Biology:Lac repressor
The lac repressor (LacI) is a DNA-binding protein that inhibits the expression of genes coding for proteins involved in the metabolism of lactose in bacteria. These genes are repressed when lactose is not available to the cell, ensuring that the bacterium only invests energy in the production of machinery necessary for uptake and utilization of lactose when lactose is present. When lactose becomes available, it is firstly converted into allolactose by β-Galactosidase (lacZ) in bacteria. The DNA binding ability of lac repressor bound with allolactose is inhibited due to allosteric regulation, thereby genes coding for proteins involved in lactose uptake and utilization can be expressed.
Function
The lac repressor (LacI) operates by a helix-turn-helix motif in its DNA-binding domain, binding base-specifically to the major groove of the operator region of the lac operon, with base contacts also made by residues of symmetry-related alpha helices, the "hinge" helices, which bind deeply in the minor groove.[1] This bound repressor can reduce transcription of the Lac proteins by occluding the RNA polymerase binding site or by prompting DNA looping.[2] When lactose is present, allolactose binds to the lac repressor, causing an allosteric change in its shape. In its changed state, the lac repressor is unable to bind tightly to its cognate operator. Thus, the gene is mostly off in the absence of inducer and mostly on in the presence of inducer, although the degree of gene expression depends on the number of repressors in the cell and on the repressor's DNA-binding affinity.[3] Isopropyl β-D-1-thiogalactopyranoside (IPTG) is a commonly used allolactose mimic which can be used to induce transcription of genes being regulated by lac repressor.
Structure
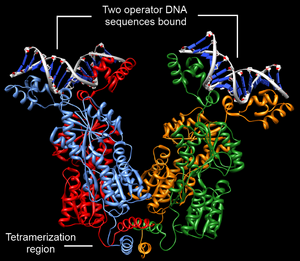
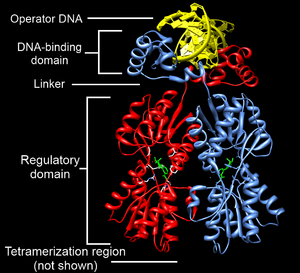
Structurally, the lac repressor protein is a homotetramer. More precisely, the tetramer contains two DNA-binding subunits composed of two monomers each (a dimer of dimers). Each monomer consists of four distinct regions:[4][5][6]
- An N-terminal DNA-binding domain (in which two LacI proteins bind a single operator site)
- A regulatory domain (sometimes called the core domain, which binds allolactose, an allosteric effector molecule)
- A linker that connects the DNA-binding domain with the core domain (sometimes called the hinge helix, which is important for allosteric communication[6])
- A C-terminal tetramerization region (which joins four monomers in an alpha-helix bundle)
DNA binding occurs via an N-terminal helix-turn-helix structural motif and is targeted to one of several operator DNA sequences (known as O1, O2 and O3). The O1 operator sequence slightly overlaps with the promoter, which increases the affinity of RNA polymerase for the promoter sequence such that it cannot enter elongation and remains in abortive initiation. Additionally, because each tetramer contains two DNA-binding subunits, binding of multiple operator sequences by a single tetramer induces DNA looping.[7]
Each monomer has 360 amino acids, so it has 1440 amino acids in total, and 154,520 Dalton of atomic mass.[8]
Kinetics of DNA binding and unbinding
File:Binding and unbinding mechanism of LacI.webm LacI finds its target operator DNA surprisingly fast. In vitro the search is 10-100 times faster than the theoretical upper limit for two particles searching for each other via diffusion in three dimensions (3D).[9] To explain the fast search, it was hypothesized that LacI and other transcription factors (TFs) find their binding sites by facilitated diffusion, a combination of free diffusion in 3D and 1D-sliding on the DNA.[10] During sliding the repressor is in contact with the DNA helix, sliding around and tracking its major groove, which speeds up the search process by extending the target length when the TF slides in onto the operator from the side. In vivo single-molecule experiments with E. coli cells have now tested and verified the facilitated diffusion model, and shown that the TF scans on average 45 bp during each sliding event, before the TF detaches spontaneously and resumes exploring the genome in 3D.[11] These experiments also suggest that LacI slides over the O1 operator several times before binding, meaning that different DNA sequences can have different probabilities to be recognized at each encounter with the TF. This implies a trade-off between fast search on nonspecific sequences and binding to specific sequences.[11] In vivo and in vitro experiments have shown that it is this probability to recognize the operator that changes with DNA sequence, while the time the TF remains in the bound conformation on the operator changes less with sequence.[12] The TF often leaves the sequence it is intended to regulate, but at a strong target site, it almost always make a very short journey before finding the way back again. On the macroscopic scale, this looks like a stable interaction. This binding mechanism explains how DNA binding proteins manage to quickly search through the genome of the cell without getting stuck too long at sequences that resemble the true target.
An all-atom molecular dynamics simulation suggests that the transcription factor encounters a barrier of 1 kBT during sliding and 12 kBT for dissociation, implying that the repressor will slide over 8 bp on average before dissociating.[13] The in vivo search model for the lac repressor includes intersegment transfer and hopping as well as crowding by other proteins which make the genome in E. coli cells less accessible for the repressor.[14] The existence of hopping, where the protein slips out of the major groove of DNA to land in another nearby groove along the DNA chain, has been proven more directly in vitro, where the lac repressor has been observed to bypass operators, flip orientation, and rotate with a longer pitch than the 10.5 bp period of DNA while moving along it.[15]
Discovery
The lac repressor was first isolated by Walter Gilbert and Benno Müller-Hill in 1966.[16] They showed that in vitro the protein bound to DNA containing the lac operon, and it released the DNA when IPTG (an analog of allolactose) was added.
See also
References
- ↑ "Crystal structure of LacI member, PurR, bound to DNA: minor groove binding by alpha helices". Science 266 (5186): 763–70. November 1994. doi:10.1126/science.7973627. PMID 7973627. Bibcode: 1994Sci...266..763S.
- ↑ "Comparison of the theoretical and real-world evolutionary potential of a genetic circuit". Physical Biology 1 (2). 2014. doi:10.1088/1478-3975/11/2/026005. PMID 24685590. Bibcode: 2014PhBio..11b6005R.
- ↑ "Tuning Transcriptional Regulation through Signaling: A Predictive Theory of Allosteric Induction". Cell Systems 6 (4): 456–469. 2018. doi:10.1016/j.cels.2018.02.004. PMID 29574055.
- ↑ "Lac Repressor". RCSB Protein Data Bank. 2003. doi:10.2210/rcsb_pdb/mom_2003_3.
- ↑ "The lac repressor". Comptes Rendus Biologies 328 (6): 521–48. June 2005. doi:10.1016/j.crvi.2005.04.004. PMID 15950160.
- ↑ 6.0 6.1 "Allostery in the LacI/GalR family: variations on a theme". Current Opinion in Microbiology 12 (2): 129–37. April 2009. doi:10.1016/j.mib.2009.01.009. PMID 19269243.
- ↑ "The three operators of the lac operon cooperate in repression". The EMBO Journal 9 (4): 973–9. April 1990. doi:10.1002/j.1460-2075.1990.tb08199.x. PMID 2182324.
- ↑ Lewis, Mitchell (2005-06-01). "The lac repressor". Comptes Rendus Biologies. Retour sur l'operon lac 328 (6): 521–548. doi:10.1016/j.crvi.2005.04.004. ISSN 1631-0691. PMID 15950160.
- ↑ Riggs, Arthur D.; Bourgeois, Suzanne; Cohn, Melvin (1970). "The lac repressor-operator interaction". Journal of Molecular Biology (Elsevier BV) 53 (3): 401–417. doi:10.1016/0022-2836(70)90074-4. ISSN 0022-2836. PMID 4924006.
- ↑ Berg, Otto G.; Winter, Robert B.; Von Hippel, Peter H. (1981-11-01). "Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory". Biochemistry (American Chemical Society (ACS)) 20 (24): 6929–6948. doi:10.1021/bi00527a028. ISSN 0006-2960. PMID 7317363.
- ↑ 11.0 11.1 Hammar, Petter; Leroy, Prune; Mahmutovic, Anel; Marklund, Erik G.; Berg, Otto G.; Elf, Johan (2012-06-22). "The lac Repressor Displays Facilitated Diffusion in Living Cells" (in en). Science 336 (6088): 1595–1598. doi:10.1126/science.1221648. ISSN 0036-8075. PMID 22723426. Bibcode: 2012Sci...336.1595H.
- ↑ Marklund, Emil; Mao, Guanzhong; Yuan, Jinwen; Zikrin, Spartak; Abdurakhmanov, Eldar; Deindl, Sebastian; Elf, Johan (2022-01-28). "Sequence specificity in DNA binding is mainly governed by association". Science (American Association for the Advancement of Science (AAAS)) 375 (6579): 442–445. doi:10.1126/science.abg7427. ISSN 0036-8075. PMID 35084952. Bibcode: 2022Sci...375..442M. http://urn.kb.se/resolve?urn=urn:nbn:se:uu:diva-466865.
- ↑ Marklund, Erik G.; Mahmutovic, Anel; Berg, Otto G.; Hammar, Petter; Spoel, David van der; Fange, David; Elf, Johan (2013-12-03). "Transcription-factor binding and sliding on DNA studied using micro- and macroscopic models" (in en). Proceedings of the National Academy of Sciences 110 (49): 19796–19801. doi:10.1073/pnas.1307905110. ISSN 0027-8424. PMID 24222688. Bibcode: 2013PNAS..11019796M.
- ↑ Mahmutovic, Anel; Berg, Otto G.; Elf, Johan (2015-03-16). "What matters for lac repressor search in vivo—sliding, hopping, intersegment transfer, crowding on DNA or recognition?" (in en). Nucleic Acids Research 43 (7): 3454–3464. doi:10.1093/nar/gkv207. ISSN 1362-4962. PMID 25779051.
- ↑ Marklund, Emil; van Oosten, Brad; Mao, Guanzhong; Amselem, Elias; Kipper, Kalle; Sabantsev, Anton; Emmerich, Andrew; Globisch, Daniel et al. (2020). "DNA surface exploration and operator bypassing during target search". Nature 583 (7818): 858–861. doi:10.1038/s41586-020-2413-7. ISSN 0028-0836. PMID 32581356. Bibcode: 2020Natur.583..858M. http://urn.kb.se/resolve?urn=urn:nbn:se:uu:diva-439327.
- ↑ "Isolation of the lac repressor". Proceedings of the National Academy of Sciences of the United States of America 56 (6): 1891–8. December 1966. doi:10.1073/pnas.56.6.1891. PMID 16591435. Bibcode: 1966PNAS...56.1891G.
External links
- Lac Repressor at the US National Library of Medicine Medical Subject Headings (MeSH)
- More information on the lac repressor molecule on protein database
- Lac Repressor in Proteopedia.
 |
