Biology:Docking (molecular)
In the field of molecular modeling, docking is a method which predicts the preferred orientation of one molecule to a second when a ligand and a target are bound to each other to form a stable complex.[1] Knowledge of the preferred orientation in turn may be used to predict the strength of association or binding affinity between two molecules using, for example, scoring functions.
File:Docking GPCR example.webm The associations between biologically relevant molecules such as proteins, peptides, nucleic acids, carbohydrates, and lipids play a central role in signal transduction. Furthermore, the relative orientation of the two interacting partners may affect the type of signal produced (e.g., agonism vs antagonism). Therefore, docking is useful for predicting both the strength and type of signal produced.
Molecular docking is one of the most frequently used methods in structure-based drug design, due to its ability to predict the binding-conformation of small molecule ligands to the appropriate target binding site. Characterisation of the binding behaviour plays an important role in rational design of drugs as well as to elucidate fundamental biochemical processes.[2][3]
Definition of problem
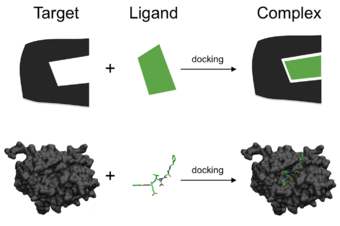
One can think of molecular docking as a problem of “lock-and-key”, in which one wants to find the correct relative orientation of the “key” which will open up the “lock” (where on the surface of the lock is the key hole, which direction to turn the key after it is inserted, etc.). Here, the protein can be thought of as the “lock” and the ligand can be thought of as a “key”. Molecular docking may be defined as an optimization problem, which would describe the “best-fit” orientation of a ligand that binds to a particular protein of interest. However, since both the ligand and the protein are flexible, a “hand-in-glove” analogy is more appropriate than “lock-and-key”.[4] During the course of the docking process, the ligand and the protein adjust their conformation to achieve an overall "best-fit" and this kind of conformational adjustment resulting in the overall binding is referred to as "induced-fit".[5]
Molecular docking research focuses on computationally simulating the molecular recognition process. It aims to achieve an optimized conformation for both the protein and ligand and relative orientation between protein and ligand such that the free energy of the overall system is minimized.
Docking approaches
Two approaches are particularly popular within the molecular docking community.
- One approach uses a matching technique that describes the protein and the ligand as complementary surfaces.[6][7][8]
- The second approach simulates the actual docking process in which the ligand-protein pairwise interaction energies are calculated.[9]
Both approaches have significant advantages as well as some limitations. These are outlined below.
Shape complementarity
Geometric matching/shape complementarity methods describe the protein and ligand as a set of features that make them dockable.[10] These features may include molecular surface/complementary surface descriptors. In this case, the receptor's molecular surface is described in terms of its solvent-accessible surface area and the ligand's molecular surface is described in terms of its matching surface description. The complementarity between the two surfaces amounts to the shape matching description that may help finding the complementary pose of docking the target and the ligand molecules. Another approach is to describe the hydrophobic features of the protein using turns in the main-chain atoms. Yet another approach is to use a Fourier shape descriptor technique.[11][12][13] Whereas the shape complementarity based approaches are typically fast and robust, they cannot usually model the movements or dynamic changes in the ligand/protein conformations accurately, although recent developments allow these methods to investigate ligand flexibility. Shape complementarity methods can quickly scan through several thousand ligands in a matter of seconds and actually figure out whether they can bind at the protein's active site, and are usually scalable to even protein-protein interactions. They are also much more amenable to pharmacophore based approaches, since they use geometric descriptions of the ligands to find optimal binding.
Simulation
Simulating the docking process is much more complicated. In this approach, the protein and the ligand are separated by some physical distance, and the ligand finds its position into the protein's active site after a certain number of “moves” in its conformational space. The moves incorporate rigid body transformations such as translations and rotations, as well as internal changes to the ligand's structure including torsion angle rotations. Each of these moves in the conformation space of the ligand induces a total energetic cost of the system. Hence, the system's total energy is calculated after every move.
The obvious advantage of docking simulation is that ligand flexibility is easily incorporated, whereas shape complementarity techniques must use ingenious methods to incorporate flexibility in ligands. Also, it more accurately models reality, whereas shape complementary techniques are more of an abstraction.
Clearly, simulation is computationally expensive, having to explore a large energy landscape. Grid-based techniques, optimization methods, and increased computer speed have made docking simulation more realistic.
Mechanics of docking
To perform a docking screen, the first requirement is a structure of the protein of interest. Usually the structure has been determined using a biophysical technique such as
- X-ray crystallography,
- NMR spectroscopy or
- cryo-electron microscopy (cryo-EM),
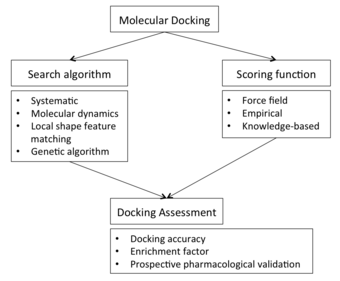
but can also derive from homology modeling construction. This protein structure and a database of potential ligands serve as inputs to a docking program. The success of a docking program depends on two components: the search algorithm and the scoring function.
Search algorithm
The search space in theory consists of all possible orientations and conformations of the protein paired with the ligand. However, in practice with current computational resources, it is impossible to exhaustively explore the search space — this would involve enumerating all possible distortions of each molecule (molecules are dynamic and exist in an ensemble of conformational states) and all possible rotational and translational orientations of the ligand relative to the protein at a given level of granularity. Most docking programs in use account for the whole conformational space of the ligand (flexible ligand), and several attempt to model a flexible protein receptor. Each "snapshot" of the pair is referred to as a pose.[14]
A variety of conformational search strategies have been applied to the ligand and to the receptor. These include:
- systematic or stochastic torsional searches about rotatable bonds
- molecular dynamics simulations
- genetic algorithms to "evolve" new low energy conformations and where the score of each pose acts as the fitness function used to select individuals for the next iteration.
Ligand flexibility
Conformations of the ligand may be generated in the absence of the receptor and subsequently docked[15] or conformations may be generated on-the-fly in the presence of the receptor binding cavity,[16] or with full rotational flexibility of every dihedral angle using fragment based docking.[17] Force field energy evaluation are most often used to select energetically reasonable conformations,[18] but knowledge-based methods have also been used.[19]
Peptides are both highly flexible and relatively large-sized molecules, which makes modeling their flexibility a challenging task. A number of methods were developed to allow for efficient modeling of flexibility of peptides during protein-peptide docking.[20]
Receptor flexibility
Computational capacity has increased dramatically over the last decade making possible the use of more sophisticated and computationally intensive methods in computer-assisted drug design. However, dealing with receptor flexibility in docking methodologies is still a thorny issue.[21] The main reason behind this difficulty is the large number of degrees of freedom that have to be considered in this kind of calculations. Neglecting it, however, in some of the cases may lead to poor docking results in terms of binding pose prediction.[22]
Multiple static structures experimentally determined for the same protein in different conformations are often used to emulate receptor flexibility.[23] Alternatively rotamer libraries of amino acid side chains that surround the binding cavity may be searched to generate alternate but energetically reasonable protein conformations.[24][25]
Scoring function
Docking programs generate a large number of potential ligand poses, of which some can be immediately rejected due to clashes with the protein. The remainder are evaluated using some scoring function, which takes a pose as input and returns a number indicating the likelihood that the pose represents a favorable binding interaction and ranks one ligand relative to another.
Most scoring functions are physics-based molecular mechanics force fields that estimate the energy of the pose within the binding site. The various contributions to binding can be written as an additive equation:
[math]\displaystyle{ \bigtriangleup G_{bind} = \bigtriangleup G_{solvent} + \bigtriangleup G_{conf} + \bigtriangleup G_{int} + \bigtriangleup G_{rot} + \bigtriangleup G_{t/t} + \bigtriangleup G_{vib} }[/math]
The components consist of solvent effects, conformational changes in the protein and ligand, free energy due to protein-ligand interactions, internal rotations, association energy of ligand and receptor to form a single complex and free energy due to changes in vibrational modes.[26] A low (negative) energy indicates a stable system and thus a likely binding interaction.
Alternative approaches use modified scoring functions to include constraints based on known key protein-ligand interactions,[27] or knowledge-based potentials derived from interactions observed in large databases of protein-ligand structures (e.g. the Protein Data Bank).[28]
There are a large number of structures from X-ray crystallography for complexes between proteins and high affinity ligands, but comparatively fewer for low affinity ligands as the latter complexes tend to be less stable and therefore more difficult to crystallize. Scoring functions trained with this data can dock high affinity ligands correctly, but they will also give plausible docked conformations for ligands that do not bind. This gives a large number of false positive hits, i.e., ligands predicted to bind to the protein that actually don't when placed together in a test tube.
One way to reduce the number of false positives is to recalculate the energy of the top scoring poses using (potentially) more accurate but computationally more intensive techniques such as Generalized Born or Poisson-Boltzmann methods.[9]
Docking assessment
The interdependence between sampling and scoring function affects the docking capability in predicting plausible poses or binding affinities for novel compounds. Thus, an assessment of a docking protocol is generally required (when experimental data is available) to determine its predictive capability. Docking assessment can be performed using different strategies, such as:
- docking accuracy (DA) calculation;
- the correlation between a docking score and the experimental response or determination of the enrichment factor (EF);[29]
- the distance between an ion-binding moiety and the ion in the active site;
- the presence of induce-fit models.
Docking accuracy
Docking accuracy[30][31] represents one measure to quantify the fitness of a docking program by rationalizing the ability to predict the right pose of a ligand with respect to that experimentally observed.[32]
Enrichment factor
Docking screens can also be evaluated by the enrichment of annotated ligands of known binders from among a large database of presumed non-binding, “decoy” molecules.[29] In this way, the success of a docking screen is evaluated by its capacity to enrich the small number of known active compounds in the top ranks of a screen from among a much greater number of decoy molecules in the database. The area under the receiver operating characteristic (ROC) curve is widely used to evaluate its performance.
Prospective
Resulting hits from docking screens are subjected to pharmacological validation (e.g. IC50, affinity or potency measurements). Only prospective studies constitute conclusive proof of the suitability of a technique for a particular target.[33] In the case of G protein-coupled receptors (GPCRs), which are targets of more than 30% of marketed drugs, molecular docking led to the discovery of more than 500 GPCR ligands.[34]
Benchmarking
The potential of docking programs to reproduce binding modes as determined by X-ray crystallography can be assessed by a range of docking benchmark sets.
For small molecules, several benchmark data sets for docking and virtual screening exist e.g. Astex Diverse Set consisting of high quality protein−ligand X-ray crystal structures[35] or the Directory of Useful Decoys (DUD) for evaluation of virtual screening performance.[29]
An evaluation of docking programs for their potential to reproduce peptide binding modes can be assessed by Lessons for Efficiency Assessment of Docking and Scoring (LEADS-PEP).[36]
Applications
A binding interaction between a small molecule ligand and an enzyme protein may result in activation or inhibition of the enzyme. If the protein is a receptor, ligand binding may result in agonism or antagonism. Docking is most commonly used in the field of drug design — most drugs are small organic molecules, and docking may be applied to:
- hit identification – docking combined with a scoring function can be used to quickly screen large databases of potential drugs in silico to identify molecules that are likely to bind to protein target of interest (see virtual screening). Reverse pharmacology routinely uses docking for target identification.
- lead optimization – docking can be used to predict in where and in which relative orientation a ligand binds to a protein (also referred to as the binding mode or pose). This information may in turn be used to design more potent and selective analogs.
- bioremediation – protein ligand docking can also be used to predict pollutants that can be degraded by enzymes.[37][38]
See also
- Drug design
- Katchalski-Katzir algorithm
- List of molecular graphics systems
- Macromolecular docking
- Molecular mechanics
- Protein structure
- Protein design
- Software for molecular mechanics modeling
- List of protein-ligand docking software
- Molecular design software
- Docking@Home
- Ibercivis
- ZINC database
- Lead Finder
- Virtual screening
- Scoring functions for docking
References
- ↑ "Computational methods for biomolecular docking". Current Opinion in Structural Biology 6 (3): 402–6. Jun 1996. doi:10.1016/S0959-440X(96)80061-3. PMID 8804827.
- ↑ "Docking and scoring in virtual screening for drug discovery: methods and applications". Nature Reviews. Drug Discovery 3 (11): 935–49. Nov 2004. doi:10.1038/nrd1549. PMID 15520816.
- ↑ "Study of CXCR4 chemokine receptor inhibitors using QSPR andmolecular docking methodologies". Journal of Theoretical and Computational Chemistry 178 (4). June 13, 2019. doi:10.1142/S0219633619500184.
- ↑ "Rusting of the lock and key model for protein-ligand binding". Science 254 (5034): 954–5. Nov 1991. doi:10.1126/science.1719636. PMID 1719636. Bibcode: 1991Sci...254..954J.
- ↑ "Testing a flexible-receptor docking algorithm in a model binding site". Journal of Molecular Biology 337 (5): 1161–82. Apr 2004. doi:10.1016/j.jmb.2004.02.015. PMID 15046985.
- ↑ "QSD quadratic shape descriptors. 2. Molecular docking using quadratic shape descriptors (QSDock)". Proteins 38 (1): 79–94. 2000. doi:10.1002/(SICI)1097-0134(20000101)38:1<79::AID-PROT9>3.0.CO;2-U. PMID 10651041.
- ↑ "Automated docking with grid-based energy evaluation". Journal of Computational Chemistry 13 (4): 505–524. 1992. doi:10.1002/jcc.540130412.
- ↑ "Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function". Journal of Computational Chemistry 19 (14): 1639–1662. 1998. doi:10.1002/(SICI)1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B.
- ↑ 9.0 9.1 "Performance comparison of generalized born and Poisson methods in the calculation of electrostatic solvation energies for protein structures". Journal of Computational Chemistry 25 (2): 265–84. Jan 2004. doi:10.1002/jcc.10378. PMID 14648625.
- ↑ "Molecular docking using shape descriptors". Journal of Computational Chemistry 13 (3): 380–397. 2004. doi:10.1002/jcc.540130311.
- ↑ "Protein-ligand recognition using spherical harmonic molecular surfaces: towards a fast and efficient filter for large virtual throughput screening". Journal of Molecular Graphics & Modelling 20 (4): 313–28. Jan 2002. doi:10.1016/S1093-3263(01)00134-6. PMID 11858640.
- ↑ "Real spherical harmonic expansion coefficients as 3D shape descriptors for protein binding pocket and ligand comparisons". Bioinformatics 21 (10): 2347–55. May 2005. doi:10.1093/bioinformatics/bti337. PMID 15728116.
- ↑ "Shape variation in protein binding pockets and their ligands". Journal of Molecular Biology 368 (1): 283–301. Apr 2007. doi:10.1016/j.jmb.2007.01.086. PMID 17337005.
- ↑ "Key Topics in Molecular Docking for Drug Design". International Journal of Molecular Sciences 20 (18): 4574. September 2019. doi:10.3390/ijms20184574. PMID 31540192.
- ↑ "Flexibases: a way to enhance the use of molecular docking methods". Journal of Computer-Aided Molecular Design 8 (5): 565–82. Oct 1994. doi:10.1007/BF00123666. PMID 7876901. Bibcode: 1994JCAMD...8..565K.
- ↑ "Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy". Journal of Medicinal Chemistry 47 (7): 1739–49. Mar 2004. doi:10.1021/jm0306430. PMID 15027865.
- ↑ "eHiTS: a new fast, exhaustive flexible ligand docking system". Journal of Molecular Graphics & Modelling 26 (1): 198–212. Jul 2007. doi:10.1016/j.jmgm.2006.06.002. PMID 16860582.
- ↑ "Preference of small molecules for local minimum conformations when binding to proteins". PLOS ONE 2 (9): e820. September 2007. doi:10.1371/journal.pone.0000820. PMID 17786192. Bibcode: 2007PLoSO...2..820W.
- ↑ "A fast and efficient method to generate biologically relevant conformations". Journal of Computer-Aided Molecular Design 8 (5): 583–606. October 1994. doi:10.1007/BF00123667. PMID 7876902. Bibcode: 1994JCAMD...8..583K.
- ↑ "Protein-peptide docking: opportunities and challenges". Drug Discovery Today 23 (8): 1530–1537. May 2018. doi:10.1016/j.drudis.2018.05.006. PMID 29733895.
- ↑ "Understanding the challenges of protein flexibility in drug design". Expert Opinion on Drug Discovery 10 (12): 1301–13. December 2015. doi:10.1517/17460441.2015.1094458. PMID 26414598. https://scholarship.rice.edu/bitstream/1911/88215/1/antunes-15-EODD.pdf.
- ↑ "MADAMM: a multistaged docking with an automated molecular modeling protocol". Proteins 74 (1): 192–206. January 2009. doi:10.1002/prot.22146. PMID 18618708.
- ↑ "Flexible ligand docking to multiple receptor conformations: a practical alternative". Current Opinion in Structural Biology 18 (2): 178–84. Apr 2008. doi:10.1016/j.sbi.2008.01.004. PMID 18302984.
- ↑ "Docking and scoring with alternative side-chain conformations". Proteins 74 (3): 712–26. Feb 2009. doi:10.1002/prot.22189. PMID 18704939.
- ↑ "FDS: flexible ligand and receptor docking with a continuum solvent model and soft-core energy function". Journal of Computational Chemistry 24 (13): 1637–56. Oct 2003. doi:10.1002/jcc.10295. PMID 12926007.
- ↑ "Computational Methods to Predict Binding Free Energy in Ligand-Receptor Complexes". Journal of Medicinal Chemistry 38 (26): 4953–67. Dec 1995. doi:10.1021/jm00026a001. PMID 8544170.
- ↑ Biased Docking for Protein-Ligand Pose Prediction. Methods in Molecular Biology. 2266. New York, NY: Springer US. 2021. pp. 39–72. doi:10.1007/978-1-0716-1209-5_3. ISBN 978-1-0716-1209-5.
- ↑ "Knowledge-based scoring function to predict protein-ligand interactions". Journal of Molecular Biology 295 (2): 337–356. January 2000. doi:10.1006/jmbi.1999.3371. PMID 10623530.
- ↑ 29.0 29.1 29.2 "Benchmarking sets for molecular docking". Journal of Medicinal Chemistry 49 (23): 6789–801. Nov 2006. doi:10.1021/jm0608356. PMID 17154509.
- ↑ "An Automated Strategy for Binding-Pose Selection and Docking Assessment in Structure-Based Drug Design". Journal of Chemical Information and Modeling 56 (1): 54–72. January 2016. doi:10.1021/acs.jcim.5b00603. PMID 26682916.
- ↑ "Comparative study of several algorithms for flexible ligand docking". Journal of Computer-Aided Molecular Design 17 (11): 755–763. November 2003. doi:10.1023/B:JCAM.0000017496.76572.6f. PMID 15072435. Bibcode: 2003JCAMD..17..755B.
- ↑ "Protein-Ligand Docking in Drug Design: Performance Assessment and Binding-Pose Selection". Rational Drug Design. Methods in Molecular Biology. 1824. 2018. pp. 67–88. doi:10.1007/978-1-4939-8630-9_5. ISBN 978-1-4939-8629-3.
- ↑ "Community benchmarks for virtual screening". Journal of Computer-Aided Molecular Design 22 (3–4): 193–199. 2008-02-14. doi:10.1007/s10822-008-9189-4. PMID 18273555. Bibcode: 2008JCAMD..22..193I.
- ↑ "Structure-Based Virtual Screening for Ligands of G Protein-Coupled Receptors: What Can Molecular Docking Do for You?". Pharmacological Reviews 73 (4): 527–565. October 2021. doi:10.1124/pharmrev.120.000246. PMID 34907092.
- ↑ "Diverse, high-quality test set for the validation of protein-ligand docking performance". Journal of Medicinal Chemistry 50 (4): 726–41. Feb 2007. doi:10.1021/jm061277y. PMID 17300160.
- ↑ "A Benchmark Data Set for Assessment of Peptide Docking Performance". Journal of Chemical Information and Modeling 56 (1): 188–200. Dec 2015. doi:10.1021/acs.jcim.5b00234. PMID 26651532.
- ↑ "An in silico [correction of insilico] approach to bioremediation: laccase as a case study". Journal of Molecular Graphics & Modelling 26 (5): 845–9. Jan 2008. doi:10.1016/j.jmgm.2007.05.005. PMID 17606396.
- ↑ "Implications of Molecular Docking Assay for Bioremediation". Data Analytics in Medicine: Concepts, Methodologies, Tools, and Applications. Advances in Environmental Engineering and Green Technologies. IGI Global. 2020. pp. 1556–1577. doi:10.4018/978-1-5225-2325-3.ch002. ISBN 978-1799812043. https://www.igi-global.com/chapter/implications-of-molecular-docking-assay-for-bioremediation/176457.
External links
- "Molecular Docking Server - Ligand Protein Docking & Molecular Modeling". Virtua Drug Ltd. http://www.dockingserver.com. "Internet service that calculates the site, geometry and energy of small molecules interacting with proteins"
- "Step by step installation of MGLTools 1.5.2 (AutoDockTools, Python Molecular Viewer and Visual Programming Environment) on Ubuntu Linux 8.04". http://users.ox.ac.uk/~jesu1458/installation_of_autodock_on_ubuntu_linux/.
- Docking@GRID Project of Conformational Sampling and Docking on Grids : one aim is to deploy some intrinsic distributed docking algorithms on computational Grids, download Docking@GRID open-source Linux version
- Click2Drug.org - Directory of computational drug design tools.
- Ligand:Receptor Docking with MOE (Molecular Operating Environment)
 |