Biology:Paired receptors
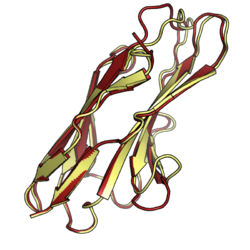
Paired receptors are pairs or clusters of receptor proteins that bind to extracellular ligands but have opposing activating and inhibitory signaling effects.[2][3][4] Traditionally, paired receptors are defined as homologous pairs with similar extracellular domains and different cytoplasmic regions, whose genes are located together in the genome as part of the same gene cluster and which evolved through gene duplication.[3][5] Homologous paired receptors often, but not always, have a shared ligand in common.[5][6] More broadly, pairs of receptors have been identified that exhibit paired functional behavior - responding to a shared ligand with opposing intracellular signals - but are not closely homologous or co-located in the genome.[4] Paired receptors are highly expressed in the cells of the immune system, especially natural killer (NK) and myeloid cells, and are involved in immune regulation.[5][7]
Structure
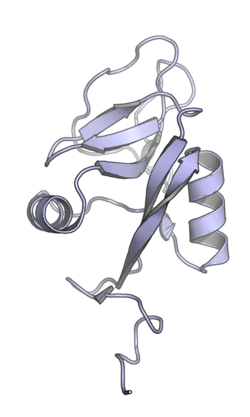
Paired receptors are membrane proteins with extracellular domains that interact with extracellular ligands. The extracellular region may contain multiple repeating protein domains and may be members of either the immunoglobulin or C-type lectin families.[5] The extracellular domains of homologous paired receptors are typically very similar in sequence but have different binding affinity for their shared ligands, with the inhibitory member of the pair binding more tightly.[4]
Homologous paired receptors have characteristic differences in their transmembrane and cytoplasmic regions that distinguish the activating and inhibiting members of the pair. Inhibitory receptors have a cytoplasmic sequence typically containing at least one immunoreceptor tyrosine-based inhibitory motif (ITIM). Activating receptors have a truncated cytoplasmic sequence compared to their corresponding inhibitory receptor and feature a positively charged amino acid residue in their transmembrane domain, enabling protein-protein interaction with an adaptor protein that possesses a immunoreceptor tyrosine-based activation motif (ITAM).[3]
Genetics and evolution
Homologous paired receptors are located in the same gene cluster and are thought to have evolved through gene duplication.[3][5] Sequence features such as the presence of an ITIM-like sequence in the 3' untranslated region of some activating receptors imply that the activating members of the pair likely evolved from the inhibitory members.[4][9] A number of pathogens interact with the inhibitory member of a pair as a means of immune evasion or viral entry, suggesting that activating members with similar binding competencies may be an evolutionary response to this mechanism.[4][10] This hypothesis is known as the "counterbalance theory"[11] and these evolutionary dynamics represent an evolutionary arms race between pathogens and the host immune system.[12] The evolutionary pressures on some paired-receptor families have been described as examples of the "Red Queen" effect.[5]
Including non-paired examples, over 300 potential immune inhibitory receptors have been identified in the human genome.[6] There are strong indications that paired receptors are rapidly and recently evolving. These genetic regions have high levels of gene polymorphism, and the gene repertoires found in the genomes of closely related lineages vary significantly.[5] The selective pressure experienced by the host from pathogens is thought to underlie this rapid evolution.[4][5]
Although paired receptors are best characterized as part of the human and mouse immune systems,[4] they have also been studied in other organisms. The chicken (Gallus gallus domesticus) genome contains a number of examples including a very large family, the chicken Ig-like receptors (CHIR) with over 100 members.[13] Paired receptor evolution has also been studied in Xenopus (clawed frog) species.[14][15] The adaptive immune system is unique to jawed vertebrates, but an example of a paired receptor family has been identified in a jawless vertebrate, termed agnathan paired receptors resembling Ag receptors (APAR) in the hagfish.[16]
Expression
Expression of paired receptors is common in many types of leukocytes, especially myeloid cells and natural killer (NK) cells.[4][5][7] Activation of NK cells is a complex regulatory process modulated by a number of different paired receptor families coexpressed in this cell type.[7] In some cases, only one member of the pair is expressed in a cell type. Expression of the paired members in a single cell type may vary with time, or the proteins may differ in subcellular localization, resulting in variations in signaling.[4] Expression in NK cells can be stochastic, resulting in unique variations in receptor repertoire.[4][12]
Some paired receptors are expressed outside the immune system, for example in neurons,[3][12] endothelium, and epithelium[5] but in many examples, wide tissue distribution can be observed.[4]
Function

Paired receptors transduce extracellular signals through opposing intracellular signaling pathways. Canonically, inhibitory receptors recruit phosphatases through their ITIM motifs, inhibiting the function of cells in which they are expressed. By contrast, activating receptors interact with adaptor proteins such as DAP-12 bearing an ITAM motif, which in turn recruit kinases such as Syk and ZAP70.[5]
Ligands for paired receptors can be very diverse. They are often proteins; the best-characterized are the MHC class I molecules, but a number of other endogenous molecules have been described as ligands for at least one family of paired receptors, and in a few cases in the LILR family, even intact bacteria or viruses can serve as ligands.[18] Lipids such as phosphatidylethanolamine and phosphatidylserine, sugars and sialylated glycans, and nucleic acids can all serve as ligands for some paired receptors.[4]
The binding affinity of paired receptors' extracellular domains for their ligands is generally fairly weak, with dissociation constants (Kd) in the micromolar (μM) range. However, the inhibitory member of a pair usually binds with higher affinity than the activating member.[3][4] This can produce a competitive inhibition effect, in which the inhibitory member of the pair out-competes its activating counterpart for ligand binding; other mechanisms of interference with activation, such as disrupting dimerization, have also been described.[4] Thus the net baseline signal from the pair is usually inhibitory, but may be modulated through differences in expression, surface density, subcellular localization, or other factors.[4]
In NK cells, ligands for inhibitory receptors are often MHC class I (MHC-I) molecules, while those for activating receptors may include signals of abnormality or infection such as proteins from pathogens or tumors, or molecules associated with cell stress.[5] Endogenous ligands for inhibitory receptors are better characterized than those for activating receptors.[3] Paired receptor signaling may represent maintenance of homeostasis such that immune responses to normal host cells are inhibited, while responses to abnormal or pathogenic molecules in the environment are activating. NK activation in the absence of inhibitory receptor signals from endogenous ligands is a molecular mechanism for the missing-self hypothesis of NK activation.[3][5][12]
Interaction with pathogens
A number of examples of molecular mimicry by pathogens, emulating natural endogenous ligands of paired receptors for immune evasion, have been described in the literature. Such interactions are particularly common with the inhibitory members of receptor pairs, bolstering the hypothesis that activating partners are a later evolutionary response to this immune escape strategy.[4][10]
The first described interaction between a paired receptor and a viral protein identified ILT-2 and ILR-4 (LILRB1 and LILRB2) as targets for herpes simplex virus UL18 protein, which resembles an MHC-I molecule.[5] Variations in susceptibility to mouse cytomegalovirus infection due to differences in Ly49-family paired receptors among mouse strains are well-characterized, and are attributed to the structural resemblance between the viral protein m157 and MHC-I molecules.[5] The pathogenic bacterium Escherichia coli K1 exposes surface polysialic acid molecules that serve as a molecular mimic for the native ligand of the inhibitory receptor Siglec-11, but induces an opposing response through interactions with the paired activating receptor Siglec-16, exemplifying the benefit of activating receptors as defense mechanisms against molecular mimicry by pathogens.[10]
Paired receptors are also used as viral entry receptors by a number of viruses and occasionally as entry mechanisms for other pathogens.[4] Sialylation is common among mammalian cell-surface proteins and a number of pathogens use sialic acid - either self-synthesized or obtained from the host cell - to evade host immunity, including by interacting with inhibitory siglec receptors.[5]
Families
There are two main groups of paired receptors, distinguished by extracellular regions containing immunoglobulin or C-type lectin domains. Nomenclature within these families is complex and has changed over time as new members were identified.[19] In general, the example of the LILR family applies; genes designated A represent the inhibitory receptor and genes designated B represent the activating receptor.[18]
Immunoglobulin-like receptors
Immunoglobulin-like receptors are members of the immunoglobulin superfamily and have one or more 70-110 residue immunoglobulin domains (Ig) in their extracellular region, typically multiple such domains in tandem. Many of the genes encoding these proteins occur in the leukocyte receptor complex (LRC), a large gene cluster on human chromosome 19.[3] Members of this group found in the human genome include:
- The killer-cell immunoglobulin-like receptor (KIR) family contains proteins with 2-3 extracellular Ig domains and long (inhibitory) or short (activating) cytoplasmic regions. Typically expressed in NK and some T cells, they interact with MHC class I.[3] This gene family located in the LRC is highly polymorphic and there is individual variation in both alleles and copy number, as well as in alternative splicing.[20] This family has undergone significant diversification in primate lineages.[21]
- The leukocyte immunoglobulin-like receptors (LILR) family contains 13 genes, including two pseudogenes. They have 2-4 Ig domains. One member, LILRA3, lacks a transmembrane region and is a soluble protein; others may be expressed in soluble form through alternative splicing.[22] Like the similar KIR family, LILR genes are found in the LRC and are polymorphic, though less so than KIR. LILR proteins are broadly expressed in immune cells and have very diverse ligands.[18]
- The paired type 2 immunoglobulin like receptor (PILR) family contains two genes, PILRA (inhibiting) and PILRB (activating).[3] They have a single extracellular Ig domain with a siglec-like structure.[1]
- The signal regulatory protein (SIRP) family contains three genes, SIRPA (inhibiting), SIRPB1 (activating), and SIRPG (non-signaling),[23] with the more distantly related SIRPD and SIRPB2 not yet well characterized.[24] SIRPA interacts with CD47, a regulator of phagocytosis.[24] This family also interacts with surfactant protein D.[3]
- The carcinoembryonic antigen-related cell adhesion (CEACAM) family contains 12 genes with one or more Ig domains.[25] They are expressed broadly, especially in endothelium and epithelium[5][25] and have roles in cell-cell recognition.[26] They have been extensively studied for their role in cancer and have been used as cancer biomarkers.[25]
- The siglec family contains 15 genes divided into two evolutionarily related groups. This family has three members with activating motifs, Siglec-14, Siglec-15, and Siglec-16.[27] These proteins bind sialic acids, and are often targeted by pathogens.[5]
- TIGIT (T cell immunoreceptor with Ig and ITIM domains) is an inhibitory receptor that forms a nonhomologous but functional pair with DNAM1 (CD226).[4][7]
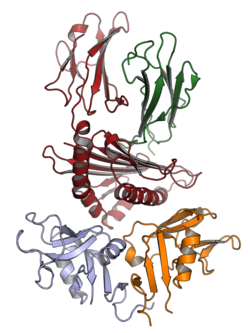
C-type lectin-like receptors
C-type lectin-like receptors (CLRs) contain one or more C-type lectin (Ca2+ dependent carbohydrate-binding lectin) domains. Example pairs include:
- CD94/NKG2, expressed in NK and some T cells and interacts with the ligand HLA-E.[3]
- Dendritic cell immunoreceptor (DCIR)/dendritic cell immunoactivating receptor (DCAR), characterized as a pair in mice, though no human DCAR has been identified.[28]
- NKR-P1 (CD161) is a member of a paired receptor group in rodents, but the human genome contains only one, inhibitory receptor, NKRP1A (KLRB1).[3]
- The Ly49 family in mice has been extensively studied for its role in NK activation using laboratory mice as a model organism, but has no homologous gene cluster in the human genome. The KIR family is the functional equivalent.[5]
References
- ↑ 1.0 1.1 Lu, Q.; Lu, G.; Qi, J.; Wang, H.; Xuan, Y.; Wang, Q.; Li, Y.; Zhang, Y. et al. (2014-06-03). "PILR and PILR have a siglec fold and provide the basis of binding to sialic acid". Proceedings of the National Academy of Sciences 111 (22): 8221–8226. doi:10.1073/pnas.1320716111. PMID 24843130.
- ↑ Lanier, Lewis L (June 2001). "Face off — the interplay between activating and inhibitory immune receptors". Current Opinion in Immunology 13 (3): 326–331. doi:10.1016/S0952-7915(00)00222-3. PMID 11406364.
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 3.13 Kuroki, Kimiko; Furukawa, Atsushi; Maenaka, Katsumi (2012). "Molecular Recognition of Paired Receptors in the Immune System". Frontiers in Microbiology 3: 429. doi:10.3389/fmicb.2012.00429. PMID 23293633.
- ↑ 4.00 4.01 4.02 4.03 4.04 4.05 4.06 4.07 4.08 4.09 4.10 4.11 4.12 4.13 4.14 4.15 4.16 4.17 Levi-Schaffer, Francesca; Mandelboim, Ofer (February 2018). "Inhibitory and Coactivating Receptors Recognising the Same Ligand: Immune Homeostasis Exploited by Pathogens and Tumours". Trends in Immunology 39 (2): 112–122. doi:10.1016/j.it.2017.10.001. PMID 29066058.
- ↑ 5.00 5.01 5.02 5.03 5.04 5.05 5.06 5.07 5.08 5.09 5.10 5.11 5.12 5.13 5.14 5.15 5.16 5.17 5.18 Akkaya, Munir; Barclay, A. Neil (February 2013). "How do pathogens drive the evolution of paired receptors?: HIGHLIGHTS". European Journal of Immunology 43 (2): 303–313. doi:10.1002/eji.201242896. PMID 23280392.
- ↑ 6.0 6.1 Rumpret, Matevž; Drylewicz, Julia; Ackermans, Laura J. E.; Borghans, José A. M.; Medzhitov, Ruslan; Meyaard, Linde (December 2020). "Functional categories of immune inhibitory receptors". Nature Reviews Immunology 20 (12): 771–780. doi:10.1038/s41577-020-0352-z. PMID 32612208.
- ↑ 7.0 7.1 7.2 7.3 Martinet, Ludovic; Smyth, Mark J. (April 2015). "Balancing natural killer cell activation through paired receptors". Nature Reviews Immunology 15 (4): 243–254. doi:10.1038/nri3799. PMID 25743219.
- ↑ 8.0 8.1 Petrie, Emma J.; Clements, Craig S.; Lin, Jie; Sullivan, Lucy C.; Johnson, Darryl; Huyton, Trevor; Heroux, Annie; Hoare, Hilary L. et al. (2008-03-17). "CD94-NKG2A recognition of human leukocyte antigen (HLA)-E bound to an HLA class I leader sequence". Journal of Experimental Medicine 205 (3): 725–735. doi:10.1084/jem.20072525. PMID 18332182.
- ↑ Arase, Hisashi; Lanier, Lewis L. (March 2004). "Specific recognition of virus-infected cells by paired NK receptors: Paired NK receptors". Reviews in Medical Virology 14 (2): 83–93. doi:10.1002/rmv.422. PMID 15027001.
- ↑ 10.0 10.1 10.2 Schwarz, Flavio; Landig, Corinna S; Siddiqui, Shoib; Secundino, Ismael; Olson, Joshua; Varki, Nissi; Nizet, Victor; Varki, Ajit (2017-03-15). "Paired Siglec receptors generate opposite inflammatory responses to a human‐specific pathogen". The EMBO Journal 36 (6): 751–760. doi:10.15252/embj.201695581. PMID 28100677.
- ↑ Barclay, A. Neil; Hatherley, Deborah (November 2008). "The Counterbalance Theory for Evolution and Function of Paired Receptors". Immunity 29 (5): 675–678. doi:10.1016/j.immuni.2008.10.004. PMID 19006692.
- ↑ 12.0 12.1 12.2 12.3 Miletić, Antonija; Krmpotić, Astrid; Jonjić, Stipan (April 2013). "The evolutionary arms race between NK cells and viruses: Who gets the short end of the stick?: HIGHLIGHTS". European Journal of Immunology 43 (4): 867–877. doi:10.1002/eji.201243101.
- ↑ Viertlboeck, Birgit C.; Hanczaruk, Matthias A.; Amann, Barbara; Bader, Sophie R.; Schmitt, Ramona; Sperling, Beatrice; Schwarz, Susanne C.N.; Schmahl, Wolfgang et al. (November 2013). "Chicken immunoregulatory Ig-like receptor families: An overview and expression details on ggTREM-A1". Developmental & Comparative Immunology 41 (3): 403–412. doi:10.1016/j.dci.2013.04.017. PMID 23648646.
- ↑ Guselnikov, Sergey V; Ramanayake, Thaminda; Erilova, Aleksandra Y; Mechetina, Ludmila V; Najakshin, Alexander M; Robert, Jacques; Taranin, Alexander V (2008). "The Xenopus FcR family demonstrates continually high diversification of paired receptors in vertebrate evolution". BMC Evolutionary Biology 8 (1): 148. doi:10.1186/1471-2148-8-148. PMID 18485190.
- ↑ Zimmermann, Wolfgang; Kammerer, Robert (December 2016). "Coevolution of paired receptors in Xenopus carcinoembryonic antigen-related cell adhesion molecule families suggests appropriation as pathogen receptors". BMC Genomics 17 (1): 928. doi:10.1186/s12864-016-3279-9. PMID 27852220.
- ↑ Suzuki, Takashi; Shin-I, Tadasu; Fujiyama, Asao; Kohara, Yuji; Kasahara, Masanori (2005-03-01). "Hagfish Leukocytes Express a Paired Receptor Family with a Variable Domain Resembling Those of Antigen Receptors". The Journal of Immunology 174 (5): 2885–2891. doi:10.4049/jimmunol.174.5.2885. PMID 15728499.
- ↑ Fan, Qing R.; Long, Eric O.; Wiley, Don C. (May 2001). "Crystal structure of the human natural killer cell inhibitory receptor KIR2DL1–HLA-Cw4 complex". Nature Immunology 2 (5): 452–460. doi:10.1038/87766. PMID 11323700.
- ↑ 18.0 18.1 18.2 Burshtyn, Deborah N.; Morcos, Chris (2016-02-01). "The Expanding Spectrum of Ligands for Leukocyte Ig-like Receptors". The Journal of Immunology 196 (3): 947–955. doi:10.4049/jimmunol.1501937. PMID 26802060.
- ↑ Yamada, Eriko; McVicar, Daniel W. (May 2008). "Paired Receptor Systems of the Innate Immune System". Current Protocols in Immunology 81 (1): Appendix-1X. doi:10.1002/0471142735.ima01xs81. PMID 18491293.
- ↑ Bruijnesteijn, Jesse; van der Wiel, Marit K. H.; de Groot, Nanine; Otting, Nel; de Vos-Rouweler, Annemiek J. M.; Lardy, Neubury M.; de Groot, Natasja G.; Bontrop, Ronald E. (2018-12-04). "Extensive Alternative Splicing of KIR Transcripts". Frontiers in Immunology 9: 2846. doi:10.3389/fimmu.2018.02846. PMID 30564240.
- ↑ Bruijnesteijn, Jesse; de Groot, Natasja G.; Bontrop, Ronald E. (2020-09-11). "The Genetic Mechanisms Driving Diversification of the KIR Gene Cluster in Primates". Frontiers in Immunology 11. doi:10.3389/fimmu.2020.582804. PMID 33013938.
- ↑ Lewis Marffy, Alexander L.; McCarthy, Alex J. (2020-05-13). "Leukocyte Immunoglobulin-Like Receptors (LILRs) on Human Neutrophils: Modulators of Infection and Immunity". Frontiers in Immunology 11: 857. doi:10.3389/fimmu.2020.00857. PMID 32477348.
- ↑ Takahashi, Shinichiro (2018-05-23). "Molecular functions of SIRPα and its role in cancer (Review)". Biomedical Reports 9 (1): 3–7. doi:10.3892/br.2018.1102. PMID 29930800.
- ↑ 24.0 24.1 Barclay, A. Neil; van den Berg, Timo K. (2014-03-21). "The Interaction Between Signal Regulatory Protein Alpha (SIRP α ) and CD47: Structure, Function, and Therapeutic Target". Annual Review of Immunology 32 (1): 25–50. doi:10.1146/annurev-immunol-032713-120142. PMID 24215318.
- ↑ 25.0 25.1 25.2 Han, Zi-Wen; Lyv, Zhi-Wu; Cui, Bin; Wang, Ying-Ying; Cheng, Jun-Ting; Zhang, Ying; Cai, Wen-Qi; Zhou, Yang et al. (December 2020). "The old CEACAMs find their new role in tumor immunotherapy". Investigational New Drugs 38 (6): 1888–1898. doi:10.1007/s10637-020-00955-w. PMID 32488569.
- ↑ Kuespert, Katharina; Pils, Stefan; Hauck, Christof R (October 2006). "CEACAMs: their role in physiology and pathophysiology". Current Opinion in Cell Biology 18 (5): 565–571. doi:10.1016/j.ceb.2006.08.008. PMID 16919437.
- ↑ Lenza, María Pia; Atxabal, Unai; Oyenarte, Iker; Jiménez-Barbero, Jesús; Ereño-Orbea, June (2020-12-15). "Current Status on Therapeutic Molecules Targeting Siglec Receptors". Cells 9 (12): 2691. doi:10.3390/cells9122691. PMID 33333862.
- ↑ Toyonaga, Kenji; Yamasaki, Sho (2020). "Recognition of Mycobacteria by Dendritic Cell Immunoactivating Receptor". C-Type Lectins in Immune Homeostasis. Current Topics in Microbiology and Immunology 429: 103–115. doi:10.1007/82_2020_203. ISBN 978-3-030-62236-7. PMID 32300915.
 |
