Biology:Microscale thermophoresis
Microscale thermophoresis (MST) is a technology for the biophysical analysis of interactions between biomolecules. Microscale thermophoresis is based on the detection of a temperature-induced change in fluorescence of a target as a function of the concentration of a non-fluorescent ligand. The observed change in fluorescence is based on two distinct effects. On the one hand it is based on a temperature related intensity change (TRIC) of the fluorescent probe, which can be affected by binding events. On the other hand, it is based on thermophoresis, the directed movement of particles in a microscopic temperature gradient. Any change of the chemical microenvironment of the fluorescent probe, as well as changes in the hydration shell of biomolecules result in a relative change of the fluorescence detected when a temperature gradient is applied and can be used to determine binding affinities. MST allows measurement of interactions directly in solution without the need of immobilization to a surface (immobilization-free technology).
Applications
Affinity
- between any kind of biomolecules[1] including proteins, DNA,[2] RNA,[3] peptides,[4] small molecules,[5] fragments[6] and ions
- for interactions with high molecular weight complexes, large molecule assemblies, even with liposomes,[7] vesicles, nanodiscs,[8] nanoparticles[9] and viruses
- in any buffer, including serum and cell lysate[4]
- in competition experiments (for example with substrate and inhibitors)
Thermodynamic parameters
MST has been used to estimate the enthalpic and entropic contributions to biomolecular interactions.[10]
Additional information
- Sample property (homogeneity, aggregation, stability)
- Multiple binding sites, cooperativity
Technology
MST is based on the quantifiable detection of a fluorescence change in a sample when a temperature change is applied. The fluorescence of a target molecule can be extrinsic or intrinsic (aromatic amino acids) and is altered in temperature gradients due to two distinct effects. On the one hand temperature related intensity change (TRIC), which describes the intrinsic property of fluorophores to change their fluorescence intensity as a function of temperature. The extent of the change in fluorescence intensity is affected by the chemical environment of the fluorescent probe, which can be altered in binding events due to conformational changes or proximity of ligands.[11][12] On the other hand, MST is also based on the directed movement of molecules along temperature gradients, an effect termed thermophoresis. A spatial temperature difference ΔT leads to a change in molecule concentration in the region of elevated temperature, quantified by the Soret coefficient ST:chot/ccold = exp(-ST ΔT).[13][14] Both, TRIC and thermophoresis contribute to the recorded signal in MST measurements in the following way: ∂/∂T(cF)=c∂F/∂T+F∂c/∂T. The first term in this equation c∂F/∂T describes TRIC as a change in fluorescence intensity (F) as a function of temperature (T), whereas the second term F∂c/∂T describes thermophoresis as the change in particle concentration (c) as a function of temperature. Thermophoresis depends on the interface between molecule and solvent. Under constant buffer conditions, thermophoresis probes the size, charge and solvation entropy of the molecules. The thermophoresis of a fluorescently labeled molecule A typically differs significantly from the thermophoresis of a molecule-target complex AT due to size, charge and solvation entropy differences. This difference in the molecule's thermophoresis is used to quantify the binding in titration experiments under constant buffer conditions.
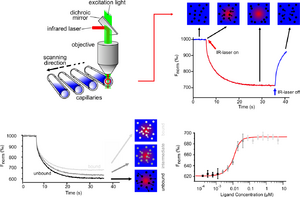
The thermophoretic movement of the fluorescently labelled molecule is measured by monitoring the fluorescence distribution F inside a capillary. The microscopic temperature gradient is generated by an IR-Laser, which is focused into the capillary and is strongly absorbed by water. The temperature of the aqueous solution in the laser spot is raised by ΔT=1-10 K. Before the IR-Laser is switched on a homogeneous fluorescence distribution Fcold is observed inside the capillary. When the IR-Laser is switched on, two effects, occur on the same time-scale, contributing to the new fluorescence distribution Fhot. The thermal relaxation induces a binding-dependent drop in the fluorescence of the dye due to its local environmental-dependent response to the temperature jump (TRIC). At the same time molecules typically move from the locally heated region to the outer cold regions. The local concentration of molecules decreases in the heated region until it reaches a steady-state distribution.
While the mass diffusion D dictates the kinetics of depletion, ST determines the steady-state concentration ratio chot/ccold=exp(-ST ΔT) ≈ 1-ST ΔT under a temperature increase ΔT. The normalized fluorescence Fnorm=Fhot/Fcold measures mainly this concentration ratio, in addition to TRIC ∂F/∂T. In the linear approximation we find: Fnorm=1+(∂F/∂T-ST)ΔT. Due to the linearity of the fluorescence intensity and the thermophoretic depletion, the normalized fluorescence from the unbound molecule Fnorm(A) and the bound complex Fnorm(AT) superpose linearly. By denoting x the fraction of molecules bound to targets, the changing fluorescence signal during the titration of target T is given by: Fnorm=(1-x) Fnorm(A)+x Fnorm(AT).[11]
Quantitative binding parameters are obtained by using a serial dilution of the binding substrate. By plotting Fnorm against the logarithm of the different concentrations of the dilution series, a sigmoidal binding curve is obtained. This binding curve can directly be fitted with the nonlinear solution of the law of mass action, with the dissociation constant KD as result.[15][16][17]
References
- ↑ "Thermophoresis for characterizing biomolecular interaction". Methods 146: 107–119. February 2018. doi:10.1016/j.ymeth.2018.02.003. PMID 29438829. http://repository.ubaya.ac.id/34334/1/Thermophoresis%20for%20characterizing%20biomolecular%20interaction.pdf.
- ↑ "Micro Scale Thermophoresis: A Rapid and Precise Method to Quantify Protein–Nucleic Acid Interactions in Solution". MicroScale Thermophoresis: A Rapid and Precise Method to Quantify Protein-Nucleic Acid Interactions in Solution. Methods in Molecular Biology. 1654. 2017. pp. 151–164. doi:10.1007/978-1-4939-7231-9_10. ISBN 978-1-4939-7230-2.
- ↑ "The extended AT-hook is a novel RNA binding motif". RNA Biology 12 (8): 864–76. 2015. doi:10.1080/15476286.2015.1060394. PMID 26156556.
- ↑ 4.0 4.1 "Microscale thermophoresis quantifies biomolecular interactions under previously challenging conditions". Methods 59 (3): 301–15. 2013. doi:10.1016/j.ymeth.2012.12.005. PMID 23270813.
- ↑ "Label-free microscale thermophoresis discriminates sites and affinity of protein-ligand binding". Angew. Chem. Int. Ed. Engl. 51 (42): 10656–9. 2012. doi:10.1002/anie.201204268. PMID 23001866.
- ↑ "An Automated Microscale Thermophoresis Screening Approach for Fragment-Based Lead Discovery". Journal of Biomolecular Screening 21 (4): 414–21. April 2016. doi:10.1177/1087057115618347. PMID 26637553.
- ↑ "Molecular interaction studies using microscale thermophoresis". Assay and Drug Development Technologies 9 (4): 342–53. August 2011. doi:10.1089/adt.2011.0380. PMID 21812660.
- ↑ "Lipid modulation of early G protein-coupled receptor signalling events". Biochimica et Biophysica Acta (BBA) - Biomembranes 1848 (11 Pt A): 2889–97. November 2015. doi:10.1016/j.bbamem.2015.08.004. PMID 26275588.
- ↑ "Understanding the Kinetics of Protein-Nanoparticle Corona Formation". ACS Nano 10 (12): 10842–10850. December 2016. doi:10.1021/acsnano.6b04858. PMID 28024351.
- ↑ "MicroScale Thermophoresis: Interaction analysis and beyond". Journal of Molecular Structure 1077: 101–113. 2014. doi:10.1016/j.molstruc.2014.03.009. Bibcode: 2014JMoSt1077..101J.
- ↑ 11.0 11.1 "Optical thermophoresis for quantifying the buffer dependence of aptamer binding". Angew. Chem. Int. Ed. Engl. 49 (12): 1–5. 2010. doi:10.1002/anie.200903998. PMID 20186894.
- "A hot road to new drugs". February 24, 2010. http://www.physorg.com/news186225693.html.
- ↑ Gupta, Amit J.; Duhr, Stefan; Baaske, Philipp (2018). "Microscale Thermophoresis (MST)". Encyclopedia of Biophysics. pp. 1–5. doi:10.1007/978-3-642-35943-9_10063-1. ISBN 9783642359439.
- ↑ "Why molecules move along a temperature gradient". Proc. Natl. Acad. Sci. U.S.A. 103 (52): 19678–82. 2006. doi:10.1073/pnas.0603873103. PMID 17164337. Bibcode: 2006PNAS..10319678D.
- ↑ "Thermophoresis of single stranded DNA". Electrophoresis 31 (2): 279–86. 2010. doi:10.1002/elps.200900505. PMID 20084627.
- ↑ "Protein-binding assays in biological liquids using microscale thermophoresis". Nat Commun 1 (7): 100. 2010. doi:10.1038/ncomms1093. PMID 20981028. Bibcode: 2010NatCo...1..100W.
- ↑ "Optisch erzeugte Thermophorese für die Bioanalytik" (in de). Biophotonik: 22–24. 2009. http://www.photonik.de/index.php?id=112&seitenid=11&fachid=2043&readpdf=biophotonik_2009_01_022.pdf.[|permanent dead link|dead link}}]
- ↑ "Thermophoretic melting curves quantify the conformation and stability of RNA and DNA". Nucleic Acids Res. 39 (8): e52. 2011. doi:10.1093/nar/gkr035. PMID 21297115.
 |
