Biology:Opisthokont
| Opisthokont | |
|---|---|
| |
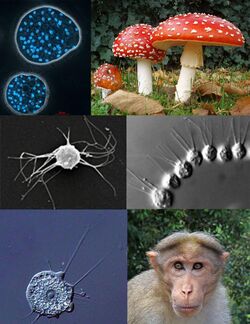
| Clockwise, from top left: Abeoforma whisleri (Mesomycetozoea); Amanita muscaria (Fungi); Desmarella moniliformis (Choanoflagellatea); bonnet macaque (Metazoa); Nuclearia thermophila (Nucleariida); Ministeria vibrans (Filasterea) | |
| Scientific classification | |
| Domain: | Eukaryota |
| Clade: | Amorphea |
| Clade: | Obazoa |
| (unranked): | Opisthokonta Copeland 1956,[2] emend. Cavalier-Smith 1987,[3] emend. Adl et al., 2005[4] |
| Subgroups | |
The opisthokonts (from grc ὀπίσθιος (opísthios) 'rear, posterior', and κοντός (kontós) 'pole, i.e. flagellum') are a broad group of eukaryotes, including both the animal and fungus kingdoms.[5] The opisthokonts, previously called the "Fungi/Metazoa group",[6] are generally recognized as a clade. Opisthokonts together with Apusomonadida and Breviata comprise the larger clade Obazoa.[7][8][9][10][11]
Flagella and other characteristics
A common characteristic of opisthokonts is that flagellate cells, such as the sperm of most animals and the spores of the chytrid fungi, propel themselves with a single posterior flagellum. It is this feature that gives the group its name. In contrast, flagellate cells in other eukaryote groups propel themselves with one or more anterior flagella. However, in some opisthokont groups, including most of the fungi, flagellate cells have been lost.[7]
Opisthokont characteristics include synthesis of extracellular chitin in exoskeleton, cyst/spore wall, or cell wall of filamentous growth and hyphae; the extracellular digestion of substrates with osmotrophic absorption of nutrients; and other cell biosynthetic and metabolic pathways. Genera at the base of each clade are amoeboid and phagotrophic.[12]
History
The close relationship between animals and fungi was suggested by Thomas Cavalier-Smith in 1987,[3] who used the informal name opisthokonta (the formal name has been used for the chytrids by Copeland in 1956), and was supported by later genetic studies.[13]
Early phylogenies placed fungi near the plants and other groups that have mitochondria with flat cristae, but this character varies. More recently, it has been said that holozoa (animals) and holomycota (fungi) are much more closely related to each other than either is to plants, because opisthokonts have a triple fusion of carbamoyl phosphate synthetase, dihydroorotase, and aspartate carbamoyltransferase that is not present in plants, and plants have a fusion of thymidylate synthase and dihydrofolate reductase not present in the opisthokonts. Animals and fungi are also more closely related to amoebas than to plants, and plants are more closely related to the SAR supergroup of protists than to animals or fungi.[citation needed] Animals and fungi are both heterotrophs, unlike plants, and while fungi are sessile like plants, there are also sessile animals.
Cavalier-Smith and Stechmann argue that the uniciliate eukaryotes such as opisthokonts and Amoebozoa, collectively called unikonts, split off from the other biciliate eukaryotes, called bikonts, shortly after they evolved.[14]
Taxonomy
Opisthokonts are divided into Holomycota or Nucletmycea (fungi and all organisms more closely related to fungi than to animals) and Holozoa (animals and all organisms more closely related to animals than to fungi); no opisthokonts basal to the Holomycota/Holozoa split have yet been identified.[citation needed] The Opisthokonts was largely resolved by Torriella et al.[15] Holomycota and Holozoa are composed of the following groups.Lua error: Internal error: The interpreter exited with status 1.
- Holomycota (Fungus-like)
- Fungi
- Includes:
- chytrids (flagellated, zoosporic fungi)
- Fonticula[16] (more recent work considers this to be part of Cristidiscoidea, a sister group to the fungi)
- Hyaloraphidium (previously thought to be a green alga, now considered a fungus)
- microsporidia (previously thought to be apicomplexia)
- Nucleariida (more recent work considers this to be part of Cristidiscoidea, a sister group to the fungi)
- Excludes:
- labyrinthulomycetes (slime nets) (now included in the SAR supergroup)
- myxomycetes (now included in amoebozoans)
- oomycetes (water molds) (now included in the SAR supergroup)
- Includes:
- Rozellida (placement uncertain)
- Fungi
- Holozoa (Animal-like)
- Corallochytrium (formerly considered a Heterokont)
- Filozoa
- Animalia (including myxozoa)
- Choanoflagellata (flagellates formerly included in protozoa)
- Filasterea
- Mesomycetozoea
- Amoebidiales (formerly considered trichomycetes)
- Dermocystida (formerly considered parasitic fungi or sporozoans)
- Eccrinales (formerly considered fungi)
- Ichthyophonida (formerly considered parasitic fungi incertae sedis)
Phylogeny
The following phylogenetic tree indicates the evolutionary relationships between the different opisthokont lineages, and the time divergence of the clades in millions of years ago (Mya).[17][18][19]
Lua error: Internal error: The interpreter exited with status 1.
Gallery
-
Microsporidian spore (Microsporidia)
-
Chytrid (flagellated fungus)
-
Sphaeroforma sp. (Mesomycetozoea)
-
Ministeria sp. (Filasterea)
-
Capsaspora sp. (Filasterea)
-
Salpingoeca (Choanoflagellatea)
-
Desmarella sp. colony (Choanoflagellatea)
-
Two opisthokonts: a human (Metazoa) and a mushroom (Fungi)
References
- ↑ Loron, Corentin C.; François, Camille; Rainbird, Robert H.; Turner, Elizabeth C.; Borensztajn, Stephan; Javaux, Emmanuelle J. (June 2019). "Early fungi from the Proterozoic era in Arctic Canada". Nature 570 (7760): 232–235. doi:10.1038/s41586-019-1217-0. PMID 31118507. Bibcode: 2019Natur.570..232L.
- ↑ The Classification of Lower Organisms. Palo Alto: Pacific Books. 1956. OCLC 477890. https://archive.org/details/classificationof00cope.
- ↑ 3.0 3.1 Rayner, Alan D. M., ed (1987). "The origin of fungi and pseudofungi". Evolutionary biology of Fungi. Cambridge: Cambridge University Press. pp. 339–353. ISBN 0-521-33050-5.
- ↑ Adl, Sina M.; Simpson, Alastair G. B.; Farmer, Mark A.; Andersen, Robert A.; Anderson, O. Roger; Barta, John R. et al. (September–October 2005). "The new higher level classification of eukaryotes with emphasis on the taxonomy of protists". The Journal of Eukaryotic Microbiology 52 (5): 399–451. doi:10.1111/j.1550-7408.2005.00053.x. PMID 16248873.
- ↑ "Multigene phylogeny of choanozoa and the origin of animals". PLOS ONE 3 (5): e2098. May 2008. doi:10.1371/journal.pone.0002098. PMID 18461162. Bibcode: 2008PLoSO...3.2098S.
- ↑ "Fungi/Metazoa group". UniProt. https://www.uniprot.org/taxonomy/33154.
- ↑ 7.0 7.1 "The protistan origins of animals and fungi". Molecular Biology and Evolution 23 (1): 93–106. January 2006. doi:10.1093/molbev/msj011. PMID 16151185.
- ↑ Huang, Jinling; Xu, Ying; Gogarten, Johann Peter (November 2005). "The presence of a haloarchaeal type tyrosyl-tRNA synthetase marks the opisthokonts as monophyletic". Molecular Biology and Evolution 22 (11): 2142–2146. doi:10.1093/molbev/msi221. PMID 16049196.
- ↑ "Evaluating support for the current classification of eukaryotic diversity". PLOS Genetics 2 (12): e220. December 2006. doi:10.1371/journal.pgen.0020220. PMID 17194223.
- ↑ Torruella, G.; Derelle, R.; Paps, J.; Lang, B. F.; Roger, A. J.; Shalchian-Tabrizi, K.; Ruiz-Trillo, I. (February 2012). "Phylogenetic relationships within the Opisthokonta based on phylogenomic analyses of conserved single-copy protein domains". Molecular Biology and Evolution 29 (2): 531–544. doi:10.1093/molbev/msr185. PMID 21771718.
- ↑ Eme, L.; Sharpe, S. C.; Brown, M. W.; Roger, A. J. (August 2014). "On the age of eukaryotes: evaluating evidence from fossils and molecular clocks". Cold Spring Harbor Perspectives in Biology 6 (8): a016139. doi:10.1101/cshperspect.a016139. PMID 25085908.
- ↑ Adl, Sina M.; Bass, David; Lane, Christopher E.; Lukeš, Julius; Schoch, Conrad L.; Smirnov, Alexey et al. (January 2019). "Revisions to the Classification, Nomenclature, and Diversity of Eukaryotes". The Journal of Eukaryotic Microbiology 66 (1): 4–119. doi:10.1111/jeu.12691. PMID 30257078.
- ↑ "Monophyletic origins of the metazoa: an evolutionary link with fungi". Science 260 (5106): 340–342. April 1993. doi:10.1126/science.8469985. PMID 8469985. Bibcode: 1993Sci...260..340W.
- ↑ Stechmann, Alexandra; Cavalier-Smith, Thomas (July 2002). "Rooting the eukaryote tree by using a derived gene fusion". Science 297 (5578): 89–91. doi:10.1126/science.1071196. PMID 12098695. Bibcode: 2002Sci...297...89S.
- ↑ Torruella, Guifré; De Mendoza, Alex; Grau-Bové, Xavier; Antó, Meritxell; Chaplin, Mark A.; Del Campo, Javier et al. (September 2015). "Phylogenomics Reveals Convergent Evolution of Lifestyles in Close Relatives of Animals and Fungi". Current Biology 25 (18): 2404–2410. doi:10.1016/j.cub.2015.07.053. PMID 26365255.
- ↑ Brown, M. W.; Spiegel, F. W.; Silberman, J. D. (December 2009). "Phylogeny of the "forgotten" cellular slime mold, Fonticula alba, reveals a key evolutionary branch within Opisthokonta". Molecular Biology and Evolution 26 (12): 2699–2709. doi:10.1093/molbev/msp185. PMID 19692665.
- ↑ Tedersoo, Leho; Sánchez-Ramírez, Santiago; Kõljalg, Urmas; Bahram, Mohammad; Döring, Markus; Schigel, Dmitry et al. (2018). "High-level classification of the Fungi and a tool for evolutionary ecological analyses". Fungal Diversity 90 (1): 135–159. doi:10.1007/s13225-018-0401-0. ISSN 1560-2745.
- ↑ Galindo, Luis Javier; Torruella, Guifré; López-García, Purificación; Ciobanu, Maria; Gutiérrez-Preciado, Ana; Karpov, Sergey A.; Moreira, David (June 2023). "Phylogenomics Supports the Monophyly of Aphelids and Fungi and Identifies New Molecular Synapomorphies". Systematic Biology 72 (3): 505–515. doi:10.1093/sysbio/syac054. PMID 35900180.
- ↑ Tikhonenkov, Denis V.; Mikhailov, Kirill V.; Hehenberger, Elisabeth; Karpov, Sergei A.; Prokina, Kristina I.; Esaulov, Anton S. et al. (November 2020). "New Lineage of Microbial Predators Adds Complexity to Reconstructing the Evolutionary Origin of Animals". Current Biology 30 (22): 4500–4509.e5. doi:10.1016/j.cub.2020.08.061. PMID 32976804.
Lua error: Internal error: The interpreter exited with status 1.
External links
Lua error: Internal error: The interpreter exited with status 1. Wikidata ☰ Q129021 entry
Lua error: Internal error: The interpreter exited with status 1.