Biology:Steroid Delta-isomerase
| Steroid Δ-isomerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
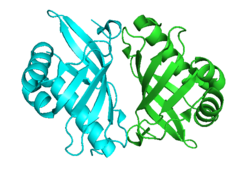 Crystallographic structure of Pseudomonas putida steroid Δ5-isomerase homodimer.[1] | |||||||||
| Identifiers | |||||||||
| EC number | 5.3.3.1 | ||||||||
| CAS number | 9031-36-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
In enzymology, a steroid Δ5-isomerase (EC 5.3.3.1) is an enzyme that catalyzes the chemical reaction
- a 3-oxo-Δ5-steroid [math]\displaystyle{ \rightleftharpoons }[/math] a 3-oxo-Δ4-steroid
Hence, this enzyme has one substrate, a 3-oxo-Δ5-steroid, and one product, a 3-oxo-Δ4-steroid.
Introduction
This enzyme belongs to the family of isomerases, specifically those intramolecular oxidoreductases transposing C=C bonds. The systematic name of this enzyme class is 3-oxosteroid Δ5-Δ4-isomerase. Other names in common use include ketosteroid isomerase (KSI), hydroxysteroid isomerase, steroid isomerase, Δ5-ketosteroid isomerase, Δ5(or Δ4)-3-keto steroid isomerase, Δ5-steroid isomerase, 3-oxosteroid isomerase, Δ5-3-keto steroid isomerase, and Δ5-3-oxosteroid isomerase.
KSI has been studied extensively from the bacteria Comamonas testosteroni (TI), formerly referred to as Pseudomonas testosteroni, and Pseudomonas putida (PI).[2] The enzymes from these two sources are 34% homologous, and structural studies have shown that the placement of the catalytic groups in the active sites is virtually identical.[3] Mammalian KSI has been studied from bovine adrenal cortex[4] and rat liver.[5] This enzyme participates in c21-steroid hormone metabolism and androgen and estrogen metabolism. An example substrate is Δ5-androstene-3,17-dione, which KSI converts to Δ4-androstene-3,17-dione.[6] The above reaction in the absence of enzyme takes 7 weeks to complete in aqueous solution.[7] KSI performs this reaction on an order of 1011 times faster, ranking it among the most proficient enzymes known.[7] Bacterial KSI also serves as a model protein for studying enzyme catalysis[8] and protein folding.[9]
Structural studies
KSI exists as a homodimer with two identical halves.[9] The interface between the two monomers is narrow and well defined, consisting of neutral or apolar amino acids, suggesting the hydrophobic interaction is important for dimerization.[9] Results show that the dimerization is essential to function.[9] The active site is highly apolar and folds around the substrate in a manner similar to other enzymes with hydrophobic substrates, suggesting this fold is characteristic for binding hydrophobic substrates.[10]
No complete atomic structure of KSI appeared until 1997, when an NMR structure of TI KSI was reported.[11] This structure showed that the active site is a deep hydrophobic pit with Asp-38 and Tyr-14 located at the bottom of this pit.[11] The structure is thus entirely consistent with the proposed mechanistic roles of Asp-38 and Tyr-14.
| Residue Role | Comamonas testosteroni (PDB: 8CHO) | Pseudomonas putida (PDB: 1OH0) |
|---|---|---|
| Oxyanion H-Bond Donor(s) | Asp-99 | Asp-103 |
| Tyr-14 | Tyr-16 | |
| General Acid/Base | Asp-38 | Asp-40 |
As of late 2007, 25 structures have been solved for this class of enzymes, with PDB accession codes 1BUQ, 1C7H, 1CQS, 1DMM, 1DMN, 1DMQ, 1E97, 1GS3, 1ISK, 1K41, 1OCV, 1OGX, 1OGZ, 1OH0, 1OHO, 1OHP, 1OHS, 1OPY, 1VZZ, 1W00, 1W01, 1W02, 1W6Y, 2PZV, and 8CHO.
Mechanism
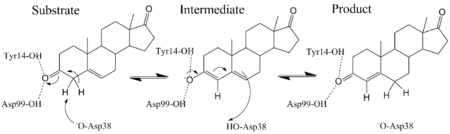
KSI catalyzes the rearrangement of a carbon-carbon double bond in ketosteroids through an enolate intermediate at a diffusion-limited rate.[2] There have been conflicting results on the ionization state of the intermediate, whether it exists as the enolate[12] or enol.[13] Pollack uses a thermodynamic argument to suggest the intermediate exists as the enolate.[2] The general base Asp-38 abstracts a proton from position 4 (alpha to the carbonyl, next to the double bond) of the steroid ring to form an enolate (the rate-limiting step)[14] that is stabilized by the hydrogen bond donating Tyr-14 and Asp-99.[2] Tyr-14 and Asp-99 are positioned deep within the hydrophobic active site and form a so-called oxanion hole.[15] Protonated Asp-38 then transfers its proton to position 6 of the steroid ring to complete the reaction.[2]
Although the mechanistic steps of the reaction are not disputed, the contributions of various factors to catalysis such as electrostatics, hydrogen bonding of the oxyanion hole, and distal binding effects are discussed below and still debated.
The Warshel group applied statistical mechanical computational methods and empirical valence bond theory to previous experimental data. It was determined that electrostatic preorganization-including ionic residues and fixed dipoles within the active site-contributes most to KSI catalysis.[16] More specifically, Tyr-14 and Asp-99 dipoles work to stabilize the growing charge which accumulates on the enolate oxygen (O-3) throughout catalysis. In a similar way, the charge on Asp38 is stabilized by surrounding residues and a water molecule during the course of the reaction.[16] The Boxer group used experimental Stark spectroscopy methods to identify the presence of H-bond-mediated electric fields within the KSI active site. These measurements quantified the electrostatic contribution to KSI catalysis (70%).[17]
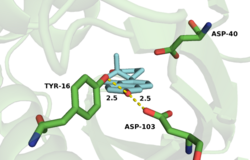
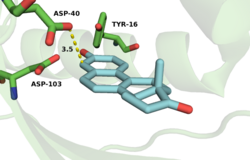
The active site is lined with hydrophobic residues to accommodate the substrate, but Asp-99 and Tyr-14 are within hydrogen bonding distance of O-3.[18] The hydrogen bonds from Tyr-14 and Asp-99 are known to significantly affect the rate of catalysis in KSI.[2] Mutagenesis of this residue to alanine (D99A) or asparagine (D99N) results in a loss in activity at pH 7 of 3000-fold and 27-fold, respectively,[11][19] implicating Asp-99 as important for enzymatic activity. Wu et al.[11] proposed a mechanism that involves both Tyr-14 and Asp-99 forming hydrogen bonds directly to O-3 of the steroid. This mechanism was challenged by Zhao et al.,[20] who postulated a hydrogen bonding network with Asp-99 hydrogen bonding to Tyr-14, which in turn forms a hydrogen bond to O-3. More recently, the Herschlag group utilized unnatural amino acid incorporation to assay the importance of Tyr-14 to KSI catalysis.[21] The natural tyrosine residue was substituted with unnatural halogenated amino acids surveying a range of pKa's. There was very little difference in KSI catalytic turnover with decreasing pKa, suggesting, in contrast to the electrostatic studies outlined above, that oxyanion hole stabilization is not primarily important for catalysis.[21]
| Wild-Type KSI Reaction Kinetics on 5-Androstenedione[22] | |
|---|---|
| kcat (s−1) | 3.0 x 104 |
| Km (μM) | 123 |
| kcat/Km (M−1s−1) | 2.4 x 108 |
Asp-38 general acidic/basic activity and effective molarity was probed by the Herschlag group through site-directed mutagenesis and exogenous base rescue.[23] Asp-38 was mutated to Gly, nullifying catalytic activity, and exogenous rescue was attempted with carboxylates of varying size and molarity. By calculating the concentration of base needed for full rescue, the Herschlag group determined the effective molarity of Asp-38 in KSI (6400 M). Thus, Asp-38 is critical for KSI catalysis.[23]
Sigala et al. found that solvent exclusion and replacement by the remote hydrophobic steroid rings negligibly alter the electrostatic environment within the KSI oxyanion hole.[24] In addition, ligand binding does not grossly alter the conformations of backbone and side chain groups observed in X-ray structures of PI KSI. However, NMR and UV studies suggest that steroid binding restricts the motions of several active-site groups, including Tyr-16.[25][26] Recently, the Herschlag group proposed that remote binding of hydrophobic regions of the substrate to distal portions of the active site contribute to KSI catalysis (>5 kcal/mol).[27] A 4-ring substrate reacted 27,000 times faster than a single ring substrate indicating the importance of distal active site binding motifs. This activity ratio persists throughout mutagenesis of residues important to oxyanion hole stabilization, implying that distal binding is what accounts for the large aforementioned reactivity difference.[27]
Numerous physical changes occur upon steroid binding within the KSI active site. In the free enzyme an ordered water molecule is positioned within hydrogen-bonding distance of Tyr-16 (the PI equivalent of TI KSI Tyr-14) and Asp-103 (the PI equivalent of TI KSI Asp-99).[28] This and additional disordered water molecules present within the unliganded active site are displaced upon steroid binding and are substantially excluded by the dense constellation of hydrophobic residues that pack around the bound, hydrophobic steroid skeleton.[28][25]
As stated above, the degree to which various factors contribute to KSI catalysis is still debated.
Function
KSI occurs in animal tissues concerned with steroid hormone biosynthesis, such as the adrenal, testis, and ovary.[29] KSI in Comamomas testosteroni is used in the degradation pathway of steroids, allowing this bacteria to utilize steroids containing a double bond at Δ5, such as testosterone, as its sole source of carbon.[30] In mammals, transfer of a double bond at Δ5 to Δ4 is catalyzed by 3-β-hydroxy-Δ5-steroid dehydrogenase at the same time as the dehydroxylation of 3-β-hydroxyl group to ketone group,[31] while in C. testosteroni and P. putida, Δ5,3-ketosteroid isomerase just transfers a double bond at Δ5 of 3-ketosteroid to Δ4.[32]
A Δ5-3-ketosteroid isomerase-disrupted mutant of strain TA441 can grow on dehydroepiandrosterone, which has a double bond at Δ5, but cannot grow on epiandrosterone, which lacks a double bond at Δ5, indicating that C. testosteroni KSI is responsible for transfer of the double bond from Δ5 to Δ4 and transfer of the double bond by hydrogenation at Δ5 and following dehydrogenation at Δ4 is not possible.[33]
Model enzyme
KSI has been used as a model system to test different theories to explain how enzymes achieve their catalytic efficiency. Low-barrier hydrogen bonds and unusual pKa values for the catalytic residues have been proposed as the basis for the fast action of KSI.[10][15] Gerlt and Gassman proposed the formation of unusually short, strong hydrogen bonds between KSI oxanion hole and the reaction intermediate as a means of catalytic rate enhancement.[34][35] In their model, high-energy states along the reaction coordinate are specifically stabilized by the formation of these bonds. Since then, the catalytic role of short, strong hydrogen bonds has been debated.[36][37] Another proposal explaining enzyme catalysis tested through KSI is the geometrical complementarity of the active site to the transition state, which proposes the active site electrostatics is complementary to the substrate transition state.[8]
KSI has also been a model system for studying protein folding. Kim et al. studied the effect of folding and tertiary structure on the function of KSI.[9]
References
- ↑ PDB: 3VSY; "Incorporation of rapid thermodynamic data in fragment-based drug discovery". Journal of Medicinal Chemistry 56 (5): 2155–9. March 2013. doi:10.1021/jm301603n. PMID 23419007.
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 "Enzymatic mechanisms for catalysis of enolization: ketosteroid isomerase". Bioorganic Chemistry 32 (5): 341–53. October 2004. doi:10.1016/j.bioorg.2004.06.005. PMID 15381400.
- ↑ "Crystal structure and enzyme mechanism of Delta 5-3-ketosteroid isomerase from Pseudomonas testosteroni". Biochemistry 37 (23): 8325–30. June 1998. doi:10.1021/bi9801614. PMID 9622484.
- ↑ "Activation of delta5-3-ketosteroid isomerase of bovine adrenal microsomes by serum albumins". Biochemical and Biophysical Research Communications 88 (3): 1158–66. June 1979. doi:10.1016/0006-291X(79)91530-4. PMID 465075.
- ↑ "Role of reduced glutathione in the delta(5)-3-kitosteroid isomerase reaction of liver". Biochemical and Biophysical Research Communications 69 (4): 1073–9. April 1976. doi:10.1016/0006-291X(76)90482-4. PMID 6023.
- ↑ Talalay, Paul; Benson, Ann M (1972). "Δ5-3-Ketosteroid Isomerase". in Boyer, Paul D.. The Enzymes. 6 (3rd ed.). Academic Press. pp. 591–618. ISBN 978-0-12-122706-7. https://books.google.com/books?id=_YP9Qu299UwC&pg=PA591.
- ↑ 7.0 7.1 "A proficient enzyme". Science 267 (5194): 90–3. January 1995. doi:10.1126/science.7809611. PMID 7809611. Bibcode: 1995Sci...267...90R.
- ↑ 8.0 8.1 "Testing electrostatic complementarity in enzyme catalysis: hydrogen bonding in the ketosteroid isomerase oxyanion hole". PLOS Biology 4 (4): e99. April 2006. doi:10.1371/journal.pbio.0040099. PMID 16602823.

- ↑ 9.0 9.1 9.2 9.3 9.4 "Roles of dimerization in folding and stability of ketosteroid isomerase from Pseudomonas putida biotype B". Protein Science 10 (4): 741–52. April 2001. doi:10.1110/ps.18501. PMID 11274465.
- ↑ 10.0 10.1 "Detection of large pKa perturbations of an inhibitor and a catalytic group at an enzyme active site, a mechanistic basis for catalytic power of many enzymes". The Journal of Biological Chemistry 275 (52): 41100–6. December 2000. doi:10.1074/jbc.M007561200. PMID 11007792.
- ↑ 11.0 11.1 11.2 11.3 "Solution structure of 3-oxo-delta5-steroid isomerase". Science 276 (5311): 415–8. April 1997. doi:10.1126/science.276.5311.415. PMID 9103200.
- ↑ "Catalytic mechanism of an active-site mutant (D38N) of delta 5-3-ketosteroid isomerase. Direct spectroscopic evidence for dienol intermediates". Biochemistry 30 (20): 4991–7. May 1991. doi:10.1021/bi00234a022. PMID 2036366.
- ↑ "Substituent effects on the binding of phenols to the D38N mutant of 3-oxo-delta5-steroid isomerase. A probe for the nature of hydrogen bonding to the intermediate". Biochemistry 37 (2): 700–5. January 1998. doi:10.1021/bi972262s. PMID 9425094.
- ↑ "A Critical Test of the Electrostatic Contribution to Catalysis with Noncanonical Amino Acids in Ketosteroid Isomerase". Journal of the American Chemical Society 138 (36): 11890–5. September 2016. doi:10.1021/jacs.6b06843. PMID 27545569.
- ↑ 15.0 15.1 "Proton affinity of the oxyanion hole in the active site of ketosteroid isomerase". Biochemistry 49 (12): 2725–31. March 2010. doi:10.1021/bi100074s. PMID 20143849.
- ↑ 16.0 16.1 "Ketosteroid isomerase provides further support for the idea that enzymes work by electrostatic preorganization". Proceedings of the National Academy of Sciences of the United States of America 107 (9): 4075–80. March 2010. doi:10.1073/pnas.0914579107. PMID 20150513. Bibcode: 2010PNAS..107.4075K.
- ↑ "Extreme electric fields power catalysis in the active site of ketosteroid isomerase". Science 346 (6216): 1510–4. December 2014. doi:10.1126/science.1259802. PMID 25525245. Bibcode: 2014Sci...346.1510F.
- ↑ "High-resolution crystal structures of delta5-3-ketosteroid isomerase with and without a reaction intermediate analogue" (in en). Biochemistry 36 (46): 14030–6. November 1997. doi:10.1021/bi971546+. PMID 9369474.
- ↑ "Electrophilic assistance by Asp-99 of 3-oxo-Delta 5-steroid isomerase". Biochemistry 37 (29): 10499–506. July 1998. doi:10.1021/bi980099a. PMID 9671521.
- ↑ "Hydrogen bonding at the active site of delta 5-3-ketosteroid isomerase". Biochemistry 36 (48): 14616–26. December 1997. doi:10.1021/bi971549m. PMID 9398180.
- ↑ 21.0 21.1 "Using unnatural amino acids to probe the energetics of oxyanion hole hydrogen bonds in the ketosteroid isomerase active site". Journal of the American Chemical Society 136 (21): 7643–54. May 2014. doi:10.1021/ja413174b. PMID 24787954.
- ↑ "Insights into the catalytic mechanism and active-site environment of Comamonas testosteroni delta 5-3-ketosteroid isomerase as revealed by site-directed mutagenesis of the catalytic base aspartate-38". Biochemistry 34 (43): 14245–53. October 1995. doi:10.1021/bi00043a032. PMID 7578024.
- ↑ 23.0 23.1 "Evaluation of the Catalytic Contribution from a Positioned General Base in Ketosteroid Isomerase". Journal of the American Chemical Society 138 (31): 9902–9. August 2016. doi:10.1021/jacs.6b04796. PMID 27410422.
- ↑ "Do ligand binding and solvent exclusion alter the electrostatic character within the oxyanion hole of an enzymatic active site?". Journal of the American Chemical Society 129 (40): 12104–5. October 2007. doi:10.1021/ja075605a. PMID 17854190.
- ↑ 25.0 25.1 "Ultraviolet spectroscopic evidence for decreased motion of the active site tyrosine residue of delta 5-3-ketosteroid isomerase by steroid binding". Biochemistry 34 (19): 6562–72. May 1995. doi:10.1021/bi00019a038. PMID 7756287.
- ↑ "13C NMR relaxation studies of backbone and side chain motion of the catalytic tyrosine residue in free and steroid-bound delta 5-3-ketosteroid isomerase". Biochemistry 35 (5): 1525–32. February 1996. doi:10.1021/bi9525381. PMID 8634283.
- ↑ 27.0 27.1 "Determining the catalytic role of remote substrate binding interactions in ketosteroid isomerase". Proceedings of the National Academy of Sciences of the United States of America 106 (34): 14271–5. August 2009. doi:10.1073/pnas.0901032106. PMID 19706511. Bibcode: 2009PNAS..10614271S.
- ↑ 28.0 28.1 "High-resolution crystal structures of delta5-3-ketosteroid isomerase with and without a reaction intermediate analogue". Biochemistry 36 (46): 14030–6. November 1997. doi:10.1021/bi971546+. PMID 9369474.
- ↑ "The preparation and properties of crystalline delta5-3-ketosteroid isomerase". The Journal of Biological Chemistry 237 (5): 1500–6. May 1962. doi:10.1016/S0021-9258(19)83730-4. PMID 14454546. http://www.jbc.org/cgi/pmidlookup?view=long&pmid=14454546.
- ↑ "Oxidative degradation of testosterone by adaptive enzymes". Nature 170 (4328): 620–1. October 1952. doi:10.1038/170620a0. PMID 13002385. Bibcode: 1952Natur.170..620T.
- ↑ "Characterization of human 3 beta-hydroxysteroid dehydrogenase/delta 5-delta 4-isomerase gene and its expression in mammalian cells". The Journal of Biological Chemistry 267 (5): 3551. February 1992. doi:10.1016/S0021-9258(19)50764-5. PMID 1737804. http://www.jbc.org/content/267/5/3551.long.
- ↑ "Steroid degradation in Comamonas testosteroni". The Journal of Steroid Biochemistry and Molecular Biology 129 (1–2): 4–14. March 2012. doi:10.1016/j.jsbmb.2010.10.008. PMID 21056662.
- ↑ "Steroid degradation genes in Comamonas testosteroni TA441: Isolation of genes encoding a Δ4(5)-isomerase and 3α- and 3β-dehydrogenases and evidence for a 100 kb steroid degradation gene hot spot". The Journal of Steroid Biochemistry and Molecular Biology 122 (4): 253–63. October 2010. doi:10.1016/j.jsbmb.2010.06.002. PMID 20554032.
- ↑ "Understanding the rates of certain enzyme-catalyzed reactions: proton abstraction from carbon acids, acyl-transfer reactions, and displacement reactions of phosphodiesters". Biochemistry 32 (45): 11943–52. November 1993. doi:10.1021/bi00096a001. PMID 8218268.
- ↑ "An explanation for rapid enzyme-catalyzed proton abstraction from carbon acids: importance of late transition states in concerted mechanisms.". Biochemistry 115 (24): 11552–11568. December 1993. doi:10.1021/ja00077a062.
- ↑ "On low-barrier hydrogen bonds and enzyme catalysis". Science 269 (5220): 102–6. July 1995. doi:10.1126/science.7661987. PMID 7661987. Bibcode: 1995Sci...269..102W.
- ↑ "Short strong hydrogen bonds: can they explain enzymic catalysis?". Chemistry & Biology 3 (3): 163–70. March 1996. doi:10.1016/s1074-5521(96)90258-6. PMID 8807842.
Further reading
- "Evidence for the presence of 17-hydroxypregnenedione isomerase in beef adrenal cortex". Biochimica et Biophysica Acta (BBA) - General Subjects 111 (1): 306–12. November 1965. doi:10.1016/0304-4165(65)90497-6. PMID 5867327.
- "Crystalline Delta 5-3-ketosteroid isomerase". The Journal of Biological Chemistry 235: PC1–2. January 1960. doi:10.1016/S0021-9258(18)69620-6. PMID 14404954. http://www.jbc.org/cgi/pmidlookup?view=long&pmid=14404954.
- "Enzymic isomerization of delta5-3-ketosteroids". Biochimica et Biophysica Acta 18 (2): 300–1. October 1955. doi:10.1016/0006-3002(55)90079-2. PMID 13276386.
- Steroid+Isomerases at the US National Library of Medicine Medical Subject Headings (MeSH)
 |