Biology:mRNA surveillance
mRNA surveillance mechanisms are pathways utilized by organisms to ensure fidelity and quality of messenger RNA (mRNA) molecules. There are a number of surveillance mechanisms present within cells. These mechanisms function at various steps of the mRNA biogenesis pathway to detect and degrade transcripts that have not properly been processed.
Overview
The translation of messenger RNA transcripts into proteins is a vital part of the central dogma of molecular biology. mRNA molecules are, however, prone to a host of fidelity errors which can cause errors in translation of mRNA into quality proteins.[1] RNA surveillance mechanisms are methods cells use to assure the quality and fidelity of the mRNA molecules.[2] This is generally achieved through marking aberrant mRNA molecule for degradation by various endogenous nucleases.[3]
mRNA surveillance has been documented in bacteria and yeast. In eukaryotes, these mechanisms are known to function in both the nucleus and cytoplasm.[4] Fidelity checks of mRNA molecules in the nucleus results in the degradation of improperly processed transcripts before export into the cytoplasm. Transcripts are subject to further surveillance once in the cytoplasm. Cytoplasmic surveillance mechanisms assess mRNA transcripts for the absence of or presence of premature stop codons.[3][4]
Three surveillance mechanisms are currently known to function within cells: the nonsense-mediated mRNA decay pathway (NMD); the nonstop mediated mRNA decay pathways (NSD); and the no-go mediated mRNA decay pathway (NGD).
Nonsense-mediated mRNA decay
Overview
Nonsense-mediated decay is involved in detection and decay of mRNA transcripts which contain premature termination codons (PTCs). PTCs can arise in cells through various mechanisms: germline mutations in DNA; somatic mutations in DNA; errors in transcription; or errors in post transcriptional mRNA processing.[5][6] Failure to recognize and decay these mRNA transcripts can result in the production of truncated proteins which may be harmful to the organism. By causing decay of C-terminally truncated polypeptides, the NMD mechanism can protect cells against deleterious dominant-negative, and gain of function effects.[7] PTCs have been implicated in approximately 30% of all inherited diseases; as such, the NMD pathway plays a vital role in assuring overall survival and fitness of an organism.[8][9]
A surveillance complex consisting of various proteins (eRF1, eRF3, Upf1, Upf2 and Upf3) is assembled and scans the mRNA for premature stop codons.[5] The assembly of this complex is triggered by premature translation termination. If a premature stop codon is detected then the mRNA transcript is signalled for degradation – the coupling of detection with degradation occurs.[3][10][11]
Seven smg genes (smg1-7) and three UPF genes (Upf1-3) have been identified in Saccharomyces cerevisiae and Caenorhabditis elegans as essential trans-acting factors contributing to NMD activity.[12][13] All of these genes are conserved in Drosophila melanogaster and further mammals where they also play critical roles in NMD. Throughout eukaryotes there are three components which are conserved in the process of NMD.[14] These are the Upf1/SMG-2, Upf2/SMG-3 and Upf3/SMG-4 complexes. Upf1/SMG-2 is a phosphoprotein in multicellular organisms and is thought to contribute to NMD via its phosphorylation activity. However, the exact interactions of the proteins and their roles in NMD are currently disputed.[11][12][14][15][16]
Mechanism in mammals
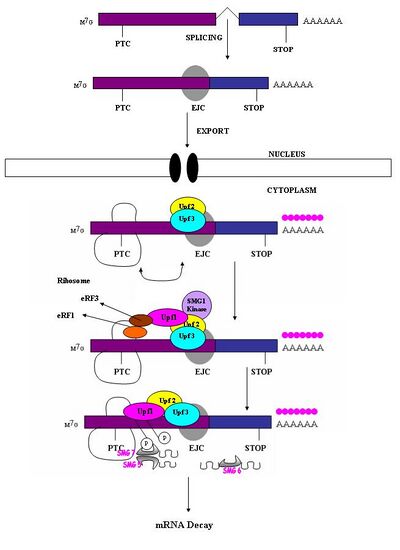
A premature stop codon must be recognized as different from a normal stop codon so that only the former triggers a NMD response. It has been observed that the ability of a nonsense codon to cause mRNA degradation depends on its relative location to the downstream sequence element and associated proteins.[1] Studies have demonstrated that nucleotides more than 50–54 nucleotides upstream of the last exon-exon junction can target mRNA for decay.[1][4][5][6][7][17] Those downstream from this region are unable to do so. Thus, nonsense codons lie more than 50-54 nucleotides upstream from the last exon boundary whereas natural stop codons are located within terminal exons.[18] Exon junction complexes (EJCs) mark the exon-exon boundaries. EJCs are multiprotein complexes that assemble during splicing at a position about 20–24 nucleotides upstream from the splice junction.[19] It is this EJC that provides position information needed to discriminate premature stop codons from natural stop codons. Recognition of PTCs appears to be dependent on the definitions of the exon-exon junctions. This suggests involvement of the spliceosome in mammalian NMD.[17][20] Research has investigated the possibility of spliceosome involvement in mammalian NMD and has determined this is a likely possibility.[18] Furthermore, it has been observed that NMD mechanisms are not activated by nonsense transcripts that are generated from genes that naturally do not contain introns (e.g., Histone H4, Hsp70, melanocortin-4-receptor).[7]
When the ribosome reaches a PTC the translation factors eRF1 and eRF3 interact with retained EJC complexes though a multiprotein bridge.[21] The interactions of UPF1 with the terminating complex and with UPF2/UPF3 of the retained EJCs are critical. It is these interactions which target the mRNA for rapid decay by endogenous nucleases[18][21]

Mechanism in invertebrates
Studies involving organisms such as S. cerevisiae, D.melanogaster and C. elegans have shown that PTC recognition involving invertebrate organisms does not involve exon-exon boundaries.[20] These studies suggest that invertebrate NMD occurs independently of splicing. As a result, EJCs which are responsible for marking exon-exon boundaries are not required in invertebrate NMD.[3] Several models have been proposed to explain how PTCs are distinguished from normal stop codons in invertebrates. One of these suggests that there may be a downstream sequence element which functions similar to the exon junctions in mammals.[11] A second model proposes that a widely present feature in mRNA, such as a 3' poly-A tail, might provide the positional information required for recognition.[22] Another model, dubbed the "faux 3'UTR model", suggests that premature translation termination may be distinguished from normal termination because of intrinsic features that may allow it to recognize its presence in an inappropriate environment.[3] These mechanisms, however, have yet to be conclusively demonstrated.
Mechanism in plants
There are two mechanisms of PTC recognition in plants: according to its distance from the EJC (like in vertebrates) or from the poly-A tail. The NMD mechanism in plants induces the decay of mRNAs containing a 3’UTR longer than 300 nucleotides, that is why the proportion of mRNAs with longer 3'UTRs is much lower in plants than in vertebrates.[23][24]
NMD avoidance
mRNAs with nonsense mutations are generally thought to be targeted for decay via the NMD pathways. The presence of this premature stop codon about 50-54 nucleotides 5' to the exon junction appears to be the trigger for rapid decay; however, it has been observed that some mRNA molecules with a premature stop codon are able to avoid detection and decay.[17][25] In general, these mRNA molecules possess the stop codon very early in the reading frame (i.e. the PTC is AUG-proximal). This appears to be a contradiction to the current accepted model of NMD as this position is significantly 5' of the exon-exon junction.[26]
This has been demonstrated in β-globulin. β-globulin mRNAs containing a nonsense mutation early in the first exon of the gene are more stable than NMD sensitive mRNA molecules. The exact mechanism of detection avoidance is currently not known. It has been suggested that the poly-A binding protein (PABP) appears to play a role in this stability.[27] It has been demonstrated in other studies that the presence of this protein near AUG-proximal PTCs appears to promote the stability of these otherwise NMD sensitive mRNAs. It has been observed that this protective effect is not limited only to the β-globulin promoter.[25] This suggests that this NMD avoidance mechanism may be prevalent in other tissue types for a variety of genes. The current model of NMD may need to be revisited upon further studies.
Nonstop mediated mRNA decay
Overview
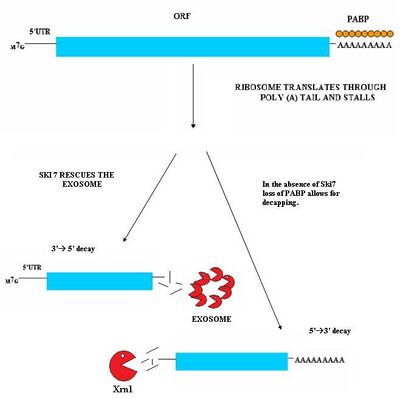
Nonstop mediated decay (NSD) is involved in the detection and decay of mRNA transcripts which lack a stop codon.[29][30] These mRNA transcripts can arise from many different mechanisms such as premature 3' adenylation or cryptic polyadenylation signals within the coding region of a gene.[31] This lack of a stop codon results a significant issue for cells. Ribosomes translating the mRNA eventually translate into the 3'poly-A tail region of transcripts and stalls. As a result, it cannot eject the mRNA.[32] Ribosomes thus may become sequestered associated with the nonstop mRNA and would not be available to translate other mRNA molecules into proteins. Nonstop mediated decay resolves this problem by both freeing the stalled ribosomes and marking the nonstop mRNA for degradation in the cell by nucleases. Nonstop mediated decay consists of two distinct pathways which likely act in concert to decay nonstop mRNA.[29][30]
Ski7 pathway
This pathway is active when Ski7 protein is available in the cell. The Ski7 protein is thought to bind to the empty A site of the ribosome. This binding allows the ribosome to eject the stuck nonstop mRNA molecule – this even frees the ribosome and allows it to translate other transcripts. The Ski7 is now associated with the nonstop mRNA and it is this association which targets the nonstop mRNA for recognition by the cytosolic exosome. The Ski7-exosome complex rapidly deadenylates the mRNA molecule which allows the exosome to decay the transcript in a 3' to 5' fashion.[29][30]
Non-Ski7 pathway
A second type of NSD has been observed in yeast. In this mechanism, the absence of Ski7 results in the loss of poly-A tail binding PABP proteins by the action of the translation ribosome. The removal of these PABP proteins then results in the loss of the protective 5'm7G cap. The loss of the cap results in rapid degradation of the transcript by an endogenous 5'-3' exonuclease such as XrnI.[30]
No-Go decay
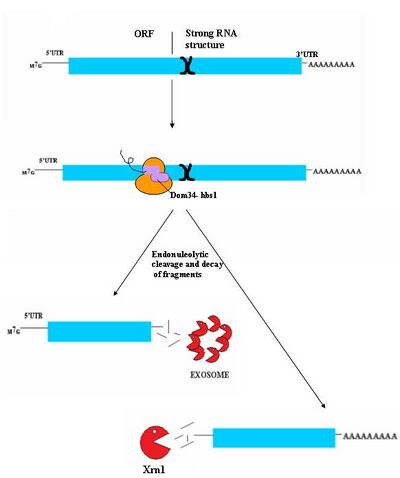
No-Go decay (NGD) is the most recently discovered surveillance mechanism.[33] As such, it is not currently well understood. While authentic targets of NGD are poorly understood, they appear to consist largely of mRNA transcripts on which ribosomes have stalled during translation. This stall can be caused by a variety of factors including strong secondary structures, which may physically block the translational machinery from moving down the transcript.[33] Dom34/Hbs1 likely binds near the A site of stalled ribosomes and may facilitate recycling of complexes.[34] In some cases, the transcript is also cleaved in an endonucleolytic fashion near the stall site; however the identity of the responsible endonuclease remains contentious. The fragmented mRNA molecules are then fully degraded by the exosome in a 3' to 5' fashion and by Xrn1 in a 5' to 3' fashion.[33] It is not currently known how this process releases the mRNA from the ribosomes, however, Hbs1 is closely related to the Ski7 protein which plays a clear role in ribosome release in Ski7 mediated NSD. It is postulated that Hbs1 may play a similar role in NGD.[5][35]
Evolution
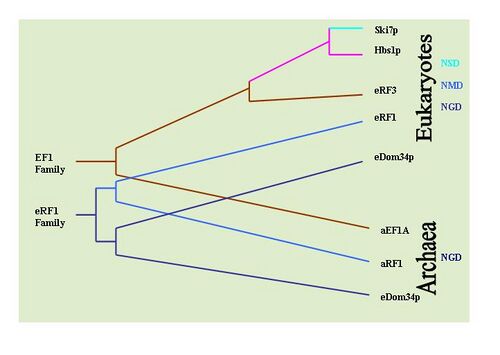
It is possible to determine the evolutionary history of these mechanisms by observing the conservation of key proteins implicated in each mechanism. For example: Dom34/Hbs1 are associated with NGD;[33] Ski7 is associated with NSD;[29] and the eRF proteins are associated with NMD.[6] To this end, extensive BLAST searches have been performed to determine the prevalence of the proteins in various types of organisms. It has been determined that NGD Hbs1 and NMD eRF3 are found only in eukaryotes. However, the NGD Dom34 is universal in eukaryotes and archaea. This suggests that NGD appears to have been the first evolved mRNA surveillance mechanism. The NSD Ski7 protein appears to be restricted strictly to yeast species which suggest that NSD is the most recently evolved surveillance mechanism. This by default leaves NMD as the second evolved surveillance mechanism.[36]
References
- ↑ 1.0 1.1 1.2 "Early nonsense: mRNA decay solves a translational problem". Nature Reviews. Molecular Cell Biology 7 (6): 415–25. June 2006. doi:10.1038/nrm1942. PMID 16723977.
- ↑ "From birth to death: the complex lives of eukaryotic mRNAs". Science 309 (5740): 1514–8. September 2005. doi:10.1126/science.1111443. PMID 16141059. Bibcode: 2005Sci...309.1514M.
- ↑ 3.0 3.1 3.2 3.3 3.4 "A faux 3'-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay". Nature 432 (7013): 112–8. November 2004. doi:10.1038/nature03060. PMID 15525991. Bibcode: 2004Natur.432..112A.
- ↑ 4.0 4.1 4.2 "Process or perish: quality control in mRNA biogenesis". Nature Structural & Molecular Biology 12 (6): 482–8. June 2005. doi:10.1038/nsmb945. PMID 15933735.
- ↑ 5.0 5.1 5.2 5.3 5.4 5.5 5.6 "The nonsense-mediated decay RNA surveillance pathway". Annual Review of Biochemistry 76: 51–74. 2007. doi:10.1146/annurev.biochem.76.050106.093909. PMID 17352659.
- ↑ 6.0 6.1 6.2 "Nonsense-mediated mRNA decay: Target genes and functional diversification of effectors". Trends in Biochemical Sciences 31 (11): 639–46. November 2006. doi:10.1016/j.tibs.2006.09.005. PMID 17010613.
- ↑ 7.0 7.1 7.2 "Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics". Nature Reviews. Molecular Cell Biology 5 (2): 89–99. February 2004. doi:10.1038/nrm1310. PMID 15040442.
- ↑ "Nonsense-mediated decay approaches the clinic". Nature Genetics 36 (8): 801–8. August 2004. doi:10.1038/ng1403. PMID 15284851.
- ↑ "Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise". Nature Genetics 36 (10): 1073–8. October 2004. doi:10.1038/ng1429. PMID 15448691.
- ↑ "Mechanistic links between nonsense-mediated mRNA decay and pre-mRNA splicing in mammalian cells". Current Opinion in Cell Biology 17 (3): 309–15. June 2005. doi:10.1016/j.ceb.2005.03.002. PMID 15901502.
- ↑ 11.0 11.1 11.2 "Nonsense-mediated mRNA decay: molecular insights and mechanistic variations across species". Current Opinion in Cell Biology 17 (3): 316–25. June 2005. doi:10.1016/j.ceb.2005.04.005. PMID 15901503.
- ↑ 12.0 12.1 "smg-7 is required for mRNA surveillance in Caenorhabditis elegans". Genetics 151 (2): 605–16. February 1999. doi:10.1093/genetics/151.2.605. PMID 9927455.
- ↑ "The role of SMG-1 in nonsense-mediated mRNA decay". Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 1754 (1–2): 305–15. December 2005. doi:10.1016/j.bbapap.2005.10.002. PMID 16289965.
- ↑ 14.0 14.1 "Mammalian Staufen1 recruits Upf1 to specific mRNA 3'UTRs so as to elicit mRNA decay". Cell 120 (2): 195–208. January 2005. doi:10.1016/j.cell.2004.11.050. PMID 15680326.
- ↑ "Mechanistic insights and identification of two novel factors in the C. elegans NMD pathway". Genes & Development 21 (9): 1075–85. May 2007. doi:10.1101/gad.417707. PMID 17437990.
- ↑ "Nonsense-mediated mRNA decay in Drosophila: at the intersection of the yeast and mammalian pathways". The EMBO Journal 22 (15): 3960–70. August 2003. doi:10.1093/emboj/cdg371. PMID 12881430.
- ↑ 17.0 17.1 17.2 "A rule for termination-codon position within intron-containing genes: when nonsense affects RNA abundance". Trends in Biochemical Sciences 23 (6): 198–9. June 1998. doi:10.1016/S0968-0004(98)01208-0. PMID 9644970.
- ↑ 18.0 18.1 18.2 "Lipid peroxidation of the microsomal fraction and extracted microsomal lipids from DAB-induced hepatomas". British Journal of Cancer 39 (6): 773–8. June 1979. doi:10.1128/mcb.18.9.5272. PMID 9710612.
- ↑ "Splicing and 3' end formation in the definition of nonsense-mediated decay-competent human beta-globin mRNPs". The EMBO Journal 20 (3): 532–40. February 2001. doi:10.1093/emboj/20.3.532. PMID 11157759.
- ↑ 20.0 20.1 "A conserved role for cytoplasmic poly(A)-binding protein 1 (PABPC1) in nonsense-mediated mRNA decay". The EMBO Journal 26 (6): 1591–601. March 2007. doi:10.1038/sj.emboj.7601588. PMID 17318186.
- ↑ 21.0 21.1 "Binding of a novel SMG-1-Upf1-eRF1-eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay". Genes & Development 20 (3): 355–67. February 2006. doi:10.1101/gad.1389006. PMID 16452507.
- ↑ "Nucleophosmin is selectively deposited on mRNA during polyadenylation". Nature Structural & Molecular Biology 13 (5): 429–35. May 2006. doi:10.1038/nsmb1080. PMID 16604083.
- ↑ "Stability of plant mRNAs depends on the length of the 3'-untranslated region". Biochemistry. Biokhimiia 71 (12): 1377–84. December 2006. doi:10.1134/s0006297906120145. PMID 17223792.
- ↑ "Plant nonsense-mediated mRNA decay is controlled by different autoregulatory circuits and can be induced by an EJC-like complex". Nucleic Acids Research 41 (13): 6715–28. July 2013. doi:10.1093/nar/gkt366. PMID 23666629.
- ↑ 25.0 25.1 "Nonsense mutations in close proximity to the initiation codon fail to trigger full nonsense-mediated mRNA decay". The Journal of Biological Chemistry 279 (31): 32170–80. July 2004. doi:10.1074/jbc.m405024200. PMID 15161914.
- ↑ "The canonical UPF1-dependent nonsense-mediated mRNA decay is inhibited in transcripts carrying a short open reading frame independent of sequence context". RNA 12 (12): 2160–70. December 2006. doi:10.1261/rna.201406. PMID 17077274.
- ↑ "Proximity of the poly(A)-binding protein to a premature termination codon inhibits mammalian nonsense-mediated mRNA decay". RNA 14 (3): 563–76. March 2008. doi:10.1261/rna.815108. PMID 18230761.
- ↑ "The highways and byways of mRNA decay". Nature Reviews. Molecular Cell Biology 8 (2): 113–26. February 2007. doi:10.1038/nrm2104. PMID 17245413.
- ↑ 29.0 29.1 29.2 29.3 "Exosome-mediated recognition and degradation of mRNAs lacking a termination codon". Science 295 (5563): 2262–4. March 2002. doi:10.1126/science.1067272. PMID 11910110. Bibcode: 2002Sci...295.2262V.
- ↑ 30.0 30.1 30.2 30.3 "An mRNA surveillance mechanism that eliminates transcripts lacking termination codons". Science 295 (5563): 2258–61. March 2002. doi:10.1126/science.1067338. PMID 11910109. Bibcode: 2002Sci...295.2258F.
- ↑ "Investigation of a pathogenic mtDNA microdeletion reveals a translation-dependent deadenylation decay pathway in human mitochondria". Human Molecular Genetics 12 (18): 2341–8. September 2003. doi:10.1093/hmg/ddg238. PMID 12915481.
- ↑ "The SsrA-SmpB system for protein tagging, directed degradation and ribosome rescue". Nature Structural Biology 7 (6): 449–55. June 2000. doi:10.1038/75843. PMID 10881189.
- ↑ 33.0 33.1 33.2 33.3 "Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation". Nature 440 (7083): 561–4. March 2006. doi:10.1038/nature04530. PMID 16554824. Bibcode: 2006Natur.440..561D.
- ↑ "Structural basis for mRNA surveillance by archaeal Pelota and GTP-bound EF1α complex". Proceedings of the National Academy of Sciences of the United States of America 107 (41): 17575–9. October 2010. doi:10.1073/pnas.1009598107. PMID 20876129. Bibcode: 2010PNAS..10717575K.
- ↑ "Structure of yeast Dom34: a protein related to translation termination factor Erf1 and involved in No-Go decay". The Journal of Biological Chemistry 283 (11): 7145–54. March 2008. doi:10.1074/jbc.M708224200. PMID 18180287.
- ↑ 36.0 36.1 "Evolution of nonstop, no-go and nonsense-mediated mRNA decay and their termination factor-derived components". BMC Evolutionary Biology 8: 290. October 2008. doi:10.1186/1471-2148-8-290. PMID 18947425.
External links
- The Daily Transcript: No-go Decay
- Chemistry of Life: non stop decay
- Notes mRNA surveillance (Yang Xu)
 |