Chemistry:Sequence-controlled polymer
}}
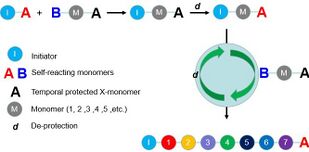
A sequence-controlled polymer is a macromolecule, in which the sequence of monomers is controlled to some degree.[1][2] This control can be absolute but not necessarily. In other words, a sequence-controlled polymer can be uniform (its dispersity Ð is equal to 1) or non-uniform (Ð>1). For example, an alternating copolymer synthesized by radical polymerization is a sequence-controlled polymer, even if it is also a non-uniform polymer, in which chains have different chain-lengths and slightly different compositions.[2] A biopolymer (for example a protein) with a perfectly defined primary structure is also a sequence-controlled polymer. However, in the case of uniform macromolecules, the term sequence-defined polymer can also be used.
With comparison to traditional polymers, the composition of sequence-controlled polymers can be precisely defined via chemical synthetic methods, such as multicomponent reactions, click reactions etc. Such tunable polymerizing manner endows sequence-controlled polymers with particular properties and thereby, sequence-controlled polymers-based applications (e.g. information storage,[3] biomaterials,[3] nanomaterials[4] etc.) are developed.
In nature, DNA, RNA, proteins and other macromolecules can also be recognized as sequence-controlled polymers for their well-ordered structural skeletons. DNA, based on A-T, C-G base pairs, are formed in well-aligned sequences. Through precise sequences of DNA, 20 amino acids are able to generate sequential peptide chains with three-dimensional structures by virtue of transcription and translation process. These ordered sequences of different constituents endow organisms with complicated and diverse functions.
Synthetic methods
Traditional polymers are usually consist of one repeating unit or several repeating units, arranged in random sequences. Sequence-controlled polymers are composed of different repeating units, which are arranged in an ordered manner. In order to control the sequence, various kinds of synthetic methodologies are developed.
Sequence-controlled biological polymerization

DNA, RNA and proteins are most common sequence-controlled polymers in living creatures. Inspired by them, polymerization methods, utilizing DNA or RNA as templates to control sequences of polymer, are developed. At first, taking DNA or RNA as templates, scientists developed a series of peptide nucleic acid (PNA)-based polymers, without using DNA polymerases.[6][7] But this method is limited to polymerization scale and yield.[1] After that, polymerase chain reaction (PCR) is developed, which currently is still the most extensively used sequence-regulated method.[8] By employing enzymes, the yields and scales are greatly increased, but the specificity of enzymes towards natural peptides limits this technique to a certain degree. Nowadays, more attention is paid to utilization of ribosomes to directly mimic the transcription and translation process.[9] This technology called protein engineering is considered as the most promising biological polymerization method for synthesis of sequence-controlled polymers.
Sequence-controlled chemical polymerizations
Other than biological polymerization methods, scientists have also developed numerous chemical synthetic methods for sequence-controlled polymers. Compared with biological polymerization, chemical polymerization can provide better diversity but most of the chemical methods cannot offer the efficiency and specificity of biological methods.[1]
Solid-phase synthesis
One of the chemical polymerization methods is solid-phase synthesis, which can be used to synthesize peptides consisted of natural and non-natural amino acids. In this method, the monomers are attached to the polymer chain via amidation between carbonyl group and amino group. For purpose of sequence control, the amino groups are usually protected by 9-fluorenylmethyloxycarbonyl group (Fmoc) and t-butyloxycarbonyl (Boc),[10] which can be removed under base and acid environment respectively to participate into next-round chain elongation.
Sequence-controlled radical polymerization
Radical polymerization is one of the most commonly used polymerization methods. About 50% of commercially available polymers are synthesized via radical polymerization.[11] However, the disadvantages of this method are apparent that sequences and polymeric features cannot be well modulated. To overcome these constraints, scientists optimized the employed protocols. The first reported example was the time-controlled sequential addition of highly-reactive N-substituted maleimides in the atom transfer radical polymerization of styrene, which led to programmed sequences of functional monomers.[12] The development of single-molecule addition into atom-transfer radical polymerization (ATRP), which enhances the sequence control of radical polymerization was also reported.[13] Other solutions include the use of intermediate purification steps to isolate the desired oligomer sequence in between subsequent reversible addition−fragmentation chain-transfer polymerization (RAFT-polymerizations). Both flash column chromatography[14] and recycling size exclusion chromatography[15] have been proven successful in this regard. RAFT single unit monomer insertion (SUMI) is recently developed as an emerging technology for precise control of monomer sequence.[16]
Sequence controlled non-radical polymerization
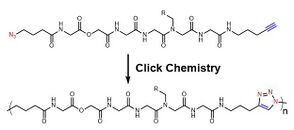
For the intrinsic shortages of radical polymerization for sequence-controlled polymers, other non-radical polymerizations are also developed. Within those non-radical methods, azide-alkyne cycloaddition (also known as click reaction),[18] olefin metathesis[19] among others are utilized to construct sequence-controlled polymers. Depending on these specific chemical reactions, monomers are accurately added to the polymer chain and a well-ordered chain is accomplished stepwise. Meanwhile, by applying multiple chemical reactions, chemists have also developed multi-component reactions[20] to accelerate the construction of polymer skeletons and also enhance variety. Beyond the aforementioned, there was a research group developing a molecule machine, which successfully achieve a sequence-controlled polymerization of oligopeptides.[21]
Methodology towards improving sequence control
The most important character of sequence-controlled polymers is its controllable sequence of polymer backbone. Nonetheless, to realize a precise sequence control and to regulate sequences in greater polymer backbones are also the most urgent issue, which needs to be addressed in the field of sequence-controlled polymers. Great efforts have been made in development and optimization of methods to improve the sequence-control properties of currently existed synthetic methods and also to further brand-new methods with better synthetic efficiency and sequence control.
Utilization of templates
One of the most significant character of sequence-controlled biosynthesis against other chemical synthetic methods is that the biomolecules (including DNA and RNA) can initiate their polymerization using highly programmed templates. Hence, biosynthetic methods, like PCR, are still considered one of the most cogent manner to develop sequence-controlled polymers.
Regulation of reactivity of monomer
To modulate the reactivity between monomer and growing polymeric chain is another approach to enhance sequence control.[22] The rationale for this method is that monomer has to be activated with first catalyst at beginning as a dormant species, which could then participate into polymerization as the second catalyst is introduced. A real example is utilization of HI as first catalyst and ZnI2 as second catalyst to achieve sequence-controlled polymerization of vinyl ethers and styrene derivatives.[23]
Recognition-oriented insertion

In this approach, a recognition site at polymer is offered to non-covalently anchor the monomer at polymer chain, which can subsequently go through a chemical insertion into polymeric backbone. One successful example demonstrates that methacrylic acid (monomer) can be radically incorporated into a backbone featuring a recognizable cationic site (protonated primary amine pendant).[24] Driven by this site-specific reaction, the sequence-controlled polymerization can be achieve by using a template adorned with differenrt recognizable pendants.
Properties and applications
The most distinguishable feature of sequence-controlled polymers is the well-ordered chains composed of different repeating units.[25] By encoding the repeating units, the correspondingly synthesized sequence-controlled polymer can be used for data storage.[26] To modify the monomer with some bioactive moieties, the obtained sequence-controlled polymer is able to treat diseases. The property of sequence control make sequence-controlled polymers an ideal platform to install various kinds of pendants (like drugs, catalyst), whereby diverse functions and applications can be realized.
References
- ↑ 1.0 1.1 1.2 Lutz, Jean-François; Ouchi, Makoto; Liu, David R.; Sawamoto, Mitsuo (2013-08-09). "Sequence-Controlled Polymers" (in en). Science 341 (6146). doi:10.1126/science.1238149. ISSN 0036-8075. PMID 23929982. Bibcode: 2013Sci...34138149L.
- ↑ 2.0 2.1 Lutz, Jean-François (2017-12-01). "Defining the Field of Sequence-Controlled Polymers" (in en). Macromolecular Rapid Communications 38 (24): n/a. doi:10.1002/marc.201700582. ISSN 1521-3927. PMID 29160615.
- ↑ 3.0 3.1 Schnitzler, Tobias; Herrmann, Andreas (2012-09-18). "DNA Block Copolymers: Functional Materials for Nanoscience and Biomedicine". Accounts of Chemical Research 45 (9): 1419–1430. doi:10.1021/ar200211a. ISSN 0001-4842. PMID 22726237. https://pure.rug.nl/ws/files/14534907/2012AccChemResSchnitzler.pdf.
- ↑ Aldaye, Faisal A.; Palmer, Alison L.; Sleiman, Hanadi F. (2008-09-26). "Assembling Materials with DNA as the Guide" (in en). Science 321 (5897): 1795–1799. doi:10.1126/science.1154533. ISSN 0036-8075. PMID 18818351. Bibcode: 2008Sci...321.1795A.
- ↑ Hibi, Yusuke; Ouchi, Makoto; Sawamoto, Mitsuo (2016-03-21). "A strategy for sequence control in vinyl polymers via iterative controlled radical cyclization" (in en). Nature Communications 7. doi:10.1038/ncomms11064. ISSN 2041-1723. PMID 26996881. Bibcode: 2016NatCo...711064H.
- ↑ Böhler, Christof; Nielsen, Peter E.; Orgel, Leslie E. (1995-08-17). "Template switching between PNA and RNA oligonucleotides" (in en). Nature 376 (6541): 578–581. doi:10.1038/376578a0. PMID 7543656. Bibcode: 1995Natur.376..578B.
- ↑ Kleiner, Ralph E.; Brudno, Yevgeny; Birnbaum, Michael E.; Liu, David R. (2008-04-01). "DNA-Templated Polymerization of Side-Chain-Functionalized Peptide Nucleic Acid Aldehydes". Journal of the American Chemical Society 130 (14): 4646–4659. doi:10.1021/ja0753997. ISSN 0002-7863. PMID 18341334. Bibcode: 2008JAChS.130.4646K.
- ↑ Saiki, R. K.; Gelfand, D. H.; Stoffel, S.; Scharf, S. J.; Higuchi, R.; Horn, G. T.; Mullis, K. B.; Erlich, H. A. (1988-01-29). "Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase" (in en). Science 239 (4839): 487–491. doi:10.1126/science.239.4839.487. ISSN 0036-8075. PMID 2448875. Bibcode: 1988Sci...239..487S. https://www.science.org/doi/10.1126/science.2448875.
- ↑ McGrath, Kevin P.; Fournier, Maurille J.; Mason, Thomas L.; Tirrell, David A. (1992-01-01). "Genetically directed syntheses of new polymeric materials. Expression of artificial genes encoding proteins with repeating -(AlaGly)3ProGluGly- elements". Journal of the American Chemical Society 114 (2): 727–733. doi:10.1021/ja00028a048. ISSN 0002-7863. Bibcode: 1992JAChS.114..727M.
- ↑ Palomo, Jose M. (2014-07-25). "Solid-phase peptide synthesis: an overview focused on the preparation of biologically relevant peptides" (in en). RSC Advances 4 (62): 32658–32672. doi:10.1039/c4ra02458c. ISSN 2046-2069. Bibcode: 2014RSCAd...432658P. https://digital.csic.es/bitstream/10261/187255/1/Solid-phase_Palomo_Publisher2014.pdf.
- ↑ Matyjaszewski, Krzysztof; Spanswick, James (2005-03-01). "Controlled/living radical polymerization". Materials Today 8 (3): 26–33. doi:10.1016/S1369-7021(05)00745-5. Bibcode: 2005MaTod...8...26M.
- ↑ Pfeifer, Sebastian; Lutz, Jean-François (2007-08-01). "A Facile Procedure for Controlling Monomer Sequence Distribution in Radical Chain Polymerizations". Journal of the American Chemical Society 129 (31): 9542–9543. doi:10.1021/ja0717616. ISSN 0002-7863. PMID 17636902. Bibcode: 2007JAChS.129.9542P.
- ↑ Tong, Xinming; Guo, Bao-hua; Huang, Yanbin (2011-02-07). "Toward the synthesis of sequence-controlled vinyl copolymers" (in en). Chem. Commun. 47 (5): 1455–1457. doi:10.1039/c0cc04807k. ISSN 1364-548X. PMID 21125120.
- ↑ Haven, Joris; De Neve, Jeroen; Junkers, Tanja (2017-06-24). "Versatile Approach for the Synthesis of Sequence-Defined Monodisperse 18- and 20-mer Oligoacrylates". ACS Macro Letters 6 (7): 743–747. doi:10.1021/acsmacrolett.7b00430. PMID 35650855.
- ↑ Vandenbergh, Joke; Reekmans, Gunter; Adriaensens, Peter; Junkers, Thomas (2013-09-17). "Synthesis of sequence controlled acrylate oligomers via consecutive RAFT monomer additions". Chemical Communications 49 (88): 10358–10360. doi:10.1039/c3cc45994b. PMID 24079009.
- ↑ Xu, Jiangtao (2019-10-21). "Single Unit Monomer Insertion: A Versatile Platform for Molecular Engineering through Radical Addition Reactions and Polymerization". Macromolecules 52 (23): 9068–9093. doi:10.1021/acs.macromol.9b01365. ISSN 0024-9297. Bibcode: 2019MaMol..52.9068X. https://unsworks.unsw.edu.au/bitstreams/589ab1fe-a24b-45c9-85c2-d84569cbf2b0/download.
- ↑ Chen, Yulin; Guan, Zhibin (2010-04-07). "Bioinspired Modular Synthesis of Elastin-Mimic Polymers To Probe the Mechanism of Elastin Elasticity". Journal of the American Chemical Society 132 (13): 4577–4579. doi:10.1021/ja9104446. ISSN 0002-7863. PMID 20235503. Bibcode: 2010JAChS.132.4577C.
- ↑ Yu, Ting-Bin; Bai, Jane Z.; Guan, Zhibin (2009-01-26). "Cycloaddition-Promoted Self-Assembly of a Polymer into Well-Defined β Sheets and Hierarchical Nanofibrils" (in en). Angewandte Chemie International Edition 48 (6): 1097–1101. doi:10.1002/anie.200805009. ISSN 1521-3773. PMID 19115358.
- ↑ Norris, Benjamin N.; Pan, Tianqi; Meyer, Tara Y. (2010-12-03). "Iterative Synthesis of Heterotelechelic Oligo(phenylene-vinylene)s by Olefin Cross-Metathesis". Organic Letters 12 (23): 5514–5517. doi:10.1021/ol102398y. ISSN 1523-7060. PMID 21069981.
- ↑ Zhang, Ze; You, Ye-Zi; Wu, De-Cheng; Hong, Chun-Yan (2015-06-09). "Syntheses of Sequence-Controlled Polymers via Consecutive Multicomponent Reactions". Macromolecules 48 (11): 3414–3421. doi:10.1021/acs.macromol.5b00463. ISSN 0024-9297. Bibcode: 2015MaMol..48.3414Z.
- ↑ Lewandowski, Bartosz; Bo, Guillaume De; Ward, John W.; Papmeyer, Marcus; Kuschel, Sonja; Aldegunde, María J.; Gramlich, Philipp M. E.; Heckmann, Dominik et al. (2013-01-11). "Sequence-Specific Peptide Synthesis by an Artificial Small-Molecule Machine" (in en). Science 339 (6116): 189–193. doi:10.1126/science.1229753. ISSN 0036-8075. PMID 23307739. Bibcode: 2013Sci...339..189L. https://www.research.manchester.ac.uk/portal/en/publications/sequencespecific-peptide-synthesis-by-an-artificial-smallmolecule-machine(dc7b8f29-3915-46cd-93a5-1b2419ee7466).html.
- ↑ Lutz, Jean-François (2010-02-09). "Sequence-controlled polymerizations: the next Holy Grail in polymer science?" (in en). Polymer Chemistry 1 (1): 55. doi:10.1039/b9py00329k. ISSN 1759-9962.
- ↑ Minoda, Masahiko; Sawamoto, Mitsuo; Higashimura, Toshinobu (1990-11-01). "Sequence-regulated oligomers and polymers by living cationic polymerization. 2. Principle of sequence regulation and synthesis of sequence-regulated oligomers of functional vinyl ethers and styrene derivatives". Macromolecules 23 (23): 4889–4895. doi:10.1021/ma00225a001. ISSN 0024-9297. Bibcode: 1990MaMol..23.4889M.
- ↑ 24.0 24.1 Ida, Shohei; Terashima, Takaya; Ouchi, Makoto; Sawamoto, Mitsuo (2009-08-12). "Selective Radical Addition with a Designed Heterobifunctional Halide: A Primary Study toward Sequence-Controlled Polymerization upon Template Effect". Journal of the American Chemical Society 131 (31): 10808–10809. doi:10.1021/ja9031314. ISSN 0002-7863. PMID 19603819. Bibcode: 2009JAChS.13110808I.
- ↑ Aksakal, R.; Mertens, C.; Soete, M.; Badi, N.; Du Prez, F. (2021). "Applications of Discrete Synthetic Macromolecules in Life and Materials Science: Recent and Future Trends". Advanced Science 2021 (2004038): 1–22. doi:10.1002/advs.202004038. PMID 33747749. Bibcode: 2021AdvSc...804038A.
- ↑ Steinkoenig, J.; Aksakal, R.; Du Prez, F. (2019). "Molecular access to multi-dimensionally encoded information". European Polymer Journal 2019 (120): 1–7. doi:10.1016/j.eurpolymj.2019.109260. Bibcode: 2019EurPJ.12009260S. https://www.sciencedirect.com/science/article/pii/S0014305719314041.
 |
