Biology:H3K56ac
H3K56ac is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the acetylation at the 56th lysine residue of the histone H3 protein.
It is a covalent modification known as a mark of newly replicated chromatin as well as replication-independent histone replacement.
H3K56ac is important for chromatin remodeling and serves as a marker of new nucleosomes during DNA replication but its role in the cell cycle is debated.
Lysine 56 is located at the amino-terminal αN-helix and close to the site where the DNA enters and exits the nucleosome. The studies on yeast might not apply to the mammals. Mammalian cells do not express HATs with high specificity to K56.
Sirtuins can catalyze the removal of the acetyl group from K56[1] H3K56ac levels are elevated in cancer and pluripotent cells.TRIM66 reads unmodified H3R2K4 and H3K56ac to respond to DNA damage.
Lysine acetylation and deacetylation
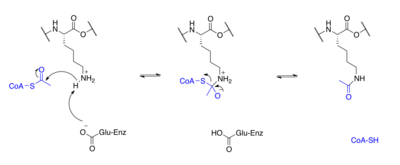
Proteins are typically acetylated on lysine residues and this reaction relies on acetyl-coenzyme A as the acetyl group donor. In histone acetylation and deacetylation, histone proteins are acetylated and deacetylated on lysine residues in the N-terminal tail as part of gene regulation. Typically, these reactions are catalyzed by enzymes with histone acetyltransferase (HAT) or histone deacetylase (HDAC) activity, although HATs and HDACs can modify the acetylation status of non-histone proteins as well.[2]
The regulation of transcription factors, effector proteins, molecular chaperones, and cytoskeletal proteins by acetylation and deacetylation is a significant post-translational regulatory mechanism[3] These regulatory mechanisms are analogous to phosphorylation and dephosphorylation by the action of kinases and phosphatases. Not only can the acetylation state of a protein modify its activity but there has been recent suggestion that this post-translational modification may also crosstalk with phosphorylation, methylation, ubiquitination, sumoylation, and others for dynamic control of cellular signaling.[4][5][6]
In the field of epigenetics, histone acetylation (and deacetylation) have been shown to be important mechanisms in the regulation of gene transcription. Histones, however, are not the only proteins regulated by posttranslational acetylation.
Nomenclature
H3K56ac indicates acetylation of lysine 56th histone H3 protein subunit: [7]
| Abbr. | Meaning |
| H3 | H3 family of histones |
| K | standard abbreviation for lysine |
| 56 | position of amino acid residue (counting from N-terminus) |
| ac | acetyl group |
Histone modifications
The genomic DNA of eukaryotic cells is wrapped around special protein molecules known as histones. The complexes formed by the looping of the DNA are known as chromatin. The basic structural unit of chromatin is the nucleosome: this consists of the core octamer of histones (H2A, H2B, H3 and H4) as well as a linker histone and about 180 base pairs of DNA. These core histones are rich in lysine and arginine residues. The carboxyl (C) terminal end of these histones contribute to histone-histone interactions, as well as histone-DNA interactions. The amino (N) terminal charged tails are the site of the post-translational modifications, such as the one seen in H3K36me3.[8][9]
Epigenetic implications
The post-translational modification of histone tails by either histone modifying complexes or chromatin remodelling complexes are interpreted by the cell and lead to complex, combinatorial transcriptional output. It is thought that a Histone code dictates the expression of genes by a complex interaction between the histones in a particular region.[10] The current understanding and interpretation of histones comes from two large scale projects: ENCODE and the Epigenomic roadmap.[11] The purpose of the epigenomic study was to investigate epigenetic changes across the entire genome. This led to chromatin states which define genomic regions by grouping the interactions of different proteins and/or histone modifications together. Chromatin states were investigated in Drosophila cells by looking at the binding location of proteins in the genome. Use of ChIP-sequencing revealed regions in the genome characterised by different banding.[12] Different developmental stages were profiled in Drosophila as well, an emphasis was placed on histone modification relevance.[13] A look in to the data obtained led to the definition of chromatin states based on histone modifications.[14]
The human genome was annotated with chromatin states. These annotated states can be used as new ways to annotate a genome independently of the underlying genome sequence. This independence from the DNA sequence enforces the epigenetic nature of histone modifications. Chromatin states are also useful in identifying regulatory elements that have no defined sequence, such as enhancers. This additional level of annotation allows for a deeper understanding of cell specific gene regulation.[15]
H3K56ac
H3K56ac is a covalent modification known as a mark of newly replicated chromatin as well as replication-independent histone replacement.[16]
H3K56ac is important for chromatin remodeling and serves as a marker of new nucleosomes during DNA replication but its role in the cell cycle is debated.[1]
Lysine 56 is located at the amino-terminal αN-helix and close to the site where the DNA enters and exits the nucleosome.[1] due to its location on the lateral surface of the nucleosome, which is close to the DNA entry/exit site and interacts with DNA29.[17]
The studies on yeast might not apply to the mammals. Mammalian cells do not express HATs with high specificity to K56.[1]
Sirtuins can catalyze the removal of the acetyl group from K56[1] H3K56ac levels are elevated in cancer and pluripotent cells[1] TRIM66 reads unmodified H3R2K4 and H3K56ac to respond to DNA damage.[1]
H3T45P promotes H3K56 acetylation.[18] Phosphorylation of the nucleosome DNA entry-exit region improves access to DNA binding complexes, and the combination of phosphorylation and acetylation has the ability to alter DNA accessibility to transcription regulatory complexes dramatically.[19]
Methods
The histone mark acetylation can be detected in a variety of ways:
1. Chromatin Immunoprecipitation Sequencing (ChIP-sequencing) measures the amount of DNA enrichment once bound to a targeted protein and immunoprecipitated. It results in good optimization and is used in vivo to reveal DNA-protein binding occurring in cells. ChIP-Seq can be used to identify and quantify various DNA fragments for different histone modifications along a genomic region.[20]
2. Micrococcal Nuclease sequencing (MNase-seq) is used to investigate regions that are bound by well positioned nucleosomes. Use of the micrococcal nuclease enzyme is employed to identify nucleosome positioning. Well positioned nucleosomes are seen to have enrichment of sequences.[21]
3. Assay for transposase accessible chromatin sequencing (ATAC-seq) is used to look in to regions that are nucleosome free (open chromatin). It uses hyperactive Tn5 transposon to highlight nucleosome localisation.[22][23][24]
See also
- Histone acetylation
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Stejskal, Stanislav; Stepka, Karel; Tesarova, Lenka; Stejskal, Karel; Matejkova, Martina; Simara, Pavel; Zdrahal, Zbynek; Koutna, Irena (2015). "Cell cycle-dependent changes in H3K56ac in human cells". Cell Cycle 14 (24): 3851–3863. doi:10.1080/15384101.2015.1106760. PMID 26645646.
- ↑ "Regulation of protein turnover by acetyltransferases and deacetylases". Biochimie 90 (2): 306–12. 2008. doi:10.1016/j.biochi.2007.06.009. PMID 17681659.
- ↑ "Acetylation and deacetylation of non-histone proteins". Gene 363: 15–23. 2005. doi:10.1016/j.gene.2005.09.010. PMID 16289629.
- ↑ "Lysine acetylation: codified crosstalk with other posttranslational modifications". Mol. Cell 31 (4): 449–61. 2008. doi:10.1016/j.molcel.2008.07.002. PMID 18722172.
- ↑ "Posttranslational modifications of tubulin in cultured mouse brain neurons and astroglia". Biol. Cell 65 (2): 109–117. 1989. doi:10.1016/0248-4900(89)90018-x. PMID 2736326.
- ↑ "The acetylation of alpha-tubulin and its relationship to the assembly and disassembly of microtubules". J. Cell Biol. 103 (2): 571–579. 1986. doi:10.1083/jcb.103.2.571. PMID 3733880.
- ↑ Huang, Suming; Litt, Michael D.; Ann Blakey, C. (2015-11-30). Epigenetic Gene Expression and Regulation. Elsevier Science. pp. 21–38. ISBN 9780127999586.
- ↑ "Multivalent engagement of chromatin modifications by linked binding modules". Nature Reviews. Molecular Cell Biology 8 (12): 983–94. December 2007. doi:10.1038/nrm2298. PMID 18037899.
- ↑ "Chromatin modifications and their function". Cell 128 (4): 693–705. February 2007. doi:10.1016/j.cell.2007.02.005. PMID 17320507.
- ↑ "Translating the histone code". Science 293 (5532): 1074–80. August 2001. doi:10.1126/science.1063127. PMID 11498575.
- ↑ "Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project". Nature 447 (7146): 799–816. June 2007. doi:10.1038/nature05874. PMID 17571346. Bibcode: 2007Natur.447..799B.
- ↑ "Systematic protein location mapping reveals five principal chromatin types in Drosophila cells". Cell 143 (2): 212–24. October 2010. doi:10.1016/j.cell.2010.09.009. PMID 20888037.
- ↑ "Identification of functional elements and regulatory circuits by Drosophila modENCODE". Science 330 (6012): 1787–97. December 2010. doi:10.1126/science.1198374. PMID 21177974. Bibcode: 2010Sci...330.1787R.
- ↑ "Comprehensive analysis of the chromatin landscape in Drosophila melanogaster". Nature 471 (7339): 480–5. March 2011. doi:10.1038/nature09725. PMID 21179089. Bibcode: 2011Natur.471..480K.
- ↑ "Integrative analysis of 111 reference human epigenomes". Nature 518 (7539): 317–30. February 2015. doi:10.1038/nature14248. PMID 25693563. Bibcode: 2015Natur.518..317..
- ↑ Kaplan, Tommy; Liu, Chih Long; Erkmann, Judith A.; Holik, John; Grunstein, Michael; Kaufman, Paul D.; Friedman, Nir; Rando, Oliver J. (2008). "Cell Cycle– and Chaperone-Mediated Regulation of H3K56ac Incorporation in Yeast". PLOS Genetics 4 (11): e1000270. doi:10.1371/journal.pgen.1000270. PMID 19023413.
- ↑ Chen, Jiajing; Wang, Zikang; Guo, Xudong; Li, Fudong; Wei, Qingtao; Chen, Xuwen; Gong, Deshun; Xu, Yanxin et al. (2019). "TRIM66 reads unmodified H3R2K4 and H3K56ac to respond to DNA damage in embryonic stem cells". Nature Communications 10 (1): 4273. doi:10.1038/s41467-019-12126-4. PMID 31537782. Bibcode: 2019NatCo..10.4273C.
- ↑ Darieva, Zoulfia; Webber, Aaron; Warwood, Stacey; Sharrocks, Andrew D. (2015). "Protein kinase C coordinates histone H3 phosphorylation and acetylation". eLife 4: e09886. doi:10.7554/eLife.09886. PMID 26468616.
- ↑ Brehove, Matthew; Wang, Tao; North, Justin; Luo, Yi; Dreher, Sarah J.; Shimko, John C.; Ottesen, Jennifer J.; Luger, Karolin et al. (2015). "Histone Core Phosphorylation Regulates DNA Accessibility". Journal of Biological Chemistry 290 (37): 22612–22621. doi:10.1074/jbc.M115.661363. PMID 26175159.
- ↑ "Whole-Genome Chromatin IP Sequencing (ChIP-Seq)". https://www.illumina.com/Documents/products/datasheets/datasheet_chip_sequence.pdf. Retrieved 23 October 2019.
- ↑ "MAINE-Seq/Mnase-Seq". https://www.illumina.com/science/sequencing-method-explorer/kits-and-arrays/maine-seq-mnase-seq-nucleo-seq.html?langsel=/us/. Retrieved 23 October 2019.
- ↑ Buenrostro, Jason D.; Wu, Beijing; Chang, Howard Y.; Greenleaf, William J. (2015). "ATAC-seq: A Method for Assaying Chromatin Accessibility Genome-Wide". Current Protocols in Molecular Biology 109: 21.29.1–21.29.9. doi:10.1002/0471142727.mb2129s109. ISBN 9780471142720. PMID 25559105.
- ↑ Schep, Alicia N.; Buenrostro, Jason D.; Denny, Sarah K.; Schwartz, Katja; Sherlock, Gavin; Greenleaf, William J. (2015). "Structured nucleosome fingerprints enable high-resolution mapping of chromatin architecture within regulatory regions". Genome Research 25 (11): 1757–1770. doi:10.1101/gr.192294.115. ISSN 1088-9051. PMID 26314830.
- ↑ Song, L.; Crawford, G. E. (2010). "DNase-seq: A High-Resolution Technique for Mapping Active Gene Regulatory Elements across the Genome from Mammalian Cells". Cold Spring Harbor Protocols 2010 (2): pdb.prot5384. doi:10.1101/pdb.prot5384. ISSN 1559-6095. PMID 20150147.
 |