Chemistry:Fructose-asparagine
| |
| Names | |
|---|---|
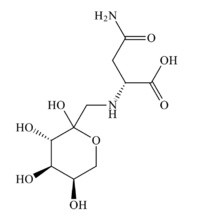
| IUPAC name
(2R)-4-amino-4-oxo-2-[ [(3S,4R,5R)-2,3,4,5-tetrahydroxyoxan-2-yl]methylamino]butanoic acid
| |
| Identifiers | |
3D model (JSmol)
|
|
PubChem CID
|
|
| |
| |
| Properties | |
| C10H18N2O8[1] | |
| Molar mass | 294.25852[1] |
| Appearance | off white crystals[1] |
| Melting point | 118 °C (244 °F; 391 K) |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| Infobox references | |
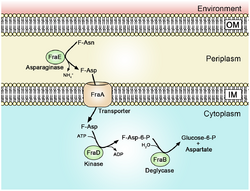
Fructose-asparagine (F-Asn) is a glycosylamine compound that is most notably used by Salmonella during Salmonella-mediated inflammation of the intestine. In addition to Salmonella, several other species of bacteria may utilize fructose-asparagine as a nutrient.[2] The name of the genetic locus that encodes the uptake capability in Salmonella is fra.[3] This fra locus has five genes: fraR (a regulator), fraB a fructose-asparagine deglycase, fraD a sugar kinase, fraA a fructose-asparagine transporter, and fraE a L-asparaginase.[3] Notably, mutations in fraB cause the buildup of the toxic intermediate 6-phosphofuctose-aspartate (6-P-F-Asp). The buildup of 6-P-F-Asp has a bacteriostatic effect on fraB mutant cells, making FraB a potential drug target.[4]
Properties
Fructose-asparagine is a aminodeoxysugar. It takes the form of a pale yellow solid. It decomposes above 120 °C.[1]
Production
Fructose-asparagine is formed when cooking food by way of the Maillard reaction. Fructose-asparagine can be prepared by refluxing D-glucose with sodium bisulfite in methanol, and then adding L-asparagine, and then acetic acid[1] or malonic acid.[3] An ion exchange resin then can separate the product from other contaminants.[1]
Fructose-asparagine is found naturally in some foods, with dried asparagus containing 1.4% and carrot containing 0.1%.[3]
F-Asn Utilization Pathway and fra Gene Locus
fraR
A regulator gene that lies upstream of the fraBDAE operon. It is predicted to encode a transcription regulator that belongs to the GntR family.[3]
fraE
fraE encodes the periplasmic fructose-asparaginase FraE that removes ammonia from F-Asn to form fructose-aspartate (F-Asp).[5] Mutations in fraE do not prevent F-Asn utilization due to redundancy conferred by the ansB gene that codes for the asparaginase AnsB.[5] It is hypothesized fraE may have resulted from a past duplication of ansB, which then underwent selection for specificity for F-Asn. FraE is specific for F-Asn and cannot function as a general periplasmic asparaginase like ansB.[5]
fraA
fraA encodes a transporter of the Dcu family, which moves F-Asp, but not F-Asn, into the cytoplasm.[5]
fraD
fraD encodes the sugar kinase FraD that phosphorylates F-Asp into 6-phosphofuctose-aspartate (6-P-F-Asp).[6] It is unusual in that it can phosphorylate both the L and D stereoisomers of F-Asp.[6]
fraB
fraB functions as a deglycase that cleaves 6-P-F-Asp into glucose-6-phosphate (G-6-P) and aspartate.[4] Mutants lacking fraD or fraA are unable to grow in media containing only F-Asn, but are able to grow in media containing F-Asn along with glucose as an alternative carbon source.[4] However, fraB mutants are unable to grow on glucose in the presence of F-Asn.[4] This is due to the accumulation of 6-P-F-Asp inside the cell. This intermediate product has been shown to be bacteriostatic to Salmonella.[4] This property makes FraB a potential target for drug treatments selective to Salmonella.[4]
Usage By Salmonella
Many gastrointestinal Salmonella serovars encode the fra locus, presumably to utilize fructose-asparagine within the inflamed gut environment. When Salmonella infects a host, it uses two Type 3 secretion systems, encoded within the genes SPI1 and SPI2, to initiate inflammation. This disrupts normal microbiota, and creates alternative electron acceptors such as nitrate and tetrathionate, and even allows oxygen to accumulate near epithelial surfaces.[7][8][9][10] Microbiota that would normally consume F-Asn are eliminated by inflammation, allowing Salmonella access to this nutrient.[11] The fra operon confers a fitness advantage to Salmonella in an inflamed intestinal tract.[3] However, this is not due to fructose-asparagine being an essential nutrient, but rather because fructose-asparagine is toxic to a fraB mutant of Salmonella,[4] Notable serovars of Salmonella enterica that do not utilize fructose-asparagine and are missing the fra operon include Typhi and Paratyphi A.[2] This is likely due to their extraintestinal niche within the host.
Usage By Other Bacteria
F-Asn is abundant in the intestinal tract of germ-free mice, but is difficult to detect in conventional mice. This suggests that there are members of the microbiota that can consume F-Asn.[11][3] It was recently discovered that several Clostridium, Citrobacter, and Klebsiella species can utilize F-Asn.[2] Notably, C. difficile cannot utilize F-Asn.[2] Salmonella is able to eliminate competition from these species by causing inflammation within the host intestinal tract. This kills other members of the microbiota that consume F-Asn and leave Salmonella the sole user of the nutrient.[11] Bioinformatic approaches suggest that Salmonella may have acquired the fra genes by horizontal transfer from the Clostridia.[2]
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 "N-(1-Deoxy-D-fructos-1-yl)-L-asparagine". Material Data Sheet. http://legacy.library.ucsf.edu/documentStore/l/j/c/ljc40d00/Sljc40d00.pdf.
- ↑ 2.0 2.1 2.2 2.3 2.4 "Identification of bacterial species that can utilize fructose-asparagine". Applied and Environmental Microbiology 84 (5): 13. 2018. doi:10.1128/AEM.01957-17. PMID 29269489. Bibcode: 2018ApEnM..84E1957S.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 "Fructose-asparagine is a primary nutrient during growth of Salmonella in the inflamed intestine". PLOS Pathogens 10 (6): e1004209. 2014. doi:10.1371/journal.ppat.1004209. PMID 24967579.
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 "A metabolic intermediate of the fructose-asparagine utilization pathway inhibits growth of a Salmonella fraB mutant". Scientific Reports 6: 28117. 2016. doi:10.1038/srep28117. PMID 27403719. Bibcode: 2016NatSR...628117S.
- ↑ 5.0 5.1 5.2 5.3 "Salmonella FraE, an asparaginase homolog, contributes to fructose-asparagine but not asparagine utilization". Journal of Bacteriology 199 (22): e00330-17. 2017. doi:10.1128/JB.00330-17. PMID 28847920.
- ↑ 6.0 6.1 "Characterization of a Salmonella sugar kinase essential for the utilization of fructose-asparagine". Biochemistry and Cell Biology 95 (2): 304–309. 2017. doi:10.1139/bcb-2016-0138. PMID 28177776.
- ↑ "Gut inflammation provides a respiratory electron acceptor for Salmonella". Nature 467 (7314): 426–429. 2010. doi:10.1038/nature09415. PMID 20864996. Bibcode: 2010Natur.467..426W.
- ↑ "Salmonella enterica serovar Typhimurium exploits inflammation to compete with the intestinal microbiota". PLOS Biology 5 (10): 2177–89. 2007. doi:10.1371/journal.pbio.0050244. PMID 17760501.
- ↑ Rivera-Chávez, Fabian; Lopez, Christopher A.; Bäumler, Andreas J. (2017). "Oxygen as a driver of gut dysbiosis". Free Radical Biology and Medicine 105: 93–101. doi:10.1016/j.freeradbiomed.2016.09.022. PMID 27677568.
- ↑ "Phage-mediated acquisition of a type III secreted effector protein boosts growth of Salmonella by nitrate respiration". mBio 3 (3): e00143-12. 2012. doi:10.1128/mBio.00143-12. PMID 22691391.
- ↑ 11.0 11.1 11.2 "Salmonella-Mediated Inflammation Eliminates Competitors for Fructose-Asparagine in the Gut". Infection and Immunity 86 (5): e00945-17. 2018. doi:10.1128/IAI.00945-17. PMID 29483291.
 |