Biology:C16orf95
Chromosome 16 open reading frame 95 (C16orf95) is a gene which in humans encodes the protein C16orf95. It has orthologs in mammals, and is expressed at a low level in many tissues. C16orf95 evolves quickly compared to other proteins.
Gene
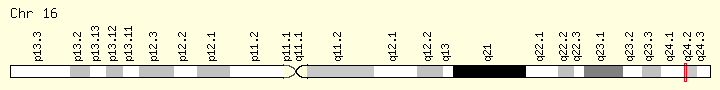
C16orf95 is a Homo sapiens gene oriented on the minus strand of chromosome 16. It is located on the cytogenic band 16q24.2 and spans 14.62 kilobases.[1] The gene contains 6 introns and 7 exons.[1]
Homology
Paralogs
There are no known paralogs of C16orf95.
Orthologs
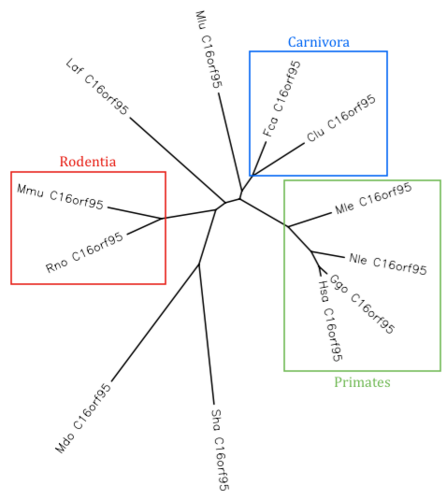
Orthologs of C16orf95 exist only in mammals (identified with BLAST).[3] The most distant orthologs are found in opossums and Tasmanian devils.
| Genus and species | Common name | NCBI accession | Date of divergence | Sequence identity |
| Homo sapiens | Human | NP_001182053 | 0 mya | 100% |
| Pan paniscus | Bonobo | XP_008972565 | 6.2 mya | 92% |
| Gorilla gorilla gorilla | Gorilla | XP_004058157 | 8.3 mya | 95% |
| Nomascus leucogenys | White-cheeked gibbon | XP_003272503 | 19.3 mya | 88% |
| Mandrillus leucophaeus | Drill | XP_011827052 | 27.3 mya | 78% |
| Propithecus coquereli | Lemur | XP_012513111 | 77.1 mya | 62% |
| Tupaia chinensis | Tree shrew | XP_006152612 | 86.5 mya | 58% |
| Oryctolagus cuniculus | European rabbit | XP_008250325 | 90.1 mya | 56% |
| Mus musculus | Mouse | NP_083873 | 90.1 mya | 54% |
| Rattus norvegicus | Rat | XP_006222844 | 90.1 mya | 51% |
| Camelus bactrianus | Camel | XP_010966555 | 95 mya | 63% |
| Canis lupus familiaris | Dog | XP_005620646 | 95 mya | 63% |
| Equus caballus | Horse | XP_005608538 | 95 mya | 60% |
| Felis catus | Cat | XP_011288582 | 95 mya | 60% |
| Bos taurus | Cattle | XP_015331266 | 95 mya | 60% |
| Lipotes vexillifer | Yangtze river dolphin | XP_007468528 | 95 mya | 50% |
| Myotis lucifugus | Brown bat | XP_014318589 | 95 mya | 56% |
| Trichechus manatus latirostris | Manatee | XP_004377854 | 102 mya | 66% |
| Loxodonta africana | Elephant | XP_003418190 | 102 mya | 59% |
| Orycteropus afer afer | Aardvark | XP_007937409 | 102 mya | 54% |
| Monodelphis domestica | Opossum | XP_007477328 | 162.4 mya | 42% |
| Sarcophilus harrisii | Tasmanian devil | XP_012395810 | 162.4 mya | 41% |

mRNA
Alternative splicing
There are three splice variants of C16orf95.[6] The longest transcript contains 1156 base pairs and 7 exons.[7] Compared to variant 1, the second transcript variant lacks exons 4 and 5.[8] This alternative splicing results in a frameshift of the 3' coding region, and a shorter, unique C-terminus. The third transcript variant lacks exons 4 and 5, and uses an alternate 5' exon and start codon.[9] The resulting peptide has unique N- and C-termini compared to variant 1.
| Size (base pairs) | |||
|---|---|---|---|
| Exon # | Variant 1 | Variant 2 | Variant 3 |
| 1 | 330 | 330 | 334 |
| 2 | 52 | 52 | 52 |
| 3 | 126 | 126 | 126 |
| 4 | 147 | – | – |
| 5 | 37 | – | – |
| 6 | 187 | 187 | 187 |
| 7 | 277 | 278 | 278 |
| Total | 1,156 | 973 | 977 |
Secondary structure
The 3' untranslated region of the C16orf95 mRNA contains binding sites for KH domain-containing, RNA-binding, signal transduction-associated protein 3 (KHDRBS3) within an internal loop structure. KHDRBS3 regulates mRNA splicing and may act as a negative regulator of cell growth.[12]
Expression
The expression of C16orf95 is not well characterized. However, it has been detected at low levels in the following tissue types: bone, brain, ear, eye, intestine, kidney, lung, lymph nodes, prostate, testes, tonsils, skin, and uterus.[13]
Protein
Structure
Primary
The longest isoform of the C16orf95 protein has 239 amino acids.[14] It has a conserved domain of unknown function spanning residues 76 to 239.[14] C16orf95 has a calculated molecular weight of 26.5 kDa, and a predicted isoelectric point of 9.8.[5] Compared to other human proteins, C16orf95 has more cysteine, arginine, and glutamine residues.[5] It has fewer aspartate, glutamate, and asparagine.[5] The high ratio of basic to acidic amino acids contributes to the protein's higher isoelectric point.
Secondary
C16orf95 is predicted to have several alpha-helices in its C-terminus.[5] This is true for the human and mouse proteins. The N-terminus does not have significant cross-program consensus for secondary structure.

Post-translational modifications
The tools available at ExPASy were used to predict post-translational modification sites on C16orf95.[16] The following modifications are predicted: palmitoylation, phosphorylation, and O-linked glycosylation. Bolded residues in the table indicate sites that are conserved in more than one species.
| Predicted modification | Sites - Homo sapiens | Sites - Mus musculus | Sites - Canis lupus familiaris | Tool |
|---|---|---|---|---|
| Palmitoylation | C77, C80, C126, C178,
C187 |
C24, C41, C90 | C64, C113, C174 | CSS-Palm[17] |
| Phosphorylation | S6, S9, S53, T57, S68,
S91, S111, T122, S166 |
S30, S76, S89, S120,
T134, S141 |
S15, S35, T39, S153 | NetPhos 2.0[18] |
| O-β-GlcNAc | S4, S6, S9, T57, S111 | None | None | NetOGlyc 4.0[19] |
Evolution
C16orf95 has a large number of amino acid changes over time, indicating it is a quickly evolving protein.

Interacting proteins
There are no proteins known to interact with C16orf95.
Clinical significance
Deletions of C16orf95 have been associated with hydronephrosis, microcephaly, distichiasis, vesicoureteral reflux, and intellectual impairment.[21][22] However, the deletions included coding regions of the following genes: F-box Protein 31 (FBXO31), Microtubule-Associated Protein 1 Light Chain 3 Beta (MAP1LC3B), and Zinc Finger CCHC Type 14 (ZCCHC14). The contributions of each of these genes to the observed phenotypes has yet to be scientifically determined.
References
- ↑ 1.0 1.1 "C16orf95 chromosome 16 open reading frame 95 [Homo sapiens (human) - Gene - NCBI"]. https://www.ncbi.nlm.nih.gov/gene?cmd=retrieve&list_uids=100506581.
- ↑ "C16orf95 Gene". Weizmann Institute of Science. https://www.genecards.org/cgi-bin/carddisp.pl?gene=C16orf95&keywords=c16orf95.
- ↑ "BLAST: Basic Local Alignment Search Tool". https://blast.ncbi.nlm.nih.gov.
- ↑ "TimeTree :: The Timescale of Life". http://timetree.org.
- ↑ 5.0 5.1 5.2 5.3 5.4 "SDSC Biology Workbench". http://workbench.sdsc.edu.
- ↑ "c16orf95 - Nucleotide - NCBI". https://www.ncbi.nlm.nih.gov/nuccore/?term=c16orf95.
- ↑ "Homo sapiens chromosome 16 open reading frame 95 (C16orf95), transcrip - Nucleotide - NCBI". https://www.ncbi.nlm.nih.gov/nuccore/NM_001195124.1.
- ↑ "Homo sapiens chromosome 16 open reading frame 95 (C16orf95), transcrip - Nucleotide - NCBI". https://www.ncbi.nlm.nih.gov/nuccore/NM_001195125.1.
- ↑ "Homo sapiens chromosome 16 open reading frame 95 (C16orf95), transcrip - Nucleotide - NCBI". https://www.ncbi.nlm.nih.gov/nuccore/NM_001256917.1.
- ↑ "RNA Folding Form". http://unafold.rna.albany.edu/?q=mfold/rna-folding-form.
- ↑ "RBPDB: The database of RNA-binding specificities". http://rbpdb.ccbr.utoronto.ca.
- ↑ "KHDRBS3 - KH domain-containing, RNA-binding, signal transduction-associated protein 3 - Homo sapiens (Human) - KHDRBS3 gene & protein". https://www.uniprot.org/uniprot/O75525.
- ↑ "EST Profile - Hs.729380". https://www.ncbi.nlm.nih.gov/UniGene/ESTProfileViewer.cgi?uglist=Hs.729380.
- ↑ 14.0 14.1 "uncharacterized protein C16orf95 isoform 1 [Homo sapiens - Protein - NCBI"]. https://www.ncbi.nlm.nih.gov/protein/NP_001182053.1.
- ↑ "SDSC Biology Workbench". http://workbench.sdsc.edu.
- ↑ "ExPASy: SIB Bioinformatics Resource Portal - Home". http://www.expasy.org.
- ↑ "CSS-Palm - Palmitoylation Site Prediction". http://csspalm.biocuckoo.org/.
- ↑ "NetPhos 2.0 Server". http://www.cbs.dtu.dk/services/NetPhos/.
- ↑ "NetOGlyc 4.0 Server". http://www.cbs.dtu.dk/services/NetOGlyc/.
- ↑ Griffiths, Anthony JF; Miller, Jeffrey H.; Suzuki, David T.; Lewontin, Richard C.; Gelbart, William M. (2000-01-01) (in en). Rate of molecular evolution. https://www.ncbi.nlm.nih.gov/books/NBK21946/.
- ↑ Handrigan, G. R., Chitayat, D., Lionel, A. C., Pinsk, M., Vaags, A. K., Marsall, C. R., ... Rosenblum, N. D. (2013). Deletions in 16q24.2 are associated with autism spectrum disorder, intellectual disability and congenital renal malformation. Journal of Medical Genetics, 50(4), 163-73. doi:10.1136/jmedgenet-2012-101288
- ↑ Butler, M. G., Dagenais, S. L., Garcia-Perez, J. L., Brouillard, P., Vikkula, M., Strouse, P., Innis, J. W., & Grover, T. W. (2012). Microcephaly, intellectual impairment, bilateral vesicoureteral reflux, distichiasis, and glomuvenous malformations associated with a 16q24.3 contiguous gene deletion and a Glomulin mutation. American Journal of Medical Genetics Part A, 158A(4), 839-49. doi:10.1002/ajmg.a.35229
 |


