Biology:Group II intron
| Group II catalytic intron, D5 | |
|---|---|
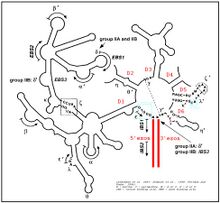 full secondary structure of group II intron | |
| Identifiers | |
| Symbol | Intron_gpII |
| Rfam | RF00029 |
| Other data | |
| PDB structures | PDBe 6cih |
| Extra information | Entry contains Splicing domain V (D5) and some consensus 3' to it. |
| Group II catalytic intron, D1-D4 | |
|---|---|
| Identifiers | |
| Symbol | group-II-D1D4 |
| Rfam | CL00102 |
| Other data | |
| PDB structures | PDBe 4fb0 |
| Extra information | Entry contains D1-D4, parts 5' to D5. |
Group II introns are a large class of self-catalytic ribozymes and mobile genetic elements found within the genes of all three domains of life. Ribozyme activity (e.g., self-splicing) can usually occur under high-salt conditions in vitro for most group II introns. Certain highly reactive group II introns can undergo self-splicing under mild, near-physiologically relevant salt conditions.[1] However, assistance from proteins is required for in vivo splicing.[2] In contrast to group I introns, intron excision occurs in the absence of GTP and involves the formation of a lariat, with an A-residue branchpoint strongly resembling that found in lariats formed during splicing of nuclear pre-mRNA. It is hypothesized that pre-mRNA splicing (see spliceosome) may have evolved from group II introns, due to the similar catalytic mechanism as well as the structural similarity of the Group II Domain V substructure to the U6/U2 extended snRNA.[3][4][5] Finally, their ability to site-specifically insert into DNA sites has been exploited as a tool for biotechnology.[6] For example, group II introns can be modified to make site-specific genome insertions and deliver cargo DNA such as reporter genes or lox sites [7]
Structure and catalysis

The secondary structure of group II introns is characterized by six typical stem-loop structures, also called domains I to VI (DI to DVI, or D1 to D6). The domains radiate from a central core that brings the 5' and 3' splice junctions into close proximity. The proximal helix structures of the six domains are connected by a few nucleotides in the central region (linker or joiner sequences). Due to its enormous size, the domain I was divided further into subdomains a, b, c, and d. Sequence differences of group II introns that led to a further division into subgroups IIA, IIB and IIC were identified, along with varying distance of the bulged adenosine in domain VI (the prospective branch point forming the lariat) from the 3' splice site, and the inclusion or omission of structural elements such as a coordination loop in domain I, which is present in IIB and IIC introns but not IIA.[2] Group II introns also form very complicated RNA Tertiary Structure.
Group II introns possess only a very few conserved nucleotides, and the nucleotides important for the catalytic function are spread over the complete intron structure. The few strictly conserved primary sequences are the consensus at the 5' and 3' splicing site (...↓GUGYG&... and ...AY↓..., with the Y representing a pyrimidine), some of the nucleotides of the central core (joiner sequences), a relatively high number of nucleotides of DV and some short-sequence stretches of DI. The unpaired adenosine in DVI (marked by an asterisk in the figure and located 7 or 8 nt away from the 3' splicing site) is also conserved and plays a central role in the splicing process. The 2' hydroxyl of the bulged adenosine attacks the 5' splice site, followed by nucleophilic attack on the 3' splice site by the 3' OH of the upstream exon. This results in a branched intron lariat connected by a 2' phosphodiester linkage at the DVI adenosine.
Protein machinery is required for splicing in vivo, and long-range intron-intron and intron-exon interactions are important for splice site positioning, as well as a number of tertiary contacts between motifs, including kissing-loop and tetraloop-receptor interactions. In 2005, A. De Lencastre et al. found that during splicing of Group II introns, all reactants are preorganized before the initiation of splicing. The branch site, both exons, the catalytically essential regions of DV and J2/3, and ε−ε' are in close proximity before the first step of splicing occurs. In addition to the bulge and AGC triad regions of DV, the J2/3 linker region, the ε−ε' nucleotides and the coordination loop in DI are crucial for the architecture and function of the active-site.[8]
The first crystal structure of a group II intron was resolved in 2008 for the Oceanobacillus iheyensis group IIC catalytic intron,[9] and was joined by the Pylaiella littoralis (P.li.LSUI2) group IIB intron in 2014. Attempts have been made to model the tertiary structure of other group II introns, such as the ai5γ group IIB intron, using a combination of programs for homology mapping onto known structures and de novo modeling of previously unresolved regions.[10] Group IIC are characterized by a catalytic triad made up by CGC, while Group IIA and Group IIB are made up by AGC catalytic triad, which is more similar to the catalytic triad of the spliceosome. It is believed that the Group IIC are also smaller, more reactive and more ancient. The first step of splicing in Group IIC intron is done by water and it form a linear structure instead of lariat without the assistance of protein cofactors.[11] Under the facilitation of intron-encoded maturase protein, Group IIC intron will form the lariat structure.[5] A series of high-resolution crystal structures revealed the different structural conformations Group II introns assume during splicing.[12][13] The mechanism of retrotransposition by Group II introns have also been revealed through Cryo - Electron Microscopy.[14]
Permuted forms of group II introns are conserved in some bacteria,[15] but their biological function is unknown. In these permuted forms of the ribozyme, the elements of the conserved group II intron structure are present, but occur in a different order.
Distribution and phylogeny
| Domain X | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Domain_X | ||||||||
| Pfam | PF01348 | ||||||||
| Pfam clan | CL0359 | ||||||||
| InterPro | IPR024937 | ||||||||
| |||||||||
| Group II intron, maturase-specific | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | GIIM | ||||||||
| Pfam | PF08388 | ||||||||
| Pfam clan | CL0359 | ||||||||
| InterPro | IPR013597 | ||||||||
| |||||||||
Group II introns are found in rRNA, tRNA, and mRNA of organelles (chloroplasts and mitochondria) in fungi, plants, and protists, and also in mRNA in bacteria. The first intron to be identified as distinct from group I was the ai5γ group IIB intron, which was isolated in 1986 from a pre-mRNA transcript of the oxi 3 mitochondrial gene of Saccharomyces cerevisiae.[16]
A subset of group II introns encode essential splicing proteins, known as intron-encoded proteins or IEPs, in intronic ORFs. The length of these introns can, as a result, be up to 3 kb. Splicing occurs in almost identical fashion to nuclear pre-mRNA splicing with two transesterification steps, with both also using magnesium ions to stabilize the leaving group in each step, which has led some to theorize a phylogenetic link between group II introns and the nuclear spliceosome. Further evidence for this link includes structural similarity between the U2/U6 junction of spliceosomal RNA and domain V of group II introns, which contains the catalytic AGC triad and much of the heart of the active site, as well as parity between conserved 5' and 3' end sequences.[17] Structural dynamics of group II intron domain VI during the two catalytic steps of splicing also resembles that of the spliceosomal U2 snRNA-intron branch helix, which further supports the evolutional link of the two splicing machineries.[5]
Many of these IEPs, including LtrA, share a reverse transcriptase domain and a "Domain X".[18] Maturase K (MatK) is a protein somewhat similar to those intron-encoded proteins, found in plant chloroplasts. It is required for in vivo splicing of Group II introns, and can be found in chloroplastic introns or in the nuclear genome. Its RT domain is broken.[18]
Protein domain
Group II IEPs share a related conserved domain, known as either "Domain X" in organelles or "GIIM" in bacteria, that is not found in other retroelements.[19][20] Domain X is essential for splicing in yeast mitochondria.[21] This domain may be responsible for recognizing and binding to intron RNA[20] or DNA.[22]
See also
- Database for bacterial group II introns
- Intron
- Splice site
- Nuclear introns
- Group I intron
- Group III intron
- Twintron
- LtrA
References
- ↑ Liu, Tianshuo; Pyle, Anna M (2021-12-02). "Discovery of highly reactive self-splicing group II introns within the mitochondrial genomes of human pathogenic fungi" (in en). Nucleic Acids Research 49 (21): 12422–12432. doi:10.1093/nar/gkab1077. ISSN 0305-1048. PMID 34850132. PMC 8643640. https://academic.oup.com/nar/article/49/21/12422/6439699.
- ↑ 2.0 2.1 "Group II introns: mobile ribozymes that invade DNA". Cold Spring Harbor Perspectives in Biology 3 (8). August 2011. doi:10.1101/cshperspect.a003616. PMID 20463000.
- ↑ "Structure of a self-splicing group II intron catalytic effector domain 5: parallels with spliceosomal U6 RNA". RNA 12 (2): 235–47. February 2006. doi:10.1261/rna.2237806. PMID 16428604.
- ↑ "Role of the snRNAs in spliceosomal active site". RNA Biology 7 (3): 345–53. May–Jun 2010. doi:10.4161/rna.7.3.12089. PMID 20458185.
- ↑ 5.0 5.1 5.2 Xu, Ling; Liu, Tianshuo; Chung, Kevin; Pyle, Anna Marie (2023-12-21). "Structural insights into intron catalysis and dynamics during splicing" (in en). Nature 624 (7992): 682–688. doi:10.1038/s41586-023-06746-6. ISSN 0028-0836. PMID 37993708. Bibcode: 2023Natur.624..682X.
- ↑ "Group II introns as controllable gene targeting vectors for genetic manipulation of bacteria". Nat Biotechnol 19 (12): 1162–7. December 2001. doi:10.1038/nbt1201-1162. PMID 11731786.
- ↑ "A Targetron-Recombinase System for Large-Scale Genome Engineering of Clostridia". mSphere 4 (6). December 2019. doi:10.1128/mSphere.00710-19. PMID 31826971.
- ↑ "A single active-site region for a group II intron". Nature Structural & Molecular Biology 12 (7): 626–7. July 2005. doi:10.1038/nsmb957. PMID 15980867.
- ↑ Toor, Navtej; Keating, Kevin S.; Taylor, Sean D.; Pyle, Anna Marie (2008-04-04). "Crystal Structure of a Self-Spliced Group II Intron" (in en). Science 320 (5872): 77–82. doi:10.1126/science.1153803. ISSN 0036-8075. PMID 18388288. PMC 4406475. https://www.science.org/doi/10.1126/science.1153803.
- ↑ "Visualizing the ai5γ group IIB intron". Nucleic Acids Research 42 (3): 1947–58. February 2014. doi:10.1093/nar/gkt1051. PMID 24203709.
- ↑ "A structural analysis of the group II intron active site and implications for the spliceosome". RNA 16 (1): 1–9. January 2010. doi:10.1261/rna.1791310. PMID 19948765.
- ↑ Chan, Russell T.; Peters, Jessica K.; Robart, Aaron R.; Wiryaman, Timothy; Rajashankar, Kanagalaghatta R.; Toor, Navtej (2018-11-08). "Structural basis for the second step of group II intron splicing" (in en). Nature Communications 9 (1): 4676. doi:10.1038/s41467-018-06678-0. ISSN 2041-1723. PMID 30410046. PMC 6224600. https://www.nature.com/articles/s41467-018-06678-0.
- ↑ Marcia, Marco; Pyle, Anna Marie (October 2012). "Visualizing Group II Intron Catalysis through the Stages of Splicing" (in en). Cell 151 (3): 497–507. doi:10.1016/j.cell.2012.09.033. PMC 3628766. https://linkinghub.elsevier.com/retrieve/pii/S0092867412011798.
- ↑ Haack, Daniel B.; Yan, Xiaodong; Zhang, Cheng; Hingey, Jason; Lyumkis, Dmitry; Baker, Timothy S.; Toor, Navtej (July 2019). "Cryo-EM Structures of a Group II Intron Reverse Splicing into DNA" (in en). Cell 178 (3): 612–623.e12. doi:10.1016/j.cell.2019.06.035. PMID 31348888. PMC 6662634. https://linkinghub.elsevier.com/retrieve/pii/S009286741930738X.
- ↑ "Natural circularly permuted group II introns in bacteria produce RNA circles". iScience 24 (12). December 2021. doi:10.1016/j.isci.2021.103431. PMID 34901790. Bibcode: 2021iSci...24j3431R.
- ↑ "A self-splicing RNA excises an intron lariat". Cell 44 (2): 213–23. January 1986. doi:10.1016/0092-8674(86)90755-5. PMID 3510741.
- ↑ "Metal ion catalysis during the exon-ligation step of nuclear pre-mRNA splicing: extending the parallels between the spliceosome and group II introns". RNA 6 (2): 199–205. February 2000. doi:10.1017/S1355838200992069. PMID 10688359. PMC 1369906. http://rnajournal.cshlp.org/content/6/2/199.long.
- ↑ 18.0 18.1 "Evolutionary origin of a plant mitochondrial group II intron from a reverse transcriptase/maturase-encoding ancestor". Journal of Plant Research 119 (4): 363–71. July 2006. doi:10.1007/s10265-006-0284-0. PMID 16763758. Bibcode: 2006JPlR..119..363A.
- ↑ "Evolutionary relationships among group II intron-encoded proteins and identification of a conserved domain that may be related to maturase function". Nucleic Acids Research 21 (22): 4991–7. November 1993. doi:10.1093/nar/21.22.4991. PMID 8255751.
- ↑ 20.0 20.1 "Presence of a group II intron in a multiresistant Serratia marcescens strain that harbors three integrons and a novel gene fusion". Antimicrobial Agents and Chemotherapy 46 (5): 1402–9. May 2002. doi:10.1128/AAC.46.5.1402-1409.2002. PMID 11959575.
- ↑ "Splicing defective mutants of the COXI gene of yeast mitochondrial DNA: initial definition of the maturase domain of the group II intron aI2". Nucleic Acids Research 22 (11): 2057–64. June 1994. doi:10.1093/nar/22.11.2057. PMID 8029012.
- ↑ "Group II intron endonucleases use both RNA and protein subunits for recognition of specific sequences in double-stranded DNA". The EMBO Journal 16 (22): 6835–48. November 1997. doi:10.1093/emboj/16.22.6835. PMID 9362497.
Further reading
- "The ins and outs of group II introns". Trends in Genetics 17 (6): 322–31. June 2001. doi:10.1016/S0168-9525(01)02324-1. PMID 11377794.
- "Control of branch-site choice by a group II intron". The EMBO Journal 20 (23): 6866–76. December 2001. doi:10.1093/emboj/20.23.6866. PMID 11726522.
- "Group II introns: structure and catalytic versatility of large natural ribozymes". Critical Reviews in Biochemistry and Molecular Biology 38 (3): 249–303. 2003. doi:10.1080/713609236. PMID 12870716.
- "Comparative and functional anatomy of group II catalytic introns--a review". Gene 82 (1): 5–30. October 1989. doi:10.1016/0378-1119(89)90026-7. PMID 2684776.
External links
 |
