Biology:Retron
A retron is a distinct DNA sequence found in the genome of many bacteria species that codes for reverse transcriptase and a unique single-stranded DNA/RNA hybrid called multicopy single-stranded DNA (msDNA). Retron msr RNA is the non-coding RNA produced by retron elements and is the immediate precursor to the synthesis of msDNA. The retron msr RNA folds into a characteristic secondary structure that contains a conserved guanosine residue at the end of a stem loop. Synthesis of DNA by the retron-encoded reverse transcriptase (RT) results in a DNA/RNA chimera which is composed of small single-stranded DNA linked to small single-stranded RNA. The RNA strand is joined to the 5′ end of the DNA chain via a 2′–5′ phosphodiester linkage that occurs from the 2′ position of the conserved internal guanosine residue.
Sequence and structure
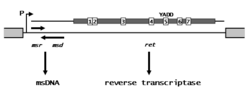
Retron elements are about 2 kb long. They contain a single operon controlling the synthesis of an RNA transcript carrying three loci, msr, msd, and ret, that are involved in msDNA synthesis. The DNA portion of msDNA is encoded by the msd gene, the RNA portion is encoded by the msr gene, while the product of the ret gene is a reverse transcriptase similar to the RTs produced by retroviruses and other types of retroelements.[1] Like other reverse transcriptases, the retron RT contains seven regions of conserved amino acids (labeled 1–7 in the figure), including a highly conserved tyr-ala-asp-asp (YADD) sequence associated with the catalytic core. The ret gene product is responsible for processing the msd/msr portion of the RNA transcript into msDNA.
Classification and occurrence
For many years after their discovery in animal viruses, reverse transcriptases were believed to be absent from prokaryotes. Currently, however, RT-encoding elements, i.e. retroelements, have been found in a wide variety of different bacteria:
- Retrons were the first family of retroelement discovered in bacteria; the other two families of known bacterial retroelements are:
- group II introns: Group II introns are the best characterized bacterial retroelement and the only type known to exhibit autonomous mobility; they consist of an RT encoded within a catalytic, self-splicing RNA structure. Group II intron mobility is mediated by a ribonucleoprotein comprising an intron lariat bound to two intron-coded proteins.[2]
- diversity-generating retroelements (DGRs).[3] The DGRs are not mobile, but function to diversify DNA sequences.[2] For example, DGRs mediate the switch between pathogenic and free-living phases of Bordetella.[4]
Function
Since retrons are not mobile, their appearance in diverse bacterial species is not a "selfish DNA" phenomenon. Rather, bacterial retrons confer some protection from phage infection to bacterial hosts. Several retrons are located in DNA regions next to certain protein effector-coding genes. When their expression is activated, most of these effectors and their associated retrons function together to block phage infection.[5][6]
Retrons in genetic engineering
Retrons have emerged as powerful tools in genetic engineering due to their unique ability to produce single-stranded DNA (ssDNA) inside cells. Here are some of the key ways retrons have been used:
In Situ DNA Production for Genome Editing
Retrons generate ssDNA through reverse transcription of a noncoding RNA. This ssDNA can serve as a donor template for genome editing, for example in recombineering and CRISPR-based systems. This approach allows for precise, targeted mutations without the need to introduce external DNA.[7][8]
Retron Library Recombineering (RLR)
RLR is a technique that enables massively parallel genome editing. It uses retrons to generate millions of unique mutations simultaneously, each tagged with a molecular "barcode."[9][10] This allows researchers to:
- Perform high-throughput genetic screens
- Simultaneously modify multiple sites on a single genome
- Study genotype-phenotype relationships
- Track mutations across large populations of cells
Biological Recording
Retrons have been engineered to act as molecular recorders, capturing information about cellular events by integrating specific DNA sequences into the genome. This could be used to monitor gene expression or environmental changes over time.[11]
Reduced Toxicity Compared to CRISPR
Unlike CRISPR-Cas9, which introduces double-stranded breaks (DSBs) that can be toxic or may lead to off-target effects, retron-based editing avoids DSBs, making it a reduced toxicity alternative for certain applications.[12][13]
Synthetic Biology and Evolutionary Engineering
Retrons are being explored for continuous evolution of synthetic genomes, enabling iterative cycles of mutation and selection to evolve new traits or functions in microbes.[14][15][16]
References
- ↑ "Retrons, msDNA, and the bacterial genome". Cytogenet Genome Res 110 (1–4): 491–499. 2005. doi:10.1159/000084982. PMID 16093702. http://gcat.davidson.edu/sybr-u/Lampson_2005.pdf.
- ↑ 2.0 2.1 "Diversity-Generating Retroelements". Current Opinion in Microbiology 10 (4): 388–395. 2007. doi:10.1016/j.mib.2007.06.004. PMID 17703991.
- ↑ "A diversity of uncharacterized reverse transcriptases in bacteria". Nucleic Acids Res. 36 (22): 7219–7229. 2008. doi:10.1093/nar/gkn867. PMID 19004871.
- ↑ "Genomic and Genetic Analysis of Bordetella Bacteriophages Encoding Reverse Transcriptase-Mediated Tropism-Switching Cassettes". J. Bacteriol. 186 (5): 1503–1517. 2004. doi:10.1128/JB.186.5.1503-1517.2004. PMID 14973019.
- ↑ Bobonis, Jacob; Mitosch, Karin; Mateus, André; Karcher, Nicolai; Kritikos, George; Selkrig, Joel; Zietek, Matylda; Monzon, Vivian et al. (2022-09-01). "Bacterial retrons encode phage-defending tripartite toxin–antitoxin systems" (in en). Nature 609 (7925): 144–150. doi:10.1038/s41586-022-05091-4. ISSN 0028-0836. PMID 35850148. Bibcode: 2022Natur.609..144B.
- ↑ "Bacterial Retrons Function In Anti-Phage Defense". Cell 183 (6): 1551–1561. 2020. doi:10.1016/j.cell.2020.09.065. PMID 33157039.
- ↑ Khan, Asim G.; Rojas-Montero, Matías; González-Delgado, Alejandro; Lopez, Santiago C.; Fang, Rebecca F.; Crawford, Kate D.; Shipman, Seth L. (2025). "An experimental census of retrons for DNA production and genome editing". Nature Biotechnology 43 (6): 914–922. doi:10.1038/s41587-024-02384-z. PMID 39289529. PMC 11911249. https://www.nature.com/articles/s41587-024-02384-z. Retrieved 6 July 2025.
- ↑ "Retrons and their applications in genome engineering". Nucleic Acids Research 47 (21): 11007–11019. 2019. doi:10.1093/nar/gkz865. PMID 31598685.
- ↑ Kaur, Navdeep; Pati, Pratap Kumar (2024). "Retron Library Recombineering: Next Powerful Tool for Genome Editing after CRISPR/Cas". ACS Synthetic Biology 13 (4): 1019–1025. doi:10.1021/acssynbio.3c00667. PMID 38480006. https://pubs.acs.org/doi/abs/10.1021/acssynbio.3c00667. Retrieved 6 July 2025.
- ↑ González-Delgado, Alejandro; Lopez, Santiago C.; Rojas-Montero, Matías; Fishman, Chloe B.; Shipman, Seth L. (2024). "Simultaneous multi-site editing of individual genomes using retron arrays". Nature Chemical Biology 20 (11): 1482–1492. doi:10.1038/s41589-024-01665-7. PMID 38982310.
- ↑ Jang, Hyeri; Yim, Sung Sun (2024). "Toward DNA-Based Recording of Biological Processes". Int. J. Mol. Sci. 25 (17): 9233. doi:10.3390/ijms25179233. PMID 39273181.
- ↑ Schubert, Max G.; Goodman, Daniel B.; Wannier, Timothy M.; Church, George M. (2021). "High-throughput functional variant screens via in vivo production of single-stranded DNA". Proc. Natl. Acad. Sci. 118 (18). doi:10.1073/pnas.2018181118. PMID 33906944. Bibcode: 2021PNAS..11818181S.
- ↑ Zhao, Bin; Chen, Shi-An A.; Lee, Jiwoo; Fraser, Hunter B. (2022). "Bacterial Retrons Enable Precise Gene Editing in Human Cells". The CRISPR Journal 5 (1): 31–39. doi:10.1089/crispr.2021.0065. PMID 35076284.
- ↑ Jiang, Wenjun; Rao, Gundra Sivakrishna; Aman, Rashid; Butt, Haroon; Kamel, Radwa; Sedeek, Khalid; Mahfouz, Magdy M. (2022). "High-efficiency retron-mediated single-stranded DNA production in plants". Synthetic Biology 7 (1). doi:10.1093/synbio/ysac025. PMID 36452068. PMC 9700382. https://academic.oup.com/synbio/article/7/1/ysac025/6786268. Retrieved 6 July 2025.
- ↑ Ellington, Adam J.; Reisch, Christopher R. (2022). "Efficient and iterative retron-mediated in vivo recombineering in Escherichia coli". Synthetic Biology 7 (1). doi:10.1093/synbio/ysac007. PMID 35673614. PMC 9165427. https://academic.oup.com/synbio/article/7/1/ysac007/6578729. Retrieved 6 July 2025.
- ↑ Liu, Wenqian; Zuo, Siqi; Shao, Youran; Bi, Ke; Zhao, Jiarun; Huang, Lei; Xu, Zhinan; Lian, Jiazhang (2023). "Retron-mediated multiplex genome editing and continuous evolution in Escherichia coli". Nucleic Acids Research 51 (15): 8293–8307. doi:10.1093/nar/gkad607. PMID 37471041. PMC 10450171. https://academic.oup.com/nar/article/51/15/8293/7226867. Retrieved 6 July 2025.
External links
- Page for Retron msr RNA at Rfam
- Mystery molecule in bacteria is revealed to be a guard, on: EurekAlert!, 5 Nov 2020. Source: WEIZMANN INSTITUTE OF SCIENCE
 |
