subtypes. Alternatively, genes can be clustered by their expression states in order to identify covarying genes. A combination of both clustering approaches, known as biclustering, has been used to simultaneously cluster by genes and cells to find genes that behave similarly within cell clusters.[58]
Clustering methods applied can be K-means clustering, forming disjoint groups or Hierarchical clustering, forming nested partitions.
Biclustering
Biclustering provides several advantages by improving the resolution of clustering. Genes that are only informative to a subset of cells and are hence only expressed there can be identified through biclustering. Moreover, similarly behaving genes that differentiate one cell cluster from another can be identified using this method.[59]
Dimensionality reduction

Dimensionality reduction algorithms such as Principal component analysis (PCA) and t-SNE can be used to simplify data for visualisation and pattern detection by transforming cells from a high to a lower dimensional space. The result of this method produces graphs with each cell as a point in a 2-D or 3-D space. Dimensionality reduction is frequently used before clustering as cells in high dimensions can wrongly appear to be close due to distance metrics behaving non-intuitively.[60]
Principal component analysis
The most frequently used technique is PCA, which identifies the directions of largest variance principal components and transforms the data so that the first principal component has the largest possible variance, and successive principle components in turn each have the highest variance possible while remaining orthogonal to the preceding components. The contribution each gene makes to each component is used to infer which genes are contributing the most to variance in the population and are involved in differentiating different subpopulations.[61]
Differential expression
Detecting differences in gene expression level between two populations is used both single-cell and bulk transcriptomic data. Specialised methods have been designed for single-cell data that considers single cell features such as technical dropouts and shape of the distribution e.g. Bimodal vs. unimodal.[62]
Gene ontology enrichment
Gene ontology terms describe gene functions and the relationships between those functions into three classes:
- Molecular function
- Cellular component
- Biological process
Gene Ontology (GO) term enrichment is a technique used to identify which GO terms are over-represented or under-represented in a given set of genes. In single-cell analysis input list of genes of interest can be selected based on differentially expressed genes or groups of genes generated from biclustering. The number of genes annotated to a GO term in the input list is normalized against the number of genes annotated to a GO term in the background set of all genes in genome to determine statistical significance.[63]
Pseudotemporal ordering

Pseudo-temporal ordering (or trajectory inference) is a technique that aims to infer gene expression dynamics from snapshot single-cell data. The method tries to order the cells in such a way that similar cells are closely positioned to each other. This trajectory of cells can be linear, but can also bifurcate or follow more complex graph structures. The trajectory, therefore, enables the inference of gene expression dynamics and the ordering of cells by their progression through differentiation or response to external stimuli. The method relies on the assumptions that the cells follow the same path through the process of interest and that their transcriptional state correlates to their progression. The algorithm can be applied to both mixed populations and temporal samples.
More than 50 methods for pseudo-temporal ordering have been developed, and each has its own requirements for prior information (such as starting cells or time course data), detectable topologies, and methodology.[64] An example algorithm is the Monocle algorithm[65] that carries out dimensionality reduction of the data, builds a minimal spanning tree using the transformed data, orders cells in pseudo-time by following the longest connected path of the tree and consequently labels cells by type. Another example is the diffusion pseudotime (DPT) algorithm,[63] which uses a diffusion map and diffusion process. Another class of methods such as MARGARET [66] employ graph partitioning for capturing complex trajectory topologies such as disconnected and multifurcating trajectories.
Network inference
Gene regulatory network inference is a technique that aims to construct a network, shown as a graph, in which the nodes represent the genes and edges indicate co-regulatory interactions. The method relies on the assumption that a strong statistical relationship between the expression of genes is an indication of a potential functional relationship.[67] The most commonly used method to measure the strength of a statistical relationship is correlation. However, correlation fails to identify non-linear relationships and mutual information is used as an alternative. Gene clusters linked in a network signify genes that undergo coordinated changes in expression.[68]
Integration
The presence or strength of technical effects and the types of cells observed often differ in single-cell transcriptomics datasets generated using different experimental protocols and under different conditions. This difference results in strong batch effects that may bias the findings of statistical methods applied across batches, particularly in the presence of confounding.[69] As a result of the aforementioned properties of single-cell transcriptomic data, batch correction methods developed for bulk sequencing data were observed to perform poorly. Consequently, researchers developed statistical methods to correct for batch effects that are robust to the properties of single-cell transcriptomic data to integrate data from different sources or experimental batches. Laleh Haghverdi performed foundational work in formulating the use of mutual nearest neighbors between each batch to define batch correction vectors.[70] With these vectors, one can merge datasets that each include at least one shared cell type. An orthogonal approach involves the projection of each dataset onto a shared low-dimensional space using canonical correlation analysis.[71] Mutual nearest neighbors and canonical correlation analysis have also been combined to define integration "anchors" comprising reference cells in one dataset, to which query cells in another dataset are normalized.[72] Another class of methods (e.g., scDREAMER[73]) uses deep generative models such as variational autoencoders for learning batch-invariant latent cellular representations which can be used for downstream tasks such as cell type clustering, denoising of single-cell gene expression vectors and trajectory inference.[66]
Cell-cell Communication
Cell-cell communication tools leverage the expression of cognate ligand and receptor pairs in distinct cell pairs to predict putative interactions between cells.[74] A pioneer tool is CellPhoneDB,[75] which included a curate database of receptor complexes and ligand-receptor interactions. The method also implemented a statistical test by evaluating the expression of the ligand-receptor in the respective cell pair. One interesting advanced analysis is at scenarios when two conditions (disease vs. controls) are available. Here, one is interested in finding cell-cell interactions specific to disease or control conditions.[76]
Analysis of Disease Atlas
With the lowering of single cell cost, it is possible to measure single cell data over large disease cohorts with up to 100 individual samples and million of cells. This brings several challenges such as individual variability and the multi-scale properties of the data, every single cell experiment represents a distribution of cells. MILO[77] tackle individual variability by encoding single cell data as a k-nn graph followed by a differential abundance test. This allows to find cellular neighboorhoods associated to a particular disease condition. Optimal transport theory is another represents another framework to tackle multi-scale nature of single cell diseae atlas. By encoding single cell experiments of an patient as a distribution of cells, optimal transport based Wasserstein distance allows to measure a distance between two patient. This can be used for sample level clustering and trajectory analysis as explored by PILOT[78] and QOT.[79]
Software frameworks
An important pilar of single cell analysis are software packages providing functionally for quality check, normalization, clustering and down-stream analysis described above. They also provide data containers for the storage and representation of analysed single cell data. Seurat is the first and possibly most complete frameworks for single cell analysis. Scanpy is an alternative for python language. Additional software package offer specific data analysis taks. Notable examples is LIANA+ for cell-cell communication analysis, scVI as a package offering variational autoencoders for analysis of single cell data, and PILOT as a package for sample level analysis of disease atlas[80][81]
See also
- RNA-Seq
- Single-cell analysis
- Single-cell multi-omics integration
- Single-cell sequencing
- Transcriptome
- Transcriptomics
References
- ↑ 1.0 1.1 Kanter, Itamar; Kalisky, Tomer (2015). "Single cell transcriptomics: methods and applications". Frontiers in Oncology 5: 53. doi:10.3389/fonc.2015.00053. ISSN 2234-943X. PMID 25806353.
- ↑ Liu, Serena; Trapnell, Cole (2016). "Single-cell transcriptome sequencing: recent advances and remaining challenges". F1000Research 5: F1000 Faculty Rev–182. doi:10.12688/f1000research.7223.1. ISSN 2046-1402. PMID 26949524.
- ↑ Szabo, David T. (2014). "Chapter 62 - Transcriptomic biomarkers in safety and risk assessment of chemicals". Biomarkers in Toxicology. Academic Press. pp. 1033–1038. ISBN 978-0-12-404630-6.
- ↑ Trapnell, Cole (October 2015). "Defining cell types and states with single-cell genomics". Genome Research 25 (10): 1491–1498. doi:10.1101/gr.190595.115. ISSN 1549-5469. PMID 26430159.
- ↑ Stegle, O.; Teichmann, S.; Marioni, J. (2015). "Computational and analytical challenges in single-cell transcriptomics". Nature Reviews Genetics 16 (3): 133–145. doi:10.1038/nrg3833. PMID 25628217.
- ↑ Kolodziejczyk, Aleksandra A.; Kim, Jong Kyoung; Svensson, Valentine; Marioni, John C.; Teichmann, Sarah A. (May 2015). "The Technology and Biology of Single-Cell RNA Sequencing". Molecular Cell 58 (4): 610–620. doi:10.1016/j.molcel.2015.04.005. PMID 26000846.
- ↑ "Cytoplasmic aspiration | Single Cell Analysis". http://www.single-cell-analysis.com/tag/cytoplasmic-aspiration/.
- ↑ Poulin, Jean-Francois; Tasic, Bosiljka; Hjerling-Leffler, Jens; Trimarchi, Jeffrey M.; Awatramani, Rajeshwar (1 September 2016). "Disentangling neural cell diversity using single-cell transcriptomics" (in en). Nature Neuroscience 19 (9): 1131–1141. doi:10.1038/nn.4366. ISSN 1097-6256. PMID 27571192.
- ↑ "Platforms for Single-Cell Collection and Analysis". International Journal of Molecular Sciences 19 (3): 807. March 2018. doi:10.3390/ijms19030807. PMID 29534489.
- ↑ Muraro, Mauro J.; Dharmadhikari, Gitanjali; Grün, Dominic; Groen, Nathalie; Dielen, Tim; Jansen, Erik; van Gurp, Leon; Engelse, Marten A. et al. (2016-10-26). "A Single-Cell Transcriptome Atlas of the Human Pancreas". Cell Systems 3 (4): 385–394.e3. doi:10.1016/j.cels.2016.09.002. ISSN 2405-4712. PMID 27693023.
- ↑ "SORT-seq Archives" (in en-US). https://www.scdiscoveries.com/publications/services/sort-seq-publications/.
- ↑ Zheng, Grace X. Y.; Terry, Jessica M.; Belgrader, Phillip; Ryvkin, Paul; Bent, Zachary W.; Wilson, Ryan; Ziraldo, Solongo B.; Wheeler, Tobias D. et al. (2017-01-16). "Massively parallel digital transcriptional profiling of single cells" (in en). Nature Communications 8 (1). doi:10.1038/ncomms14049. ISSN 2041-1723. PMID 28091601. Bibcode: 2017NatCo...814049Z.
- ↑ Cao, Junyue; Packer, Jonathan S.; Ramani, Vijay; Cusanovich, Darren A.; Huynh, Chau; Daza, Riza; Qiu, Xiaojie; Lee, Choli et al. (2017-08-18). "Comprehensive single-cell transcriptional profiling of a multicellular organism". Science 357 (6352): 661–667. doi:10.1126/science.aam8940. ISSN 1095-9203. PMID 28818938. Bibcode: 2017Sci...357..661C.
- ↑ Rosenberg, Alexander B.; Roco, Charles M.; Muscat, Richard A.; Kuchina, Anna; Sample, Paul; Yao, Zizhen; Graybuck, Lucas T.; Peeler, David J. et al. (2018-04-13). "Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding". Science 360 (6385): 176–182. doi:10.1126/science.aam8999. ISSN 1095-9203. PMID 29545511. Bibcode: 2018Sci...360..176R.
- ↑ Gaisser, Karl D.; Skloss, Sophie N.; Brettner, Leandra M.; Paleologu, Luana; Roco, Charles M.; Rosenberg, Alexander B.; Hirano, Matthew; DePaolo, R. William et al. (October 2024). "High-throughput single-cell transcriptomics of bacteria using combinatorial barcoding". Nature Protocols 19 (10): 3048–3084. doi:10.1038/s41596-024-01007-w. ISSN 1750-2799. PMID 38886529.
- ↑ Radonić, Aleksandar; Thulke, Stefanie; Mackay, Ian M.; Landt, Olfert; Siegert, Wolfgang; Nitsche, Andreas (23 January 2004). "Guideline to reference gene selection for quantitative real-time PCR". Biochemical and Biophysical Research Communications 313 (4): 856–862. doi:10.1016/j.bbrc.2003.11.177. ISSN 0006-291X. PMID 14706621. Bibcode: 2004BBRC..313..856R.
- ↑ Wildsmith, S. E.; Archer, G. E.; Winkley, A. J.; Lane, P. W.; Bugelski, P. J. (1 January 2001). "Maximization of signal derived from cDNA microarrays". BioTechniques 30 (1): 202–206, 208. doi:10.2144/01301dd04. ISSN 0736-6205. PMID 11196312.
- ↑ Wang, Zhong; Gerstein, Mark; Snyder, Michael (23 March 2017). "RNA-Seq: a revolutionary tool for transcriptomics". Nature Reviews. Genetics 10 (1): 57–63. doi:10.1038/nrg2484. ISSN 1471-0056. PMID 19015660.
- ↑ "Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells". Cell 161 (5): 1187–1201. May 2015. doi:10.1016/j.cell.2015.04.044. PMID 26000487. Bibcode: 2015Cell..161.1187K.
- ↑ "Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets". Cell 161 (5): 1202–1214. May 2015. doi:10.1016/j.cell.2015.05.002. PMID 26000488. Bibcode: 2015Cell..161.1202M.
- ↑ Jiang, Lichun; Schlesinger, Felix; Davis, Carrie A.; Zhang, Yu; Li, Renhua; Salit, Marc; Gingeras, Thomas R.; Oliver, Brian (23 March 2017). "Synthetic spike-in standards for RNA-seq experiments". Genome Research 21 (9): 1543–1551. doi:10.1101/gr.121095.111. ISSN 1088-9051. PMID 21816910.
- ↑ 22.0 22.1 Islam, Saiful; Zeisel, Amit; Joost, Simon; La Manno, Gioele; Zajac, Pawel; Kasper, Maria; Lönnerberg, Peter; Linnarsson, Sten (1 February 2014). "Quantitative single-cell RNA-seq with unique molecular identifiers" (in en). Nature Methods 11 (2): 163–166. doi:10.1038/nmeth.2772. ISSN 1548-7091. PMID 24363023. https://infoscience.epfl.ch/handle/20.500.14299/156221.
- ↑ "Quantitative single-cell RNA-seq with unique molecular identifiers". Nature Methods 11 (2): 163–6. February 2014. doi:10.1038/nmeth.2772. PMID 24363023. https://infoscience.epfl.ch/handle/20.500.14299/156221.
- ↑ "Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain". Nature Biotechnology 36 (5): 442–450. June 2018. doi:10.1038/nbt.4103. PMID 29608178.
- ↑ "Circulating tumour cell (CTC) counts as intermediate end points in castration-resistant prostate cancer (CRPC): a single-centre experience". Annals of Oncology 20 (1): 27–33. January 2009. doi:10.1093/annonc/mdn544. PMID 18695026.
- ↑ "Single-Cell Transcriptomic Analysis of Tumor Heterogeneity" (in en). Trends in Cancer 4 (4): 264–268. April 2018. doi:10.1016/j.trecan.2018.02.003. PMID 29606308.
- ↑ "A Cancer Cell Program Promotes T Cell Exclusion and Resistance to Checkpoint Blockade" (in en). Cell 175 (4): 984–997.e24. November 2018. doi:10.1016/j.cell.2018.09.006. PMID 30388455. Bibcode: 2018Cell..175..984J.
- ↑ "Single-cell RNA-seq of rheumatoid arthritis synovial tissue using low-cost microfluidic instrumentation". Nature Communications 9 (1). February 2018. doi:10.1038/s41467-017-02659-x. PMID 29476078. Bibcode: 2018NatCo...9..791S.
- ↑ "Pathogen Cell-to-Cell Variability Drives Heterogeneity in Host Immune Responses". Cell 162 (6): 1309–21. September 2015. doi:10.1016/j.cell.2015.08.027. PMID 26343579.
- ↑ "Spatiotemporal transcriptomic maps of mouse intracerebral hemorrhage at single-cell resolution" (in en). Neuron 113 (13): 2102–2122.e7. July 2025. doi:10.1016/j.neuron.2025.04.026. PMID 40412375.
- ↑ Fromm, Amir; Hevroni, Gur; Vincent, Flora; Schatz, Daniella; Martinez-Gutierrez, Carolina A.; Aylward, Frank O.; Vardi, Assaf (2024). "Single-cell RNA-seq of the rare virosphere reveals the native hosts of giant viruses in the marine environment". Nature Microbiology 9 (6): 1619–1629. doi:10.1038/s41564-024-01669-y. PMID 38605173.
- ↑ Fromm, Amir; Shaler, Talia S.; Aylward, Frank O.; Vardi, Assaf (2025). "A single-cell perspective on host–virus dynamics in the ocean". Trends in Microbiology 33 (12): 1286–1292. doi:10.1016/j.tim.2025.05.005. PMID 40480855.
- ↑ "mRNA-Seq whole-transcriptome analysis of a single cell". Nature Methods 6 (5): 377–82. May 2009. doi:10.1038/NMETH.1315. PMID 19349980.
- ↑ "Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq". Genome Research 21 (7): 1160–7. July 2011. doi:10.1101/gr.110882.110. PMID 21543516.
- ↑ "Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells". Nature Biotechnology 30 (8): 777–82. August 2012. doi:10.1038/nbt.2282. PMID 22820318.
- ↑ "CEL-Seq: single-cell RNA-Seq by multiplexed linear amplification". Cell Reports 2 (3): 666–73. September 2012. doi:10.1016/j.celrep.2012.08.003. PMID 22939981. Bibcode: 2012CellR...2..666H.
- ↑ "High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes". Nature Communications 10 (1). 2019. doi:10.1038/s41467-019-11049-4. PMID 31311926.
- ↑ "Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity". Genome Biology 14 (4): R31. April 2013. doi:10.1186/gb-2013-14-4-r31. PMID 23594475.
- ↑ "C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution". Nature Communications 10 (1). January 2019. doi:10.1038/s41467-018-08126-5. PMID 30664627. Bibcode: 2019NatCo..10..360K.
- ↑ "How to design a single-cell RNA-sequencing experiment: pitfalls, challenges and perspectives". Briefings in Bioinformatics 20 (4): 1384–1394. 2019. doi:10.1093/bib/bby007. PMID 29394315.
- ↑ "Multiplexed quantification of proteins and transcripts in single cells". Nature Biotechnology 35 (10): 936–939. October 2017. doi:10.1038/nbt.3973. PMID 28854175.
- ↑ "Simultaneous epitope and transcriptome measurement in single cells". Nature Methods 14 (9): 865–868. September 2017. doi:10.1038/nmeth.4380. PMID 28759029. Bibcode: 2017NatCB..14..865S.
- ↑ "Comprehensive single-cell transcriptional profiling of a multicellular organism". Science 357 (6352): 661–667. August 2017. doi:10.1126/science.aam8940. PMID 28818938. Bibcode: 2017Sci...357..661C.
- ↑ "Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics". Science 360 (6391). May 2018. doi:10.1126/science.aaq1723. PMID 29674432.
- ↑ "Schmidtea mediterranea". Science 360 (6391). May 2018. doi:10.1126/science.aaq1736. PMID 29674431.
- ↑ "Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo". Science 360 (6392): 981–987. June 2018. doi:10.1126/science.aar4362. PMID 29700229. Bibcode: 2018Sci...360..981W.
- ↑ "Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis". Science 360 (6392). June 2018. doi:10.1126/science.aar3131. PMID 29700225.
- ↑ "The dynamics of gene expression in vertebrate embryogenesis at single-cell resolution". Science 360 (6392). June 2018. doi:10.1126/science.aar5780. PMID 29700227.
- ↑ "Informatics for RNA Sequencing: A Web Resource for Analysis on the Cloud". PLOS Computational Biology 11 (8). August 2015. doi:10.1371/journal.pcbi.1004393. PMID 26248053. Bibcode: 2015PLSCB..11E4393G.
- ↑ "Science's 2018 Breakthrough of the Year: tracking development cell by cell". Science Magazine. American Association for the Advancement of Science. https://vis.sciencemag.org/breakthrough2018/finalists/.
- ↑ Kharchenko, Peter V.; Silberstein, Lev; Scadden, David T. (1 July 2014). "Bayesian approach to single-cell differential expression analysis" (in en). Nature Methods 11 (7): 740–742. doi:10.1038/nmeth.2967. ISSN 1548-7091. PMID 24836921. Bibcode: 2014NatCB..11..740K.
- ↑ Svensson, Valentine; Natarajan, Kedar Nath; Ly, Lam-Ha; Miragaia, Ricardo J.; Labalette, Charlotte; Macaulay, Iain C.; Cvejic, Ana; Teichmann, Sarah A. (6 March 2017). "Power analysis of single-cell RNA-sequencing experiments" (in en). Nature Methods advance online publication (4): 381–387. doi:10.1038/nmeth.4220. ISSN 1548-7105. PMID 28263961.
- ↑ "The promise of single-cell sequencing". Nature Methods 11 (1): 25–7. January 2014. doi:10.1038/nmeth.2769. PMID 24524134. Bibcode: 2014NatCB..11...25E.
- ↑ ""Single-cell sequencing-based technologies will revolutionize whole-organism science". Nature Reviews. Genetics 14 (9): 618–30. September 2013. doi:10.1038/nrg3542. PMID 23897237."
- ↑ ""Methods, Challenges and Potentials of Single Cell RNA-seq". Biology 1 (3): 658–67. November 2012. doi:10.3390/biology1030658. PMID 24832513."
- ↑ Ament, Seth A.; Poulopoulos, Alexandros (2023). "The brain's dark transcriptome: Sequencing RNA in distal compartments of neurons and glia". Current Opinion in Neurobiology 81. doi:10.1016/j.conb.2023.102725. PMID 37196598.
- ↑ Stegle, Oliver; Teichmann, Sarah A.; Marioni, John C. (1 March 2015). "Computational and analytical challenges in single-cell transcriptomics" (in en). Nature Reviews Genetics 16 (3): 133–145. doi:10.1038/nrg3833. ISSN 1471-0056. PMID 25628217.
- ↑ Buettner, Florian; Natarajan, Kedar N.; Casale, F. Paolo; Proserpio, Valentina; Scialdone, Antonio; Theis, Fabian J.; Teichmann, Sarah A.; Marioni, John C. et al. (1 February 2015). "Computational analysis of cell-to-cell heterogeneity in single-cell RNA-sequencing data reveals hidden subpopulations of cells" (in en). Nature Biotechnology 33 (2): 155–160. doi:10.1038/nbt.3102. ISSN 1087-0156. PMID 25599176.
- ↑ Ntranos, Vasilis; Kamath, Govinda M.; Zhang, Jesse M.; Pachter, Lior; Tse, David N. (26 May 2016). "Fast and accurate single-cell RNA-seq analysis by clustering of transcript-compatibility counts". Genome Biology 17 (1): 112. doi:10.1186/s13059-016-0970-8. ISSN 1474-7596. PMID 27230763.
- ↑ Pierson, Emma; Yau, Christopher (1 January 2015). "ZIFA: Dimensionality reduction for zero-inflated single-cell gene expression analysis". Genome Biology 16. doi:10.1186/s13059-015-0805-z. ISSN 1474-760X. PMID 26527291.
- ↑ Treutlein, Barbara; Brownfield, Doug G.; Wu, Angela R.; Neff, Norma F.; Mantalas, Gary L.; Espinoza, F. Hernan; Desai, Tushar J.; Krasnow, Mark A. et al. (15 May 2014). "Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq" (in en). Nature 509 (7500): 371–375. doi:10.1038/nature13173. PMID 24739965. Bibcode: 2014Natur.509..371T.
- ↑ Korthauer, Keegan D.; Chu, Li-Fang; Newton, Michael A.; Li, Yuan; Thomson, James; Stewart, Ron; Kendziorski, Christina (1 January 2016). "A statistical approach for identifying differential distributions in single-cell RNA-seq experiments". Genome Biology 17 (1): 222. doi:10.1186/s13059-016-1077-y. ISSN 1474-760X. PMID 27782827.
- ↑ 63.0 63.1 Haghverdi, Laleh; Büttner, Maren; Wolf, F. Alexander; Buettner, Florian; Theis, Fabian J. (1 October 2016). "Diffusion pseudotime robustly reconstructs lineage branching" (in en). Nature Methods 13 (10): 845–848. doi:10.1038/nmeth.3971. ISSN 1548-7091. PMID 27571553. http://edoc.mdc-berlin.de/19027/1/19027oa.pdf.
- ↑ Saelens, Wouter; Cannoodt, Robrecht; Todorov, Helena; Saeys, Yvan (2019). "A comparison of single-cell trajectory inference methods". Nature Biotechnology 37 (5): 547–554. doi:10.1038/s41587-019-0071-9. PMID 30936559.
- ↑ Trapnell, Cole; Cacchiarelli, Davide; Grimsby, Jonna; Pokharel, Prapti; Li, Shuqiang; Morse, Michael; Lennon, Niall J.; Livak, Kenneth J. et al. (23 March 2017). "Pseudo-temporal ordering of individual cells reveals dynamics and regulators of cell fate decisions". Nature Biotechnology 32 (4): 381–386. doi:10.1038/nbt.2859. ISSN 1087-0156. PMID 24658644.
- ↑ 66.0 66.1 Pandey, Kushagra; Zafar, Hamim (2022). "Inference of cell state transitions and cell fate plasticity from single-cell with MARGARET" (in en). Nucleic Acids Research 50 (15): e86. doi:10.1093/nar/gkac412. ISSN 0305-1048. PMID 35639499.
- ↑ Wei, J.; Hu, X.; Zou, X.; Tian, T. (1 December 2016). "Inference of genetic regulatory network for stem cell using single cells expression data". 2016 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). pp. 217–222. doi:10.1109/BIBM.2016.7822521. ISBN 978-1-5090-1611-2.
- ↑ Moignard, Victoria; Macaulay, Iain C.; Swiers, Gemma; Buettner, Florian; Schütte, Judith; Calero-Nieto, Fernando J.; Kinston, Sarah; Joshi, Anagha et al. (1 April 2013). "Characterization of transcriptional networks in blood stem and progenitor cells using high-throughput single-cell gene expression analysis" (in en). Nature Cell Biology 15 (4): 363–372. doi:10.1038/ncb2709. ISSN 1465-7392. PMID 23524953.
- ↑ Hicks, Stephanie C; Townes, William F; Teng, Mingxiang; Irizarry, Rafael A (6 November 2017). "Missing data and technical variability in single-cell RNA-sequencing experiments" (in en). Biostatistics 19 (4): 562–578. doi:10.1093/biostatistics/kxx053. PMID 29121214.
- ↑ Haghverdi, Laleh; Lun, Aaron T L; Morgan, Michael D; Marioni, John C (2 April 2018). "Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors" (in en). Nature Biotechnology 36 (5): 421–427. doi:10.1038/nbt.4091. PMID 29608177. Bibcode: 2018NatBi..36..421H.
- ↑ Butler, Andrew; Hoffman, Paul; Smibert, Peter; Papalexi, Efthymia; Satija, Rahul (2 April 2018). "Integrating single-cell transcriptomic data across different conditions, technologies, and species" (in en). Nature Biotechnology 36 (5): 421–427. doi:10.1038/nbt.4096. PMID 29608179.
- ↑ Stuart, Tim; Butler, Andrew; Hoffman, Paul; Hafemeister, Christoph; Papalexia, Efthymia; Mauck, William M III; Hao, Yuhan; Marlon, Stoeckius et al. (6 June 2019). "Comprehensive Integration of Single-Cell Data" (in en). Cell 177 (7): 1888–1902. doi:10.1016/j.cell.2019.05.031. PMID 31178118.
- ↑ Shree, Ajita; Pavan, Musale Krushna; Zafar, Hamim (27 November 2023). "scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier" (in en). Nature Communications 14 (1): 7781. doi:10.1038/s41467-023-43590-8. PMID 38012145. Bibcode: 2023NatCo..14.7781S.
- ↑ Cesaro, Giulia; Nagai, James Shiniti; Gnoato, Nicolò; Chiodi, Alice; Tussardi, Gaia; Klöker, Vanessa; Musumarra, Carmelo Vittorio; Mosca, Ettore et al. (2025-05-01). "Advances and challenges in cell–cell communication inference: a comprehensive review of tools, resources, and future directions" (in en). Briefings in Bioinformatics 26 (3). doi:10.1093/bib/bbaf280. ISSN 1467-5463. PMID 40536815.
- ↑ Efremova, Mirjana; Vento-Tormo, Miquel; Teichmann, Sarah A.; Vento-Tormo, Roser (April 2020). "CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes" (in en). Nature Protocols 15 (4): 1484–1506. doi:10.1038/s41596-020-0292-x. ISSN 1750-2799. PMID 32103204. https://www.nature.com/articles/s41596-020-0292-x.
- ↑ Nagai, James S; Leimkühler, Nils B; Schaub, Michael T; Schneider, Rebekka K; Costa, Ivan G (2021-11-18). Mathelier, Anthony. ed. "CrossTalkeR: analysis and visualization of ligand–receptorne tworks" (in en). Bioinformatics 37 (22): 4263–4265. doi:10.1093/bioinformatics/btab370. ISSN 1367-4803. PMID 35032393. PMC 9502146. https://academic.oup.com/bioinformatics/article/37/22/4263/6275746.
- ↑ Dann, Emma; Henderson, Neil C.; Teichmann, Sarah A.; Morgan, Michael D.; Marioni, John C. (February 2022). "Differential abundance testing on single-cell data using k-nearest neighbor graphs" (in en). Nature Biotechnology 40 (2): 245–253. doi:10.1038/s41587-021-01033-z. ISSN 1546-1696. PMID 34594043.
- ↑ Joodaki, Mehdi; Shaigan, Mina; Parra, Victor; Bülow, Roman D; Kuppe, Christoph; Hölscher, David L; Cheng, Mingbo; Nagai, James S et al. (2023-12-19). "Detection of PatIent-Level distances from single cell genomics and pathomics data with Optimal Transport (PILOT)" (in en). Molecular Systems Biology 20 (2): 57–74. doi:10.1038/s44320-023-00003-8. ISSN 1744-4292. PMID 38177382.
- ↑ Wang, Zexuan; Zhan, Qipeng; Yang, Shu; Mu, Shizhuo; Chen, Jiong; Garai, Sumita; Orzechowski, Patryk; Wagenaar, Joost et al. (2024-11-22). "QOT: Quantized Optimal Transport for sample-level distance matrix in single-cell omics" (in en). Briefings in Bioinformatics 26 (1). doi:10.1093/bib/bbae713. ISSN 1467-5463. PMID 39808114.
- ↑ Wolf, F. Alexander; Angerer, Philipp; Theis, Fabian J. (2018-02-06). "SCANPY: large-scale single-cell gene expression data analysis". Genome Biology 19 (1): 15. doi:10.1186/s13059-017-1382-0. ISSN 1474-760X. PMID 29409532.
- ↑ Butler, Andrew; Hoffman, Paul; Smibert, Peter; Papalexi, Efthymia; Satija, Rahul (May 2018). "Integrating single-cell transcriptomic data across different conditions, technologies, and species" (in en). Nature Biotechnology 36 (5): 411–420. doi:10.1038/nbt.4096. ISSN 1546-1696. PMID 29608179.
External links
- Dissecting Tumor Heterogeneity with Single-Cell Transcriptomics
- The ultimate single-cell RNA sequencing guide by single-cell RNA sequencing service provider Single Cell Discoveries.
 |
