Biology:Exosome complex

The exosome complex (or PM/Scl complex, often just called the exosome) is a multi-protein intracellular complex capable of degrading various types of RNA (ribonucleic acid) molecules. Exosome complexes are found in both eukaryotic cells and archaea, while in bacteria a simpler complex called the degradosome carries out similar functions.
The core of the exosome contains a six-membered ring structure to which other proteins are attached. In eukaryotic cells, the exosome complex is present in the cytoplasm, nucleus, and especially the nucleolus, although different proteins interact with the exosome complex in these compartments regulating the RNA degradation activity of the complex to substrates specific to these cell compartments. Substrates of the exosome include messenger RNA, ribosomal RNA, and many species of small RNAs. The exosome has an exoribonucleolytic function, meaning it degrades RNA starting at one end (the 3′ end in this case), and in eukaryotes also an endoribonucleolytic function, meaning it cleaves RNA at sites within the molecule.
Several proteins in the exosome are the target of autoantibodies in patients with specific autoimmune diseases (especially the PM/Scl overlap syndrome) and some antimetabolic chemotherapies for cancer function by blocking the activity of the exosome. In addition, mutations in exosome component 3 cause pontocerebellar hypoplasia and spinal motor neuron disease.
Discovery
The exosome was first discovered as an RNase in 1997 in the budding yeast Saccharomyces cerevisiae, an often-used model organism.[1] Not long after, in 1999, it was realized that the exosome was in fact the yeast equivalent of an already described complex in human cells called the PM/Scl complex, which had been identified as an autoantigen in patients with certain autoimmune diseases years earlier (see below).[2] Purification of this "PM/Scl complex" allowed the identification of more human exosome proteins and eventually the characterization of all components in the complex.[3][4] In 2001, the increasing amount of genome data that had become available allowed the prediction of exosome proteins in archaea, although it would take another 2 years before the first exosome complex from an archaeal organism was purified.[5][6]
Structure
Core proteins
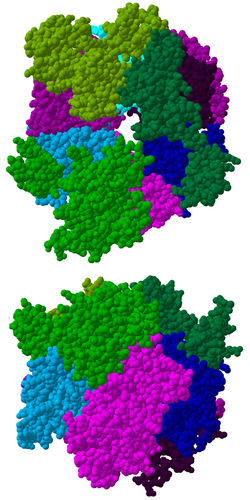
The core of the complex has a ring structure consisting of six proteins that all belong to the same class of RNases, the RNase PH-like proteins.[7] In archaea there are two different PH-like proteins (called Rrp41 and Rrp42), each present three times in an alternating order. Eukaryotic exosome complexes have six different proteins that form the ring structure.[8][9] Of these six eukaryotic proteins, three resemble the archaeal Rrp41 protein and the other three proteins are more similar to the archaeal Rrp42 protein.[10]

Located on top of this ring are three proteins that have an S1 RNA binding domain (RBD). Two proteins in addition have a K-homology (KH) domain.[7] In eukaryotes, three different "S1" proteins are bound to the ring, whereas in archaea either one or two different "S1" proteins can be part of the exosome (although there are always three S1 subunits attached to the complex).[11]
This ring structure is very similar to that of the proteins RNase PH and PNPase. In bacteria, the protein RNase PH, which is involved in tRNA processing, forms a hexameric ring consisting of six identical RNase PH proteins.[12][13] In the case of PNPase, which is a phosphorolytic RNA-degrading protein found in bacteria and the chloroplasts and mitochondria of some eukaryotic organisms, two RNase PH domains, and both an S1 and KH RNA binding domain are part of a single protein, which forms a trimeric complex that adopts a structure almost identical to that of the exosome.[14] Because of this high similarity in both protein domains and structure, these complexes are thought to be evolutionarily related and have a common ancestor.[15] The RNase PH-like exosome proteins, PNPase and RNase PH all belong to the RNase PH family of RNases and are phosphorolytic exoribonucleases, meaning that they use inorganic phosphate to remove nucleotides from the 3' end of RNA molecules.[7]
Associated proteins
Besides these nine core exosome proteins, two other proteins often associate with the complex in eukaryotic organisms. One of these is Rrp44, a hydrolytic RNase, which belongs to the RNase R family of hydrolytic exoribonucleases (nucleases that use water to cleave the nucleotide bonds). In addition to being an exoribonucleolytic enzyme, Rrp44 also has endoribonucleolytic activity, which resides in a separate domain of the protein.[16][17] In yeast, Rrp44 is associated with all exosome complexes and has a crucial role in the activity of the yeast exosome complex.[18] While a human homologue of the protein exists, no evidence was found for a long time that its human homologue was associated with the human exosome complex.[7] In 2010, however, it was discovered that humans have three Rrp44 homologues and two of these can be associated with the exosome complex. These two proteins most likely degrade different RNA substrates due to their different cellular localization, with one being localized in the cytoplasm (DIS3L1) and the other in the nucleus (DIS3).[19][20]
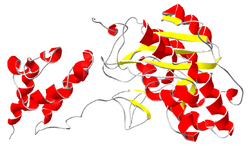
The second common associated protein is called Rrp6 (in yeast) or PM/Scl-100 (in human). Like Rrp44, this protein is a hydrolytic exoribonuclease, but in this case of the RNase D protein family.[21] The protein PM/Scl-100 is most commonly part of exosome complexes in the nucleus of cells, but can form part of the cytoplasmic exosome complex as well.[22]
Regulatory proteins
Apart from these two tightly bound protein subunits, many proteins interact with the exosome complex in both the cytoplasm and nucleus of cells. These loosely associated proteins may regulate the activity and specificity of the exosome complex. In the cytoplasm, the exosome interacts with AU-rich element (ARE) binding proteins (e.g. KRSP and TTP), which can promote or prevent degradation of mRNAs. The nuclear exosome associates with RNA binding proteins (e.g. MPP6/Mpp6 and C1D/Rrp47 in humans/yeast) that are required for processing certain substrates.[7]
In addition to single proteins, other protein complexes interact with the exosome. One of those is the cytoplasmic Ski complex, which includes an RNA helicase (Ski2) and is involved in mRNA degradation.[23] In the nucleus, the processing of rRNA and snoRNA by the exosome is mediated by the TRAMP complex, which contains both RNA helicase (Mtr4) and polyadenylation (Trf4) activity.[24]
Function
Enzymatic function
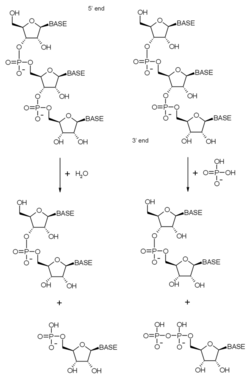
As stated above, the exosome complex contains many proteins with ribonuclease domains. The exact nature of these ribonuclease domains has changed across evolution from bacterial to archaeal to eukaryotic complexes as various activities have been gained and lost. The exosome is primarily a 3'-5' exoribonuclease, meaning that it degrades RNA molecules from their 3' end. Exoribonucleases contained in exosome complexes are either phosphorolytic (the RNase PH-like proteins) or, in eukaryotes, hydrolytic (the RNase R and RNase D domain proteins). The phosphorolytic enzymes use inorganic phosphate to cleave the phosphodiester bonds – releasing nucleotide diphosphates. The hydrolytic enzymes use water to hydrolyse these bonds – releasing nucleotide monophosphates.
In archaea, the Rrp41 subunit of the complex is a phosphorolytic exoribonuclease. Three copies of this protein are present in the ring and are responsible for the activity of the complex.[9] In eukaryotes, none of the RNase PH subunits have retained this catalytic activity, meaning the core ring structure of the human exosome has no enzymatically active protein.[25] Despite this loss of catalytic activity, the structure of the core exosome is highly conserved from archaea to humans, suggesting that the complex performs a vital cellular function. In eukaryotes, the absence of the phosphorolytic activity is compensated by the presence of the hydrolytic enzymes, which are responsible for the ribonuclease activity of the exosome in such organisms.[26][27][28]
As stated above, the hydrolytic proteins Rrp6 and Rrp44 are associated with the exosome in yeast and in humans, besides Rrp6, two different proteins, Dis3 and Dis3L1 can be associated at the position of the yeast Rrp44 protein.[19][20] Although originally the S1 domain proteins were thought to have 3'-5' hydrolytic exoribonuclease activity as well, the existence of this activity has recently been questioned and these proteins might have just a role in binding substrates prior to their degradation by the complex.[26]

Substrates
The exosome is involved in the degradation and processing of a wide variety of RNA species. In the cytoplasm of cells, it is involved in the turn-over of messenger RNA (mRNA) molecules. The complex can degrade mRNA molecules that have been tagged for degradation because they contain errors, through interactions with proteins from the nonsense mediated decay or non-stop decay pathways. In alternative fashion, mRNAs are degraded as part of their normal turnover. Several proteins that stabilize or destabilize mRNA molecules through binding to AU-rich elements in the 3' untranslated region of mRNAs interact with the exosome complex.[29][30][31] In the nucleus, the exosome is required for the correct processing of several small nuclear RNA molecules.[32] Finally, the nucleolus is the compartment where the majority of the exosome complexes are found. There it plays a role in the processing of the 5.8S ribosomal RNA (the first identified function of the exosome) and of several small nucleolar RNAs.[1][32][33]
Although most cells have other enzymes that can degrade RNA, either from the 3' or from the 5' end of the RNA, the exosome complex is essential for cell survival. When the expression of exosome proteins is artificially reduced or stopped, for example by RNA interference, growth stops and the cells eventually die. Both the core proteins of the exosome complex, as well as the two main associated proteins, are essential proteins.[34] Bacteria do not have an exosome complex; however, similar functions are performed by a simpler complex that includes the protein PNPase, called the degradosome.[35]

The exosome is a key complex in cellular RNA quality control. Unlike prokaryotes, eukaryotes possess highly active RNA surveillance systems that recognise unprocessed and mis-processed RNA-protein complexes (such as ribosomes) prior to their exit from the nucleus. It is presumed that this system prevents aberrant complexes from interfering with important cellular processes such as protein synthesis.[36]
In addition to RNA processing, turnover and surveillance activities, the exosome is important for the degradation of so-called cryptic unstable transcripts (CUTs) that are produced from thousands of loci within the yeast genome.[37][38] The importance of these unstable RNAs and their degradation are still unclear, but similar RNA species have also been detected in human cells.[39]
Disease
Autoimmunity
The exosome complex is the target of autoantibodies in patients with various autoimmune diseases. These autoantibodies are mainly found in people with the PM/Scl overlap syndrome, an autoimmune disease in which patients have symptoms from both scleroderma and either polymyositis or dermatomyositis.[40] Autoantibodies can be detected in the serum of patients by a variety of assays. In the past, the most commonly used methods were double immunodiffusion using calf thymus extracts, immunofluorescence on HEp-2 cells or immunoprecipitation from human cell extracts. In immunoprecipitation assays with sera from anti-exosome positive sera, a distinctive set of proteins is precipitated. Already years before the exosome complex was identified, this pattern was termed the PM/Scl complex.[41] Immunofluorescence using sera from these patients usually shows a typical staining of the nucleolus of cells, which sparked the suggestion that the antigen recognized by autoantibodies might be important in ribosome synthesis.[42] More recently, recombinant exosome proteins have become available and these have been used to develop line immunoassays (LIAs) and enzyme linked immunosorbent assays (ELISAs) for detecting these antibodies.[7]
In these diseases, antibodies are mainly directed against two of the proteins of the complex, called PM/Scl-100 (the RNase D like protein) and PM/Scl-75 (one of the RNase PH like proteins from the ring) and antibodies recognizing these proteins are found in approximately 30% of patients with the PM/Scl overlap syndrome.[43] Although these two proteins are the main target of the autoantibodies, other exosome subunits and associated proteins (like C1D) can be targeted in these patients.[44][45] At the current time, the most sensitive way to detect these antibodies is by using a peptide, derived from the PM/Scl-100 protein, as the antigen in an ELISA, instead of complete proteins. By this method, autoantibodies are found in up to 55% of patients with the PM/Scl overlap syndrome, but they can also be detected in patients with either scleroderma, polymyositis, or dermatomyositis alone.[46]
As the autobodies are found mainly in patients that have characteristics of several different autoimmune diseases, the clinical symptoms of these patients can vary widely. The symptoms that are seen most often are the typical symptoms of the individual autoimmune diseases and include Raynaud's phenomenon, arthritis, myositis and scleroderma.[47] Treatment of these patients is symptomatic and is similar to treatment for the individual autoimmune disease, often involving either immunosuppressive or immunomodulating drugs.[48]
Cancer treatment
The exosome has been shown to be inhibited by the antimetabolite fluorouracil, a drug used in the chemotherapy of cancer. It is one of the most successful drugs for treating solid tumors. In yeast cells treated with fluorouracil, defects were found in the processing of ribosomal RNA identical to those seen when the activity of the exosome was blocked by molecular biological strategies. Lack of correct ribosomal RNA processing is lethal to cells, explaining the antimetabolic effect of the drug.[49]
Neurological disorders
Mutations in exosome component 3 cause infantile spinal motor neuron disease, cerebellar atrophy, progressive microcephaly and profound global developmental delay, consistent with pontocerebellar hypoplasia type 1B (PCH1B; MIM 614678).[50]
List of subunits
| Legend | General name | Domains | Human | Yeast (S. cerevisiae) | Archaea | MW (kD) | Human gene | Yeast gene |
|---|---|---|---|---|---|---|---|---|
| 1 | Csl4 | S1 RBD | hCsl4 | Csl4p/Ski4p | Csl4 | 21–32 | EXOSC1 | Template:Yeast Gene |
| 2 | Rrp4 | S1/KH RBD | hRrp4 | Rrp4p | Rrp4 | 28–39 | EXOSC2 | Template:Yeast Gene |
| 3 | Rrp40 | S1/KH RBD | hRrp40 | Rrp40p | (Rrp4)A | 27–32 | EXOSC3 | Template:Yeast Gene |
| 4 | Rrp41 | RNase PH | hRrp41 | Rrp41p/Ski6p | Rrp41C | 26–28 | EXOSC4 | Template:Yeast Gene |
| 5 | Rrp46 | RNase PH | hRrp46 | Rrp46p | (Rrp41)A,C | 25–28 | EXOSC5 | Template:Yeast Gene |
| 6 | Mtr3 | RNase PH | hMtr3 | Mtr3p | (Rrp41)A,C | 24–37 | EXOSC6 | Template:Yeast Gene |
| 7 | Rrp42 | RNase PH | hRrp42 | Rrp42p | Rrp42 | 29–32 | EXOSC7 | Template:Yeast Gene |
| 8 | Rrp43 | RNase PH | OIP2 | Rrp43p | (Rrp42)A | 30–44 | EXOSC8 | Template:Yeast Gene |
| 9 | Rrp45 | RNase PH | PM/Scl-75 | Rrp45p | (Rrp42)A | 34–49 | EXOSC9 | Template:Yeast Gene |
| 10 | Rrp6 | RNase D | PM/Scl-100C | Rrp6pC | n/a | 84–100 | EXOSC10 | Template:Yeast Gene |
| 11 | Rrp44 | RNase R | Dis3B,C | Rrp44p/Dis3pC | n/a | 105–113 | DIS3
DIS3L1 |
- A In archaea several exosome proteins are present in multiple copies, to form the full core of the exosome complex.
- B In humans, two different proteins can be associated in this position. In the cytoplasm of cells, Dis3L1 is associated with the exosome, whereas in the nucleus, Dis3 can bind to the core complex.
- C Contributes to the ribonucleolytic activity of the complex.
See also
- The proteasome, the main protein degrading machinery of cells
- The spliceosome, a complex involved in RNA splicing, that also contains an RNA binding ring structure
References
- ↑ 1.0 1.1 Mitchell, P; Petfalski, E; Shevchenko, A; Mann, M; Tollervey, D (1997). "The Exosome: A Conserved Eukaryotic RNA Processing Complex Containing Multiple 3′→5′ Exoribonucleases". Cell 91 (4): 457–466. doi:10.1016/S0092-8674(00)80432-8. PMID 9390555.
- ↑ Allmang, C; Petfalski, E; Podtelejnikov, A; Mann, M; Tollervey, D; Mitchell, P (1999). "The yeast exosome and human PM-Scl are related complexes of 3' --> 5' exonucleases". Genes & Development 13 (16): 2148–58. doi:10.1101/gad.13.16.2148. PMID 10465791.
- ↑ Brouwer, R; Allmang, C; Raijmakers, R; Van Aarssen, Y; Egberts, WV; Petfalski, E; Van Venrooij, WJ; Tollervey, D et al. (2001). "Three novel components of the human exosome". Journal of Biological Chemistry 276 (9): 6177–84. doi:10.1074/jbc.M007603200. PMID 11110791.
- ↑ Chen, CY; Gherzi, R; Ong, SE; Chan, EL; Raijmakers, R; Pruijn, GJ; Stoecklin, G; Moroni, C et al. (2001). "AU binding proteins recruit the exosome to degrade ARE-containing mRNAs". Cell 107 (4): 451–64. doi:10.1016/S0092-8674(01)00578-5. PMID 11719186.
- ↑ Koonin, EV; Wolf, YI; Aravind, L (2001). "Prediction of the archaeal exosome and its connections with the proteasome and the translation and transcription machineries by a comparative-genomic approach". Genome Research 11 (2): 240–52. doi:10.1101/gr.162001. PMID 11157787.
- ↑ Evguenieva-Hackenberg, E; Walter, P; Hochleitner, E; Lottspeich, F; Klug, G (2003). "An exosome-like complex in Sulfolobus solfataricus". EMBO Reports 4 (9): 889–93. doi:10.1038/sj.embor.embor929. PMID 12947419.
- ↑ 7.0 7.1 7.2 7.3 7.4 7.5 Schilders, G; Van Dijk, E; Raijmakers, R; Pruijn, GJ (2006). Cell and molecular biology of the exosome: how to make or break an RNA. International Review of Cytology. 251. pp. 159–208. doi:10.1016/S0074-7696(06)51005-8. ISBN 9780123646552.
- ↑ Lorentzen, E; Walter, P; Fribourg, S; Evguenieva-Hackenberg, E; Klug, G; Conti, E (2005). "The archaeal exosome core is a hexameric ring structure with three catalytic subunits". Nature Structural & Molecular Biology 12 (7): 575–81. doi:10.1038/nsmb952. PMID 15951817.
- ↑ 9.0 9.1 Shen, V; Kiledjian, M (2006). "A view to a kill: structure of the RNA exosome". Cell 127 (6): 1093–5. doi:10.1016/j.cell.2006.11.035. PMID 17174886.
- ↑ Raijmakers, R; Egberts, WV; Van Venrooij, WJ; Pruijn, GJ (2002). "Protein-protein interactions between human exosome components support the assembly of RNase PH-type subunits into a six-membered PNPase-like ring". Journal of Molecular Biology 323 (4): 653–63. doi:10.1016/S0022-2836(02)00947-6. PMID 12419256.
- ↑ Walter, P; Klein, F; Lorentzen, E; Ilchmann, A; Klug, G; Evguenieva-Hackenberg, E (2006). "Characterization of native and reconstituted exosome complexes from the hyperthermophilic archaeon Sulfolobus solfataricus". Molecular Microbiology 62 (4): 1076–89. doi:10.1111/j.1365-2958.2006.05393.x. PMID 17078816.
- ↑ Ishii, R; Nureki, O; Yokoyama, S (2003). "Crystal structure of the tRNA processing enzyme RNase PH from Aquifex aeolicus". Journal of Biological Chemistry 278 (34): 32397–404. doi:10.1074/jbc.M300639200. PMID 12746447.
- ↑ Harlow, LS; Kadziola, A; Jensen, KF; Larsen, S (2004). "Crystal structure of the phosphorolytic exoribonuclease RNase PH from Bacillus subtilis and implications for its quaternary structure and tRNA binding". Protein Science 13 (3): 668–77. doi:10.1110/ps.03477004. PMID 14767080.
- ↑ Symmons, MF; Jones, GH; Luisi, BF (2000). "A duplicated fold is the structural basis for polynucleotide phosphorylase catalytic activity, processivity, and regulation". Structure 8 (11): 1215–26. doi:10.1016/S0969-2126(00)00521-9. PMID 11080643.
- ↑ Lin-Chao, S; Chiou, NT; Schuster, G (2007). "The PNPase, exosome and RNA helicases as the building components of evolutionarily-conserved RNA degradation machines". Journal of Biomedical Science 14 (4): 523–32. doi:10.1007/s11373-007-9178-y. PMID 17514363.
- ↑ Lebreton, A; Tomecki, R; Dziembowski, A; Séraphin, B (2008). "Endonucleolytic RNA cleavage by a eukaryotic exosome". Nature 456 (7224): 993–6. doi:10.1038/nature07480. PMID 19060886. Bibcode: 2008Natur.456..993L. https://hal-ens.archives-ouvertes.fr/hal-01350769/file/Exosome_endo_post-print.pdf.
- ↑ Schneider, C; Leung, E; Brown, J; Tollervey, D (2009). "The N-terminal PIN domain of the exosome subunit Rrp44 harbors endonuclease activity and tethers Rrp44 to the yeast core exosome.". Nucleic Acids Research 37 (4): 1127–40. doi:10.1093/nar/gkn1020. PMID 19129231.
- ↑ Schneider, C; Anderson, JT; Tollervey, D (2007). "The exosome subunit Rrp44 plays a direct role in RNA substrate recognition". Molecular Cell 27 (2): 324–31. doi:10.1016/j.molcel.2007.06.006. PMID 17643380. PMC 7610968. https://epublications.marquette.edu/cgi/viewcontent.cgi?article=1439&context=bio_fac.
- ↑ 19.0 19.1 Staals, RH; Bronkhorst, AW; Schilders, G; Slomovic, S; Schuster, G; Heck, AJ; Raijmakers, R; Pruijn, GJ (2010). "Dis3-like 1: a novel exoribonuclease associated with the human exosome.". The EMBO Journal 29 (14): 2358–67. doi:10.1038/emboj.2010.122. PMID 20531389.
- ↑ 20.0 20.1 Tomecki, R; Kristiansen, MS; Lykke-Andersen, S; Chlebowski, A; Larsen, KM; Szczesny, RJ; Drazkowska, K; Pastula, A et al. (2010). "The human core exosome interacts with differentially localized processive RNases: hDIS3 and hDIS3L". The EMBO Journal 29 (14): 2342–57. doi:10.1038/emboj.2010.121. PMID 20531386.
- ↑ Mian, IS (1997). "Comparative sequence analysis of ribonucleases HII, III, II PH and D". Nucleic Acids Research 25 (16): 3187–3195. doi:10.1093/nar/25.16.3187. PMID 9241229.
- ↑ Raijmakers, R; Schilders, G; Pruijn, GJ (2004). "The exosome, a molecular machine for controlled RNA degradation in both nucleus and cytoplasm". European Journal of Cell Biology 83 (5): 175–83. doi:10.1078/0171-9335-00385. PMID 15346807.
- ↑ Wang, L; Lewis, MS; Johnson, AW (2005). "Domain interactions within the Ski2/3/8 complex and between the Ski complex and Ski7p". RNA 11 (8): 1291–302. doi:10.1261/rna.2060405. PMID 16043509.
- ↑ LaCava, J; Houseley, J; Saveanu, C; Petfalski, E; Thompson, E; Jacquier, A; Tollervey, D (2005). "RNA degradation by the exosome is promoted by a nuclear polyadenylation complex". Cell 121 (5): 713–24. doi:10.1016/j.cell.2005.04.029. PMID 15935758.
- ↑ Liu, Q; Greimann, JC; Lima, CD (2007). "Erratum: Reconstitution, activities, and structure of the eukaryotic RNA exosome". Cell 131 (1): 188–189. doi:10.1016/j.cell.2007.09.019.
- ↑ 26.0 26.1 Dziembowski, A; Lorentzen, E; Conti, E; Séraphin, B (2007). "A single subunit, Dis3, is in essence responsible for yeast exosome core activity". Nature Structural & Molecular Biology 14 (1): 15–22. doi:10.1038/nsmb1184. PMID 17173052.
- ↑ Liu, Q; Greimann, JC; Lima, CD (2006). "Reconstitution, activities, and structure of the eukaryotic RNA exosome". Cell 127 (6): 1223–37. doi:10.1016/j.cell.2006.10.037. PMID 17174896.
- ↑ Lorentzen, E; Conti, E (2005). "Structural basis of 3' end RNA recognition and exoribonucleolytic cleavage by an exosome RNase PH core". Molecular Cell 20 (3): 473–81. doi:10.1016/j.molcel.2005.10.020. PMID 16285928.
- ↑ LeJeune, F; Li, X; Maquat, LE (2003). "Nonsense-mediated mRNA decay in mammalian cells involves decapping, deadenylating, and exonucleolytic activities". Molecular Cell 12 (3): 675–87. doi:10.1016/S1097-2765(03)00349-6. PMID 14527413.
- ↑ Wilson, MA; Meaux, S; Van Hoof, A (2007). "A genomic screen in yeast reveals novel aspects of nonstop mRNA metabolism". Genetics 177 (2): 773–84. doi:10.1534/genetics.107.073205. PMID 17660569.
- ↑ Lin, WJ; Duffy, A; Chen, CY (2007). "Localization of AU-rich element-containing mRNA in cytoplasmic granules containing exosome subunits". Journal of Biological Chemistry 282 (27): 19958–68. doi:10.1074/jbc.M702281200. PMID 17470429.
- ↑ 32.0 32.1 Allmang, C; Kufel, J; Chanfreau, G; Mitchell, P; Petfalski, E; Tollervey, D (1999). "Functions of the exosome in rRNA, snoRNA and snRNA synthesis". EMBO Journal 18 (19): 5399–410. doi:10.1093/emboj/18.19.5399. PMID 10508172.
- ↑ Schilders, G; Raijmakers, R; Raats, JM; Pruijn, GJ (2005). "MPP6 is an exosome-associated RNA-binding protein involved in 5.8S rRNA maturation". Nucleic Acids Research 33 (21): 6795–804. doi:10.1093/nar/gki982. PMID 16396833.
- ↑ van Dijk, EL; Schilders, G; Pruijn, GJ (2007). "Human cell growth requires a functional cytoplasmic exosome, which is involved in various mRNA decay pathways". RNA 13 (7): 1027–35. doi:10.1261/rna.575107. PMID 17545563.
- ↑ Carpousis AJ, AJ (2002). "The Escherichia coli RNA degradosome: structure, function and relationship in other ribonucleolytic multienzyme complexes". Biochem. Soc. Trans. 30 (2): 150–5. doi:10.1042/BST0300150. PMID 12035760.
- ↑ "RNA-quality control by the exosome". Nat. Rev. Mol. Cell Biol. 7 (7): 529–39. July 2006. doi:10.1038/nrm1964. PMID 16829983.
- ↑ Wyers F, F; Rougemaille, M; Badis, G et al. (June 2005). "Cryptic pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase". Cell 121 (5): 725–37. doi:10.1016/j.cell.2005.04.030. PMID 15935759.
- ↑ "Widespread bidirectional promoters are the major source of cryptic transcripts in yeast". Nature 457 (7232): 1038–42. February 2009. doi:10.1038/nature07747. PMID 19169244. Bibcode: 2009Natur.457.1038N.
- ↑ Preker P, P; Nielsen, J; Kammler, S et al. (December 2008). "RNA exosome depletion reveals transcription upstream of active human promoters". Science 322 (5909): 1851–4. doi:10.1126/science.1164096. PMID 19056938. Bibcode: 2008Sci...322.1851P.
- ↑ J.E. Pope, JE (2002). "Scleroderma overlap syndromes". Current Opinion in Rheumatology 14 (6): 704–10. doi:10.1097/00002281-200211000-00013. PMID 12410095.
- ↑ Gelpi, C; Algueró, A; Angeles Martinez, M; Vidal, S; Juarez, C; Rodriguez-Sanchez, JL (1991). "Identification of protein components reactive with anti-PM/Scl autoantibodies". Clinical and Experimental Immunology 81 (1): 59–64. doi:10.1111/j.1365-2249.1990.tb05291.x. PMID 2199097.
- ↑ Targoff, IN; Reichlin, M (1985). "Nucleolar localization of the PM-Scl antigen". Arthritis & Rheumatism 28 (2): 226–30. doi:10.1002/art.1780280221. PMID 3918546.
- ↑ Raijmakers, R; Renz, M; Wiemann, C; Egberts, WV; Seelig, HP; Van Venrooij, WJ; Pruijn, GJ (2004). "PM-Scl-75 is the main autoantigen in patients with the polymyositis/scleroderma overlap syndrome". Arthritis & Rheumatism 50 (2): 565–9. doi:10.1002/art.20056. PMID 14872500.
- ↑ Brouwer, R; Vree Egberts, WT; Hengstman, GJ; Raijmakers, R; Van Engelen, BG; Seelig, HP; Renz, M; Mierau, R et al. (2002). "Autoantibodies directed to novel components of the PM/Scl complex, the human exosome". Arthritis Research & Therapy 4 (2): 134–8. doi:10.1186/ar389. PMID 11879549.
- ↑ Schilders, G; Egberts, WV; Raijmakers, R; Pruijn, GJ (2007). "C1D is a major autoantibody target in patients with the polymyositis-scleroderma overlap syndrome". Arthritis & Rheumatism 56 (7): 2449–54. doi:10.1002/art.22710. PMID 17599775.
- ↑ Mahler, M; Raijmakers, R; Dähnrich, C; Blüthner, M; Fritzler, MJ (2005). "Clinical evaluation of autoantibodies to a novel PM/Scl peptide antigen". Arthritis Research & Therapy 7 (3): R704–13. doi:10.1186/ar1729. PMID 15899056.
- ↑ Mahler, M; Raijmakers, R (2007). "Novel aspects of autoantibodies to the PM/Scl complex: Clinical, genetic and diagnostic insights". Autoimmunity Reviews 6 (7): 432–7. doi:10.1016/j.autrev.2007.01.013. PMID 17643929.
- ↑ Jablonska, S; Blaszczyk, M (1998). "Scleromyositis: a scleroderma/polymyositis overlap syndrome". Clinical Rheumatology 17 (6): 465–7. doi:10.1007/BF01451281. PMID 9890673.
- ↑ Lum, PY; Armour, CD; Stepaniants, SB; Cavet, G; Wolf, MK; Butler, JS; Hinshaw, JC; Garnier, P et al. (2004). "Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes". Cell 116 (1): 121–37. doi:10.1016/S0092-8674(03)01035-3. PMID 14718172.
- ↑ Wan, J.; Yourshaw, M.; Mamsa, H.; Rudnik-Schöneborn, S.; Menezes, M. P.; Hong, J. E.; Leong, D. W.; Senderek, J. et al. (2012). "Mutations in the RNA exosome component gene EXOSC3 cause pontocerebellar hypoplasia and spinal motor neuron degeneration". Nature Genetics 44 (6): 704–708. doi:10.1038/ng.2254. PMID 22544365.
Further reading
- Schilders, G; Pruijn, GJ (2008). "Chapter 11 Biochemical Studies of the Mammalian Exosome with Intact Cells". RNA Turnover in Eukaryotes: Nucleases, Pathways and Analysis of mRNA Decay. Methods in Enzymology. 448. 211–226. doi:10.1016/S0076-6879(08)02611-6. ISBN 9780123743787.
- Houseley, J; Tollervey, D (2008). "The nuclear RNA surveillance machinery: the link between ncRNAs and genome structure in budding yeast?". Biochim Biophys Acta 1779 (4): 239–246. doi:10.1016/j.bbagrm.2007.12.008. PMID 18211833.
- Vanacova, S; Stefl, R (2007). "The exosome and RNA quality control in the nucleus". EMBO Reports 8 (7): 651–657. doi:10.1038/sj.embor.7401005. PMID 17603538.
- Büttner, K; Wenig, K; Hopfner, KP (2006). "The exosome: a macromolecular cage for controlled RNA degradation". Molecular Microbiology 61 (6): 1372–1379. doi:10.1111/j.1365-2958.2006.05331.x. PMID 16968219.
- Lorentzen, E; Conti, E (2006). "The Exosome and the Proteasome: Nano-Compartments for Degradation". Cell 125 (4): 651–654. doi:10.1016/j.cell.2006.05.002. PMID 16713559.
- Pruijn, GJ (2005). "Doughnuts dealing with RNA". Nature Structural & Molecular Biology 12 (7): 562–564. doi:10.1038/nsmb0705-562. PMID 15999107.
External links
- Structure of the human exosome at the RCSB Protein Data Bank
- Structure of an archaeal exosome at the RCSB Protein Data Bank
- Structure of an archaeal exosome bound to RNA at the RCSB Protein Data Bank
- Structure of the yeast exosome protein Rrp6 at the RCSB Protein Data Bank
- 3D macromolecular structures of exosomes at the EM Data Bank(EMDB)
 |
