Biology:Twitching motility
Template:Microbial and microbot movement
Twitching motility is a form of crawling bacterial motility used to move over surfaces. Twitching is mediated by the activity of hair-like filaments called type IV pili which extend from the cell's exterior, bind to surrounding solid substrates, and retract, pulling the cell forwards in a manner similar to the action of a grappling hook.[1][2][3] The name twitching motility is derived from the characteristic jerky and irregular motions of individual cells when viewed under the microscope.[4] It has been observed in many bacterial species, but is most well studied in Pseudomonas aeruginosa, Neisseria gonorrhoeae and Myxococcus xanthus. Active movement mediated by the twitching system has been shown to be an important component of the pathogenic mechanisms of several species.[2]
Mechanisms
Pilus structure
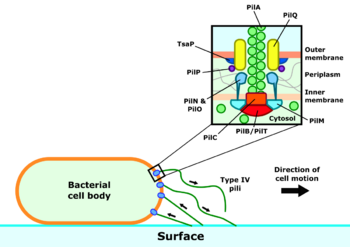
The type IV pilus complex consists of both the pilus itself and the machinery required for its construction and motor activity. The pilus filament is largely composed of the PilA protein, with more uncommon minor pilins at the tip. These are thought to play a role in initiation of pilus construction.[5] Under normal conditions, the pilin subunits are arranged as a helix with five subunits in each turn,[5][6] but pili under tension are able to stretch and rearrange their subunits into a second configuration with around 1 2⁄3 subunits per turn.[7]
Three subcomplexes form the apparatus responsible for assembling and retracting the type IV pili.[8] The core of this machinery is the motor subcomplex, consisting of the PilC protein and the cytosolic ATPases PilB and PilT. These ATPases drive pilus extension or retraction respectively, depending on which of the two is currently bound to the pilus complex. Surrounding the motor complex is the alignment subcomplex, formed from the PilM, PilN, PilO and PilP proteins. These proteins form a bridge between the inner and outer membranes and create a link between the inner membrane motor subcomplex and the outer membrane secretion subcomplex. This consists of a pore formed from the PilQ protein, through which the assembled pilus can exit the cell.[9]
Regulation
Regulatory proteins associated with the twitching motility system have strong sequence and structural similarity to those that regulate bacterial chemotaxis using flagellae.[2][10] In P. aeruginosa for example, a total of four homologous chemosensory pathways are present, three regulating swimming motility and one regulating twitching motility.[11] These chemotactic systems allow cells to regulate twitching so as to move towards chemoattractants such as phospholipids and fatty acids.[12] In contrast to the run-and-tumble model of chemotaxis associated with flagellated cells however, movement towards chemoattractants in twitching cells appears to be mediated via regulation of the timing of directional reversals.[13]
Motility patterns

Twitching motility is capable of driving the movement of individual cells.[1][13] The pattern of motility that results is highly dependent upon cell shape and the distribution of pili over the cell surface.[14] In N. gonorrhoeae for example, the roughly spherical cell shape and uniform distribution of pili results in cells adopting a 2D random walk over the surface they are attached to.[15] In contrast, species such as P. aeruginosa and M. xanthus exist as elongated rods with pili localised at their poles, and show much greater directional persistence during crawling due to the resulting bias in force generation direction.[16] P. aeruginosa and M. xanthus are also able to reverse direction during crawling by switching the pole of pilus localization.[13][14] Type IV pili also mediate a form of walking motility in P. aeruginosa, where pili are used to pull the cell rod into a vertical orientation and move it at much higher speeds than during horizontal crawling motility.[16][17]
The existence of many pili pulling simultaneously on the cell body results in a balance of forces determining the movement of the cell body. This is known as the tug-of-war model of twitching motility.[14][15] Sudden changes in the balance of forces caused by detachment or release of individual pili results in a fast jerk (or 'slingshot') that combines fast rotational and lateral movements, in contrast to the slower lateral movements seen during the longer periods between slingshots.[18]
Roles
Pathogenesis
Both presence of type IV pili and active pilar movement appear to be important contributors to the pathogenicity of several species.[8] In P. aeruginosa, loss of pilus retraction results in a reduction of bacterial virulence in pneumonia[19] and reduces colonisation of the cornea.[20] Some bacteria are also able to twitch along vessel walls against the direction of fluid flow within them,[21] which is thought to permit colonisation of otherwise inaccessible sites in the vasculatures of plants and animals.
Bacterial cells can also be targeted by twitching: during the cell invasion phase of the lifecycle of Bdellovibrio, type IV pili are used by cells to pull themselves through gaps formed in the cell wall of prey bacteria.[22] Once inside, the Bdellovibrio are able to use the host cell's resources to grow and reproduce, eventually lysing the cell wall of the prey bacterium and escaping to invade other cells.
Biofilms
Twitching motility is also important during the formation of biofilms.[8] During biofilm establishment and growth, motile bacteria are able to interact with secreted extracellular polymeric substances (EPSs) such as Psl, alginate and extracellular DNA.[23] As they encounter sites of high EPS deposition, P. aeruginosa cells slow down, accumulate and deposit further EPS components. This positive feedback is an important initiating factor for the establishment of microcolonies, the precursors to fully fledged biofilms.[24] In addition, once biofilms have become established, their twitching-mediated spread is facilitated and organised by components of the EPS.[25]
Twitching can also influence the structure of biofilms. During their establishment, twitching-capable cells are able to crawl on top of cells lacking twitching motility and dominate the fast-growing external surface of the biofilm.[23][26]
Taxonomic distribution and evolution
Type IV pili and related structures can be found across almost all phyla of Bacteria and Archaea,[27] however definitive twitching motility has been shown in a more limited range of prokaryotes. Most well studied and wide spread are the twitching Pseudomonadota, such as Neisseria gonorrhoeae, Myxococcus xanthus and Pseudomonas aeruginosa.[14][8] Nevertheless, twitching has been observed in other phyla as well. For example, twitching motility has been observed in the cyanobacterium Synechocystis,[28] as well as the gram-positive Bacillota Streptococcus sanguinis.[29]
Other structures and systems closely related to type IV pili have also been observed in prokaryotes. In Archea, for example, bundles of type IV-like filaments have been observed to form helical structures similar in both form and function to the bacterial flagellum. These swimming associated structures have been termed archaella.[30] Also closely related to the type IV pilus is the type II secretion system,[31] itself widely distributed amongst gram-negative bacteria. In this secretion system, cargo destined for export is associated with tips of type IV-like pseudopili in the periplasm. Extension of the pseudopili through secretin proteins similar to PilQ permits these cargo proteins to cross the outer membrane and enter the extracellular environment.
Because of this wide but patchy distribution of type IV pilus-like machinery, it has been suggested that the genetic material encoding it has been transferred between species via horizontal gene transfer following its initial development in a single species of Pseudomonadota.[6]
See also
References
- ↑ 1.0 1.1 Skerker, J. M.; Berg, H. C. (2001-06-05). "Direct observation of extension and retraction of type IV pili". Proceedings of the National Academy of Sciences of the United States of America 98 (12): 6901–6904. doi:10.1073/pnas.121171698. ISSN 0027-8424. PMID 11381130. Bibcode: 2001PNAS...98.6901S.
- ↑ 2.0 2.1 2.2 Mattick, John S. (2002). "Type IV pili and twitching motility". Annual Review of Microbiology 56: 289–314. doi:10.1146/annurev.micro.56.012302.160938. ISSN 0066-4227. PMID 12142488.
- ↑ Merz, A. J.; So, M.; Sheetz, M. P. (2000-09-07). "Pilus retraction powers bacterial twitching motility". Nature 407 (6800): 98–102. doi:10.1038/35024105. ISSN 0028-0836. PMID 10993081. Bibcode: 2000Natur.407...98M.
- ↑ Henrichsen, J. (December 1972). "Bacterial surface translocation: a survey and a classification". Bacteriological Reviews 36 (4): 478–503. doi:10.1128/br.36.4.478-503.1972. ISSN 0005-3678. PMID 4631369.
- ↑ 5.0 5.1 Leighton, Tiffany L.; Buensuceso, Ryan N. C.; Howell, P. Lynne; Burrows, Lori L. (2015-11-01). "Biogenesis of Pseudomonas aeruginosa type IV pili and regulation of their function" (in en). Environmental Microbiology 17 (11): 4148–4163. doi:10.1111/1462-2920.12849. ISSN 1462-2920. PMID 25808785.
- ↑ 6.0 6.1 Nudleman, Eric; Kaiser, Dale (2004). "Pulling together with type IV pili". Journal of Molecular Microbiology and Biotechnology 7 (1–2): 52–62. doi:10.1159/000077869. ISSN 1464-1801. PMID 15170403.
- ↑ Biais, Nicolas; Higashi, Dustin L.; Brujic, Jasna; So, Magdalene; Sheetz, Michael P. (2010-06-22). "Force-dependent polymorphism in type IV pili reveals hidden epitopes". Proceedings of the National Academy of Sciences of the United States of America 107 (25): 11358–11363. doi:10.1073/pnas.0911328107. ISSN 1091-6490. PMID 20534431. Bibcode: 2010PNAS..10711358B.
- ↑ 8.0 8.1 8.2 8.3 Burrows, Lori L. (2012). "Pseudomonas aeruginosa twitching motility: type IV pili in action". Annual Review of Microbiology 66: 493–520. doi:10.1146/annurev-micro-092611-150055. ISSN 1545-3251. PMID 22746331.
- ↑ Chang, Yi-Wei; Rettberg, Lee A.; Treuner-Lange, Anke; Iwasa, Janet; Søgaard-Andersen, Lotte; Jensen, Grant J. (2016-03-11). "Architecture of the type IVa pilus machine". Science 351 (6278). doi:10.1126/science.aad2001. ISSN 1095-9203. PMID 26965631. Bibcode: 2016BpJ...110..468C.
- ↑ Sampedro, Inmaculada; Parales, Rebecca E.; Krell, Tino; Hill, Jane E. (January 2015). "Pseudomonas chemotaxis". FEMS Microbiology Reviews 39 (1): 17–46. doi:10.1111/1574-6976.12081. ISSN 1574-6976. PMID 25100612.
- ↑ Ortega, Davi R.; Fleetwood, Aaron D.; Krell, Tino; Harwood, Caroline S.; Jensen, Grant J.; Zhulin, Igor B. (2017-11-13). "Assigning chemoreceptors to chemosensory pathways in Pseudomonas aeruginosa". Proceedings of the National Academy of Sciences of the United States of America 114 (48): 12809–12814. doi:10.1073/pnas.1708842114. ISSN 1091-6490. PMID 29133402.
- ↑ Miller, Rhea M.; Tomaras, Andrew P.; Barker, Adam P.; Voelker, Dennis R.; Chan, Edward D.; Vasil, Adriana I.; Vasil, Michael L. (2008-06-01). "Pseudomonas aeruginosa Twitching Motility-Mediated Chemotaxis towards Phospholipids and Fatty Acids: Specificity and Metabolic Requirements" (in en). Journal of Bacteriology 190 (11): 4038–4049. doi:10.1128/jb.00129-08. ISSN 0021-9193. PMID 18390654.
- ↑ 13.0 13.1 13.2 Oliveira, Nuno M.; Foster, Kevin R.; Durham, William M. (2016-06-07). "Single-cell twitching chemotaxis in developing biofilms" (in en). Proceedings of the National Academy of Sciences 113 (23): 6532–6537. doi:10.1073/pnas.1600760113. ISSN 0027-8424. PMID 27222583.
- ↑ 14.0 14.1 14.2 14.3 Maier, Berenike; Wong, Gerard C. L. (December 2015). "How Bacteria Use Type IV Pili Machinery on Surfaces". Trends in Microbiology 23 (12): 775–788. doi:10.1016/j.tim.2015.09.002. ISSN 1878-4380. PMID 26497940.
- ↑ 15.0 15.1 Marathe, Rahul; Meel, Claudia; Schmidt, Nora C.; Dewenter, Lena; Kurre, Rainer; Greune, Lilo; Schmidt, M. Alexander; Müller, Melanie J. I. et al. (2014-05-07). "Bacterial twitching motility is coordinated by a two-dimensional tug-of-war with directional memory". Nature Communications 5: 3759. doi:10.1038/ncomms4759. ISSN 2041-1723. PMID 24806757. Bibcode: 2014NatCo...5.3759M.
- ↑ 16.0 16.1 Conrad, Jacinta C.; Gibiansky, Maxsim L.; Jin, Fan; Gordon, Vernita D.; Motto, Dominick A.; Mathewson, Margie A.; Stopka, Wiktor G.; Zelasko, Daria C. et al. (2011-04-06). "Flagella and pili-mediated near-surface single-cell motility mechanisms in P. aeruginosa". Biophysical Journal 100 (7): 1608–1616. doi:10.1016/j.bpj.2011.02.020. ISSN 1542-0086. PMID 21463573. Bibcode: 2011BpJ...100.1608C.
- ↑ Gibiansky, Maxsim L.; Conrad, Jacinta C.; Jin, Fan; Gordon, Vernita D.; Motto, Dominick A.; Mathewson, Margie A.; Stopka, Wiktor G.; Zelasko, Daria C. et al. (2010-10-08). "Bacteria use type IV pili to walk upright and detach from surfaces". Science 330 (6001): 197. doi:10.1126/science.1194238. ISSN 1095-9203. PMID 20929769. Bibcode: 2010Sci...330..197G.
- ↑ Jin, Fan; Conrad, Jacinta C.; Gibiansky, Maxsim L.; Wong, Gerard C. L. (2011-08-02). "Bacteria use type-IV pili to slingshot on surfaces". Proceedings of the National Academy of Sciences of the United States of America 108 (31): 12617–12622. doi:10.1073/pnas.1105073108. ISSN 1091-6490. PMID 21768344.
- ↑ Comolli, J. C.; Hauser, A. R.; Waite, L.; Whitchurch, C. B.; Mattick, J. S.; Engel, J. N. (July 1999). "Pseudomonas aeruginosa gene products PilT and PilU are required for cytotoxicity in vitro and virulence in a mouse model of acute pneumonia". Infection and Immunity 67 (7): 3625–3630. doi:10.1128/IAI.67.7.3625-3630.1999. ISSN 0019-9567. PMID 10377148.
- ↑ Zolfaghar, Irandokht; Evans, David J.; Fleiszig, Suzanne M. J. (2003-09-01). "Twitching Motility Contributes to the Role of Pili in Corneal Infection Caused by Pseudomonas aeruginosa" (in en). Infection and Immunity 71 (9): 5389–5393. doi:10.1128/iai.71.9.5389-5393.2003. ISSN 0019-9567. PMID 12933890.
- ↑ Shen, Yi; Siryaporn, Albert; Lecuyer, Sigolene; Gitai, Zemer; Stone, Howard A. (2012-07-03). "Flow directs surface-attached bacteria to twitch upstream". Biophysical Journal 103 (1): 146–151. doi:10.1016/j.bpj.2012.05.045. ISSN 1542-0086. PMID 22828341. Bibcode: 2012BpJ...103..146S.
- ↑ Sockett, Renee Elizabeth (2009). "Predatory lifestyle of Bdellovibrio bacteriovorus". Annual Review of Microbiology 63: 523–539. doi:10.1146/annurev.micro.091208.073346. ISSN 1545-3251. PMID 19575566.
- ↑ 23.0 23.1 Parsek, Matthew R.; Tolker-Nielsen, Tim (December 2008). "Pattern formation in Pseudomonas aeruginosa biofilms". Current Opinion in Microbiology 11 (6): 560–566. doi:10.1016/j.mib.2008.09.015. ISSN 1879-0364. PMID 18935979.
- ↑ Zhao, Kun; Tseng, Boo Shan; Beckerman, Bernard; Jin, Fan; Gibiansky, Maxsim L.; Harrison, Joe J.; Luijten, Erik; Parsek, Matthew R. et al. (2013-05-16). "Psl trails guide exploration and microcolony formation in Pseudomonas aeruginosa biofilms". Nature 497 (7449): 388–391. doi:10.1038/nature12155. ISSN 1476-4687. PMID 23657259. Bibcode: 2013Natur.497..388Z.
- ↑ Gloag, Erin S.; Turnbull, Lynne; Huang, Alan; Vallotton, Pascal; Wang, Huabin; Nolan, Laura M.; Mililli, Lisa; Hunt, Cameron et al. (2013-07-09). "Self-organization of bacterial biofilms is facilitated by extracellular DNA". Proceedings of the National Academy of Sciences of the United States of America 110 (28): 11541–11546. doi:10.1073/pnas.1218898110. ISSN 1091-6490. PMID 23798445. Bibcode: 2013PNAS..11011541G.
- ↑ Klausen, Mikkel; Aaes-Jørgensen, Anders; Molin, Søren; Tolker-Nielsen, Tim (2003-10-01). "Involvement of bacterial migration in the development of complex multicellular structures in Pseudomonas aeruginosa biofilms" (in en). Molecular Microbiology 50 (1): 61–68. doi:10.1046/j.1365-2958.2003.03677.x. ISSN 1365-2958. PMID 14507363.
- ↑ Berry, Jamie-Lee; Pelicic, Vladimir (January 2015). "Exceptionally widespread nanomachines composed of type IV pilins: the prokaryotic Swiss Army knives". FEMS Microbiology Reviews 39 (1): 134–154. doi:10.1093/femsre/fuu001. ISSN 1574-6976. PMID 25793961.
- ↑ Bhaya, D.; Bianco, N. R.; Bryant, D.; Grossman, A. (August 2000). "Type IV pilus biogenesis and motility in the cyanobacterium Synechocystis sp. PCC6803". Molecular Microbiology 37 (4): 941–951. doi:10.1046/j.1365-2958.2000.02068.x. ISSN 0950-382X. PMID 10972813.
- ↑ Gurung, Ishwori; Spielman, Ingrid; Davies, Mark R.; Lala, Rajan; Gaustad, Peter; Biais, Nicolas; Pelicic, Vladimir (2016-01-01). "Functional analysis of an unusual type IV pilus in the Gram-positive Streptococcus sanguinis" (in en). Molecular Microbiology 99 (2): 380–392. doi:10.1111/mmi.13237. ISSN 1365-2958. PMID 26435398.
- ↑ Ng, Sandy Y. M.; Chaban, Bonnie; Jarrell, Ken F. (2006). "Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications". Journal of Molecular Microbiology and Biotechnology 11 (3–5): 167–191. doi:10.1159/000094053. ISSN 1464-1801. PMID 16983194.
- ↑ Peabody, Christopher R.; Chung, Yong Joon; Yen, Ming-Ren; Vidal-Ingigliardi, Dominique; Pugsley, Anthony P.; Saier, Milton H. (November 2003). "Type II protein secretion and its relationship to bacterial type IV pili and archaeal flagella". Microbiology 149 (Pt 11): 3051–3072. doi:10.1099/mic.0.26364-0. ISSN 1350-0872. PMID 14600218.
 |
