Biology:SurE, survival protein E
In molecular biology, the protein domain surE refers to survival protein E. It was originally found that cells that did not contain this protein, could not survive in the stationary phase, at above normal temperatures, and in high-salt media. Hence the name, survival protein E.[1] It is a metal ion-dependent phosphatase that is found in bacteria, and eukaryotes. It is an important stress response protein.[2] This domain is found in acid phosphatases (EC), 5'-nucleotidases (EC), 3'-nucleotidases (EC) and exopolyphosphatases (EC).
Interaction with pcm gene
The gene, surE, is part of a bicistronic operon found upstream of the pcm gene. When mutated, their phenotypes, or physical characteristics, are very similar and indicate that both gene products are important for survival under stressful conditions.[3]
Function
The C-terminal domain is important mainly for maintaining the oligomeric state of the protein, SurE. The N-terminal domain is thought to be part of the functional domain.[3] Since the SurE is a phosphatase enzyme it removes a phosphate group from a substance, affecting that substance's role in signal transduction.[4]
Structure
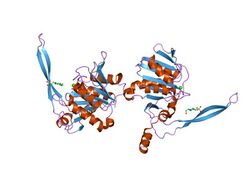
This protein consists of two protein domain. One is a large, globular N-terminal domain and the other is a smaller C-terminal domain.
N-terminal domain
The N-terminal domain contains a three-layer alpha/beta/alpha sandwich that is homologous with the Rossmann fold (CATH class 3.40.50.170) of which the major feature is a long beta sheet that is composed of nine mostly parallel beta strands.[4] SurEstructural domain has a similar topology to the N-terminal protein domain of the glutaminase/asparaginase family.[5]
C-terminal domain
The C-terminal domain, has 3 beta strands and two protrusions; one of which is a C-terminal alpha helix, and the second is a beta hairpin.[3]
References
- ↑ "A new gene involved in stationary-phase survival located at 59 minutes on the Escherichia coli chromosome.". J Bacteriol 176 (19): 6015–22. 1994. doi:10.1128/jb.176.19.6015-6022.1994. PMID 7928962.
- ↑ "Crystal structure of the stationary phase survival protein SurE with metal ion and AMP.". J Mol Biol 371 (1): 123–36. 2007. doi:10.1016/j.jmb.2007.05.007. PMID 17561111.
- ↑ Jump up to: 3.0 3.1 3.2 "Structure of Thermotoga maritima stationary phase survival protein SurE: a novel acid phosphatase.". Structure 9 (11): 1095–106. 2001. doi:10.1016/s0969-2126(01)00675-x. PMID 11709173.
- ↑ Jump up to: 4.0 4.1 "Crystal structure and functional analysis of the SurE protein identify a novel phosphatase family.". Nat Struct Biol 8 (9): 789–94. 2001. doi:10.1038/nsb0901-789. PMID 11524683.
- ↑ "Structure and function of an archaeal homolog of survival protein E (SurEalpha): an acid phosphatase with purine nucleotide specificity". J. Mol. Biol. 326 (5): 1559–75. March 2003. doi:10.1016/S0022-2836(03)00056-1. PMID 12595266.
 |