Kruskal's algorithm
 Animation of Kruskal's algorithm on a complete graph with weights based on Euclidean distance | |
| Class | Minimum spanning tree algorithm |
|---|---|
| Data structure | Graph |
| Worst-case performance | |
| Worst-case space complexity | {{{space}}} |
Kruskal's algorithm[1] finds a minimum spanning forest of an undirected edge-weighted graph. If the graph is connected, it finds a minimum spanning tree. It is a greedy algorithm that in each step adds to the forest the lowest-weight edge that will not form a cycle.[2] The key steps of the algorithm are sorting and the use of a disjoint-set data structure to detect cycles. Its running time is dominated by the time to sort all of the graph edges by their weight.
A minimum spanning tree of a connected weighted graph is a connected subgraph, without cycles, for which the sum of the weights of all the edges in the subgraph is minimal. For a disconnected graph, a minimum spanning forest is composed of a minimum spanning tree for each connected component.
This algorithm was first published by Joseph Kruskal in 1956,[3] and was rediscovered soon afterward by (Loberman Weinberger).[4] Other algorithms for this problem include Borůvka's algorithm, Jarník's algorithm, and the reverse-delete algorithm.
Algorithm
The algorithm performs the following steps:
- Create a forest (a set of trees) initially consisting of a separate single-vertex tree for each vertex in the input graph.
- Sort the graph edges by weight.
- Loop through the edges of the graph, in ascending sorted order by their weight. For each edge:
- Test whether adding the edge to the current forest would create a cycle
- If not, add the edge to the forest, combining two trees into a single tree
At the termination of the algorithm, the forest forms a minimum spanning forest of the graph. If the graph is connected, the forest has a single component and forms a minimum spanning tree.
Pseudocode
The following code is implemented with a disjoint-set data structure. It represents the forest F as a set of undirected edges, and uses the disjoint-set data structure to efficiently determine whether two vertices are part of the same tree.
algorithm Kruskal(G) is
F:= ∅
for each v in G.V do
MAKE-SET(v)
for each {u, v} in G.E ordered by weight({u, v}), increasing do
if FIND-SET(u) ≠ FIND-SET(v) then
F := F ∪ { {u, v} }
UNION(FIND-SET(u), FIND-SET(v))
return F
Complexity
For a graph with E edges and V vertices, Kruskal's algorithm can be shown to run in time O(E log E) time, with simple data structures. Here, O expresses the time in big O notation, and log is the logarithm, which could be taken as either the natural logarithm or binary logarithm: inside O-notation, the two kinds of logarithm are equivalent, because they differ by a constant factor. This time bound is often written instead as O(E log V), which is equivalent for graphs with no isolated vertices, because for these graphs V/2 ≤ E < V2 and the logarithms of V and E are again within a constant factor of each other.
To achieve this bound, first sort the edges by weight using a comparison sort in O(E log E) time. Once sorted, it is possible to loop through the edges in sorted order in constant time per edge. Next, use a disjoint-set data structure, with a set of vertices for each component, to keep track of which vertices are in which components. Creating this structure, with a separate set for each vertex, takes V operations and O(V) time. The final iteration through all edges performs two find operations and possibly one union operation per edge. These operations take amortized time O(α(V)) time per operation, giving worst-case total time O(E α(V)) for this loop, where α is the extremely slowly growing inverse Ackermann function. This part of the time bound is much smaller than the time for the sorting step, so the total time for the algorithm can be simplified to the time for the sorting step.
In cases where the edges are already sorted, or where they have small enough integer weight to allow integer sorting algorithms such as counting sort or radix sort to sort them in linear time, the disjoint set operations are the slowest remaining part of the algorithm and the total time is O(E α(V)).
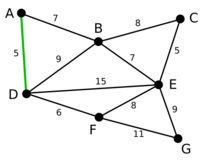
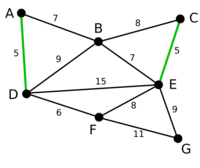
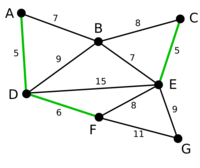
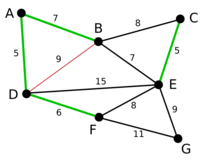
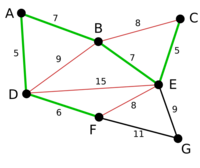
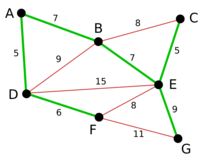
Example
Proof of correctness
The proof consists of two parts. First, it is proved that the algorithm produces a spanning tree. Second, it is proved that the constructed spanning tree is of minimal weight.
Spanning tree
Let be a connected, weighted graph and let be the subgraph of produced by the algorithm. cannot have a cycle, as by definition an edge is not added if it results in a cycle. cannot be disconnected, since the first encountered edge that joins two components of would have been added by the algorithm. Thus, is a spanning tree of .
Minimality
We show that the following proposition P is true by induction: If F is the set of edges chosen at any stage of the algorithm, then there is some minimum spanning tree that contains F and none of the edges rejected by the algorithm.
- Clearly P is true at the beginning, when F is empty: any minimum spanning tree will do, and there exists one because a weighted connected graph always has a minimum spanning tree.
- Now assume P is true for some non-final edge set F and let T be a minimum spanning tree that contains F.
- If the next chosen edge e is also in T, then P is true for F + e.
- Otherwise, if e is not in T then T + e has a cycle C. The cycle C contains edges which do not belong to F + e, since e does not form a cycle when added to F but does in T. Let f be an edge which is in C but not in F + e. Note that f also belongs to T, since f belongs to T + e but not F + e. By P, f has not been considered by the algorithm. f must therefore have a weight at least as large as e. Then T − f + e is a tree, and it has the same or less weight as T. However since T is a minimum spanning tree then T − f + e has the same weight as T, otherwise we get a contradiction and T would not be a minimum spanning tree. So T − f + e is a minimum spanning tree containing F + e and again P holds.
- Therefore, by the principle of induction, P holds when F has become a spanning tree, which is only possible if F is a minimum spanning tree itself.
Parallel algorithm
Kruskal's algorithm is inherently sequential and hard to parallelize. It is, however, possible to perform the initial sorting of the edges in parallel or, alternatively, to use a parallel implementation of a binary heap to extract the minimum-weight edge in every iteration.[5] As parallel sorting is possible in time on processors,[6] the runtime of Kruskal's algorithm can be reduced to O(E α(V)), where α again is the inverse of the single-valued Ackermann function.
A variant of Kruskal's algorithm, named Filter-Kruskal, has been described by Osipov et al.[7] and is better suited for parallelization. The basic idea behind Filter-Kruskal is to partition the edges in a similar way to quicksort and filter out edges that connect vertices of the same tree to reduce the cost of sorting. The following pseudocode demonstrates this.
function filter_kruskal(G) is
if |G.E| < kruskal_threshold:
return kruskal(G)
pivot = choose_random(G.E)
E≤, E> = partition(G.E, pivot)
A = filter_kruskal(E≤)
E> = filter(E>)
A = A ∪ filter_kruskal(E>)
return A
function partition(E, pivot) is
E≤ = ∅, E> = ∅
foreach (u, v) in E do
if weight(u, v) ≤ pivot then
E≤ = E≤ ∪ {(u, v)}
else
E> = E> ∪ {(u, v)}
return E≤, E>
function filter(E) is
Ef = ∅
foreach (u, v) in E do
if find_set(u) ≠ find_set(v) then
Ef = Ef ∪ {(u, v)}
return Ef
Filter-Kruskal lends itself better to parallelization as sorting, filtering, and partitioning can easily be performed in parallel by distributing the edges between the processors.[7]
Finally, other variants of a parallel implementation of Kruskal's algorithm have been explored. Examples include a scheme that uses helper threads to remove edges that are definitely not part of the MST in the background,[8] and a variant which runs the sequential algorithm on p subgraphs, then merges those subgraphs until only one, the final MST, remains.[9]
See also
- Prim's algorithm
- Dijkstra's algorithm
- Borůvka's algorithm
- Reverse-delete algorithm
- Single-linkage clustering
- Greedy geometric spanner
References
- ↑ Kleinberg, Jon (2006). Algorithm design. Éva Tardos. Boston: Pearson/Addison-Wesley. pp. 142–151. ISBN 0-321-29535-8. OCLC 57422612. https://www.worldcat.org/oclc/57422612.
- ↑ Cormen, Thomas; Charles E Leiserson, Ronald L Rivest, Clifford Stein (2009). Introduction To Algorithms (Third ed.). MIT Press. pp. 631. ISBN 978-0262258104. https://archive.org/details/introductiontoal00corm_805.
- ↑ Kruskal, J. B. (1956). "On the shortest spanning subtree of a graph and the traveling salesman problem". Proceedings of the American Mathematical Society 7 (1): 48–50. doi:10.1090/S0002-9939-1956-0078686-7.
- ↑ Loberman, H.; Weinberger, A. (October 1957). "Formal Procedures for connecting terminals with a minimum total wire length". Journal of the ACM 4 (4): 428–437. doi:10.1145/320893.320896.
- ↑ Quinn, Michael J.; Deo, Narsingh (1984). "Parallel graph algorithms". ACM Computing Surveys 16 (3): 319–348. doi:10.1145/2514.2515.
- ↑ Grama, Ananth; Gupta, Anshul; Karypis, George; Kumar, Vipin (2003). Introduction to Parallel Computing. pp. 412–413. ISBN 978-0201648652.
- ↑ 7.0 7.1 Osipov, Vitaly; Sanders, Peter; Singler, Johannes (2009). "The filter-kruskal minimum spanning tree algorithm". Proceedings of the Eleventh Workshop on Algorithm Engineering and Experiments (ALENEX). Society for Industrial and Applied Mathematics: 52–61. http://algo2.iti.kit.edu/documents/fkruskal.pdf.
- ↑ Katsigiannis, Anastasios; Anastopoulos, Nikos; Konstantinos, Nikas; Koziris, Nectarios (2012). "An Approach to Parallelize Kruskal's Algorithm Using Helper Threads". 2012 IEEE 26th International Parallel and Distributed Processing Symposium Workshops & PhD Forum. pp. 1601–1610. doi:10.1109/IPDPSW.2012.201. ISBN 978-1-4673-0974-5. http://tarjomefa.com/wp-content/uploads/2017/10/7793-English-TarjomeFa.pdf.
- ↑ Lončar, Vladimir; Škrbić, Srdjan; Balaž, Antun (2014). "Parallelization of Minimum Spanning Tree Algorithms Using Distributed Memory Architectures". Transactions on Engineering Technologies.: 543–554. doi:10.1007/978-94-017-8832-8_39. ISBN 978-94-017-8831-1. https://www.researchgate.net/publication/235994104.
- Thomas H. Cormen, Charles E. Leiserson, Ronald L. Rivest, and Clifford Stein. Introduction to Algorithms, Second Edition. MIT Press and McGraw-Hill, 2001. ISBN 0-262-03293-7. Section 23.2: The algorithms of Kruskal and Prim, pp. 567–574.
- Michael T. Goodrich and Roberto Tamassia. Data Structures and Algorithms in Java, Fourth Edition. John Wiley & Sons, Inc., 2006. ISBN 0-471-73884-0. Section 13.7.1: Kruskal's Algorithm, pp. 632..
External links
- Data for the article's example.
- Gephi Plugin For Calculating a Minimum Spanning Tree source code.
- Kruskal's Algorithm with example and program in c++
- Kruskal's Algorithm code in C++ as applied to random numbers
- Kruskal's Algorithm code in Python with explanation
 |