Biology:Comparative genomics

Comparative genomics is a branch of biological research that examines genome sequences across a spectrum of species, spanning from humans and mice to a diverse array of organisms from bacteria to chimpanzees.[2][3] This large-scale holistic approach compares two or more genomes to discover the similarities and differences between the genomes and to study the biology of the individual genomes.[4] Comparison of whole genome sequences provides a highly detailed view of how organisms are related to each other at the gene level. By comparing whole genome sequences, researchers gain insights into genetic relationships between organisms and study evolutionary changes.[2] The major principle of comparative genomics is that common features of two organisms will often be encoded within the DNA that is evolutionarily conserved between them. Therefore, Comparative genomics provides a powerful tool for studying evolutionary changes among organisms, helping to identify genes that are conserved or common among species, as well as genes that give unique characteristics of each organism. Moreover, these studies can be performed at different levels of the genomes to obtain multiple perspectives about the organisms.[4]
The comparative genomic analysis begins with a simple comparison of the general features of genomes such as genome size, number of genes, and chromosome number. Table 1 presents data on several fully sequenced model organisms, and highlights some striking findings. For instance, while the tiny flowering plant Arabidopsis thaliana has a smaller genome than that of the fruit fly Drosophila melanogaster (157 million base pairs v. 165 million base pairs, respectively) it possesses nearly twice as many genes (25,000 v. 13,000). In fact, A. thaliana has approximately the same number of genes as humans (25,000). Thus, a very early lesson learned in the genomic era is that genome size does not correlate with evolutionary status, nor is the number of genes proportionate to genome size.[5]
| Organism | Estimated size (base pairs) | Chromosome number | Estimated gene number |
|---|---|---|---|
| Human (Homo sapiens) | 3.1 billion | 46 | 25,000 |
| Mouse (Mus musculus) | 2.9 billion | 40 | 25,000 |
| Bovine (Bos taurus) | 2.86 billion[6] | 60[7] | 22,000[8] |
| Fruit fly (Drosophila melanogaster) | 165 million | 8 | 13,000 |
| Plant (Arabidopsis thaliana) | 157 million | 10 | 25,000 |
| Roundworm (Caenorhabditis elegans) | 97 million | 12 | 19,000 |
| Yeast (Saccharomyces cerevisiae) | 12 million | 32 | 6,000 |
| Bacteria (Escherichia coli) | 4.6 million | 1 | 3,200 |
In comparative genomics, synteny is the preserved order of genes on chromosomes of related species indicating their descent from a common ancestor. Synteny provides a framework in which the conservation of homologous genes and gene order is identified between genomes of different species.[9] Synteny blocks are more formally defined as regions of chromosomes between genomes that share a common order of homologous genes derived from a common ancestor.[10][11] Alternative names such as conserved synteny or collinearity have been used interchangeably.[12] Comparisons of genome synteny between and within species have provided an opportunity to study evolutionary processes that lead to the diversity of chromosome number and structure in many lineages across the tree of life;[13][14] early discoveries using such approaches include chromosomal conserved regions in nematodes and yeast,[15][16] evolutionary history and phenotypic traits of extremely conserved Hox gene clusters across animals and MADS-box gene family in plants,[17][18] and karyotype evolution in mammals and plants.[19]
Furthermore, comparing two genomes not only reveals conserved domains or synteny but also aids in detecting copy number variations, single nucleotide polymorphisms (SNPs), indels, and other genomic structural variations.
Virtually started as soon as the whole genomes of two organisms became available (that is, the genomes of the bacteria Haemophilus influenzae and Mycoplasma genitalium) in 1995, comparative genomics is now a standard component of the analysis of every new genome sequence.[2][20] With the explosion in the number of genome projects due to the advancements in DNA sequencing technologies, particularly the next-generation sequencing methods in late 2000s, this field has become more sophisticated, making it possible to deal with many genomes in a single study.[21] Comparative genomics has revealed high levels of similarity between closely related organisms, such as humans and chimpanzees, and, more surprisingly, similarity between seemingly distantly related organisms, such as humans and the yeast Saccharomyces cerevisiae.[22] It has also showed the extreme diversity of the gene composition in different evolutionary lineages.[20]
History
Comparative genomics has a root in the comparison of virus genomes in the early 1980s.[20] For example, small RNA viruses infecting animals (picornaviruses) and those infecting plants (cowpea mosaic virus) were compared and turned out to share significant sequence similarity and, in part, the order of their genes.[23] In 1986, the first comparative genomic study at a larger scale was published, comparing the genomes of varicella-zoster virus and Epstein-Barr virus that contained more than 100 genes each.[24]
The first complete genome sequence of a cellular organism, that of Haemophilus influenzae Rd, was published in 1995.[25] The second genome sequencing paper was of the small parasitic bacterium Mycoplasma genitalium published in the same year.[26] Starting from this paper, reports on new genomes inevitably became comparative-genomic studies.[20]
Microbial genomes. The first high-resolution whole genome comparison system of microbial genomes of 10-15kbp was developed in 1998 by Art Delcher, Simon Kasif and Steven Salzberg and applied to the comparison of entire highly related microbial organisms with their collaborators at the Institute for Genomic Research (TIGR). The system is called MUMMER and was described in a publication in Nucleic Acids Research in 1999. The system helps researchers to identify large rearrangements, single base mutations, reversals, tandem repeat expansions and other polymorphisms. In bacteria, MUMMER enables the identification of polymorphisms that are responsible for virulence, pathogenicity, and anti-biotic resistance. The system was also applied to the Minimal Organism Project at TIGR and subsequently to many other comparative genomics projects.
Eukaryote genomes. Saccharomyces cerevisiae, the baker's yeast, was the first eukaryote to have its complete genome sequence published in 1996.[27] After the publication of the roundworm Caenorhabditis elegans genome in 1998[15] and together with the fruit fly Drosophila melanogaster genome in 2000,[28] Gerald M. Rubin and his team published a paper titled "Comparative Genomics of the Eukaryotes", in which they compared the genomes of the eukaryotes D. melanogaster, C. elegans, and S. cerevisiae, as well as the prokaryote H. influenzae.[29] At the same time, Bonnie Berger, Eric Lander, and their team published a paper on whole-genome comparison of human and mouse.[30]
With the publication of the large genomes of vertebrates in the 2000s, including human, the Japanese pufferfish Takifugu rubripes, and mouse, precomputed results of large genome comparisons have been released for downloading or for visualization in a genome browser. Instead of undertaking their own analyses, most biologists can access these large cross-species comparisons and avoid the impracticality caused by the size of the genomes.[31]
Next-generation sequencing methods, which were first introduced in 2007, have produced an enormous amount of genomic data and have allowed researchers to generate multiple (prokaryotic) draft genome sequences at once. These methods can also quickly uncover single-nucleotide polymorphisms, insertions and deletions by mapping unassembled reads against a well annotated reference genome, and thus provide a list of possible gene differences that may be the basis for any functional variation among strains.[21]
Evolutionary principles
One character of biology is evolution, evolutionary theory is also the theoretical foundation of comparative genomics, and at the same time the results of comparative genomics unprecedentedly enriched and developed the theory of evolution. When two or more of the genome sequence are compared, one can deduce the evolutionary relationships of the sequences in a phylogenetic tree. Based on a variety of biological genome data and the study of vertical and horizontal evolution processes, one can understand vital parts of the gene structure and its regulatory function.
Similarity of related genomes is the basis of comparative genomics. If two creatures have a recent common ancestor, the differences between the two species genomes are evolved from the ancestors' genome. The closer the relationship between two organisms, the higher the similarities between their genomes. If there is close relationship between them, then their genome will display a linear behaviour (synteny), namely some or all of the genetic sequences are conserved. Thus, the genome sequences can be used to identify gene function, by analyzing their homology (sequence similarity) to genes of known function. Human [[FOXP2 gene and evolutionary conservation is shown in and multiple alignment (at bottom of figure) in this image from the UCSC Genome Browser. Note that conservation tends to cluster around coding regions (exons). |thumb|right]] Orthologous sequences are related sequences in different species: a gene exists in the original species, the species divided into two species, so genes in new species are orthologous to the sequence in the original species. Paralogous sequences are separated by gene cloning (gene duplication): if a particular gene in the genome is copied, then the copy of the two sequences is paralogous to the original gene. A pair of orthologous sequences is called orthologous pairs (orthologs), a pair of paralogous sequence is called collateral pairs (paralogs). Orthologous pairs usually have the same or similar function, which is not necessarily the case for collateral pairs. In collateral pairs, the sequences tend to evolve into having different functions.
Comparative genomics exploits both similarities and differences in the proteins, RNA, and regulatory regions of different organisms to infer how selection has acted upon these elements. Those elements that are responsible for similarities between different species should be conserved through time (stabilizing selection), while those elements responsible for differences among species should be divergent (positive selection). Finally, those elements that are unimportant to the evolutionary success of the organism will be unconserved (selection is neutral).
One of the important goals of the field is the identification of the mechanisms of eukaryotic genome evolution. It is however often complicated by the multiplicity of events that have taken place throughout the history of individual lineages, leaving only distorted and superimposed traces in the genome of each living organism. For this reason comparative genomics studies of small model organisms (for example the model Caenorhabditis elegans and closely related Caenorhabditis briggsae) are of great importance to advance our understanding of general mechanisms of evolution.[32][33]
Role of CNVs in evolution
Comparative genomics plays a crucial role in identifying copy number variations (CNVs) and understanding their significance in evolution. CNVs, which involve deletions or duplications of large segments of DNA, are recognized as a major source of genetic diversity, influencing gene structure, dosage, and regulation. While single nucleotide polymorphisms (SNPs) are more common, CNVs impact larger genomic regions and can have profound effects on phenotype and diversity.[34] Recent studies suggest that CNVs constitute around 4.8–9.5% of the human genome and have a substantial functional and evolutionary impact. In mammals, CNVs contribute significantly to population diversity, influencing gene expression and various phenotypic traits.[35] Comparative genomics analyses of human and chimpanzee genomes have revealed that CNVs may play a greater role in evolutionary change compared to single nucleotide changes. Research indicates that CNVs affect more nucleotides than individual base-pair changes, with about 2.7% of the genome affected by CNVs compared to 1.2% by SNPs. Moreover, while many CNVs are shared between humans and chimpanzees, a significant portion is unique to each species. Additionally, CNVs have been associated with genetic diseases in humans, highlighting their importance in human health. Despite this, many questions about CNVs remain unanswered, including their origin and contributions to evolutionary adaptation and disease. Ongoing research aims to address these questions using techniques like comparative genomic hybridization, which allows for a detailed examination of CNVs and their significance. When investigators examined the raw sequence data of the human and chimpanzee.[36]
Significance of comparative genomics
Comparative genomics holds profound significance across various fields, including medical research, basic biology, and biodiversity conservation. For instance, in medical research, predicting how genomic variants limited ability to predict which genomic variants lead to changes in organism-level phenotypes, such as increased disease risk in humans, remains challenging due to the immense size of the genome, comprising about three billion nucleotides.[37][38][39]
To tackle this challenge, comparative genomics offers a solution by pinpointing nucleotide positions that have remained unchanged over millions of years of evolution. These conserved regions indicate potential sites where genetic alterations could have detrimental effects on an organism's fitness, thus guiding the search for disease-causing variants. Moreover, comparative genomics holds promise in unraveling the mechanisms of gene evolution, environmental adaptations, gender-specific differences, and population variations across vertebrate lineages.[40]
Furthermore, comparative studies enable the identification of genomic signatures of selection—regions in the genome that have undergone preferential increase and fixation in populations due to their functional significance in specific processes.[41] For instance, in animal genetics, indigenous cattle exhibit superior disease resistance and environmental adaptability but lower productivity compared to exotic breeds. Through comparative genomic analyses, significant genomic signatures responsible for these unique traits can be identified. Using insights from this signature, breeders can make informed decisions to enhance breeding strategies and promote breed development.[42]
Methods
Computational approaches are necessary for genome comparisons, given the large amount of data encoded in genomes. Many tools are now publicly available, ranging from whole genome comparisons to gene expression analysis.[43] This includes approaches from systems and control, information theory, string analysis and data mining.[44] Computational approaches will remain critical for research and teaching, especially when information science and genome biology is taught in conjunction.[45]

Comparative genomics starts with basic comparisons of genome size and gene density. For instance, genome size is important for coding capacity and possibly for regulatory reasons. High gene density facilitates genome annotation, analysis of environmental selection. By contrast, low gene density hampers the mapping of genetic disease as in the human genome.
Sequence alignment
Alignments are used to capture information about similar sequences such as ancestry, common evolutionary descent, or common structure and function. Alignments can be done for both nucleotide and protein sequences.[47][48] Alignments consist of local or global pairwise alignments, and multiple sequence alignments. One way to find global alignments is to use a dynamic programming algorithm known as Needleman-Wunsch algorithm whereas Smith–Waterman algorithm used to find local alignments. With the exponential growth of sequence databases and the emergence of longer sequences, there's a heightened interest in faster, approximate, or heuristic alignment procedures. Among these, the FASTA and BLAST algorithms are prominent for local pairwise alignment. Recent years have witnessed the development of programs tailored to aligning lengthy sequences, such as MUMmer (1999), BLASTZ (2003), and AVID (2003). While BLASTZ adopts a local approach, MUMmer and AVID are geared towards global alignment. To harness the benefits of both local and global alignment approaches, one effective strategy involves integrating them. Initially, a rapid variant of BLAST known as BLAT is employed to identify homologous "anchor" regions. These anchors are subsequently scrutinized to identify sets exhibiting conserved order and orientation. Such sets of anchors are then subjected to alignment using a global strategy.
Additionally, ongoing efforts focus on optimizing existing algorithms to handle the vast amount of genome sequence data by enhancing their speed. Furthermore, MAVID stands out as another noteworthy pairwise alignment program specifically designed for aligning multiple genomes.
Pairwise Comparison: The Pairwise comparison of genomic sequence data is widely utilized in comparative gene prediction. Many studies in comparative functional genomics lean on pairwise comparisons, wherein traits of each gene are compared with traits of other genes across species. his method yields many more comparisons than unique observations, making each comparison dependent on others.[49][50]
Multiple comparisons: The comparison of multiple genomes is a natural extension of pairwise inter-specific comparisons. Such comparisons typically aim to identify conserved regions across two phylogenetic scales: 1. Deep comparisons, often referred to as phylogenetic footprinting[51] reveal conservation across higher taxonomic units like vertebrates.[52] 2. Shallow comparisons, recently termed Phylogenetic shadowing,[53] probe conservation across a group of closely related species.

Whole-genome alignment
Whole-genome alignment (WGA) involves predicting evolutionary relationships at the nucleotide level between two or more genomes. It integrates elements of colinear sequence alignment and gene orthology prediction, presenting a greater challenge due to the vast size and intricate nature of whole genomes. Despite its complexity, numerous methods have emerged to tackle this problem because WGAs play a crucial role in various genome-wide analyses, such as phylogenetic inference, genome annotation, and function prediction.[54] Thereby, SyRI (Synteny and Rearrangement Identifier) is one such method that utilizes whole genome alignment and it is designed to identify both structural and sequence differences between two whole-genome assemblies. By taking WGAs as input, SyRI initially scans for disparities in genome structures. Subsequently, it identifies local sequence variations within both rearranged and non-rearranged (syntenic) regions.[55]

Phylogenetic reconstruction
Another computational method for comparative genomics is phylogenetic reconstruction. It is used to describe evolutionary relationships in terms of common ancestors. The relationships are usually represented in a tree called a phylogenetic tree. Similarly, coalescent theory is a retrospective model to trace alleles of a gene in a population to a single ancestral copy shared by members of the population. This is also known as the most recent common ancestor. Analysis based on coalescence theory tries predicting the amount of time between the introduction of a mutation and a particular allele or gene distribution in a population. This time period is equal to how long ago the most recent common ancestor existed. The inheritance relationships are visualized in a form similar to a phylogenetic tree. Coalescence (or the gene genealogy) can be visualized using dendrograms.[56]
Genome maps
An additional method in comparative genomics is genetic mapping. In genetic mapping, visualizing synteny is one way to see the preserved order of genes on chromosomes. It is usually used for chromosomes of related species, both of which result from a common ancestor.[58] This and other methods can shed light on evolutionary history. A recent study used comparative genomics to reconstruct 16 ancestral karyotypes across the mammalian phylogeny. The computational reconstruction showed how chromosomes rearranged themselves during mammal evolution. It gave insight into conservation of select regions often associated with the control of developmental processes. In addition, it helped to provide an understanding of chromosome evolution and genetic diseases associated with DNA rearrangements.

Tools
Computational tools for analyzing sequences and complete genomes are developing quickly due to the availability of large amount of genomic data. At the same time, comparative analysis tools are progressed and improved. In the challenges about these analyses, it is very important to visualize the comparative results.[59]
Visualization of sequence conservation is a tough task of comparative sequence analysis. As we know, it is highly inefficient to examine the alignment of long genomic regions manually. Internet-based genome browsers provide many useful tools for investigating genomic sequences due to integrating all sequence-based biological information on genomic regions. When we extract large amount of relevant biological data, they can be very easy to use and less time-consuming.[59]
- UCSC Browser: This site contains the reference sequence and working draft assemblies for a large collection of genomes.[60]
- Ensembl: The Ensembl project produces genome databases for vertebrates and other eukaryotic species, and makes this information freely available online.[61]
- MapView: The Map Viewer provides a wide variety of genome mapping and sequencing data.[62]
- VISTA is a comprehensive suite of programs and databases for comparative analysis of genomic sequences. It was built to visualize the results of comparative analysis based on DNA alignments. The presentation of comparative data generated by VISTA can easily suit both small and large scale of data.[63]
- BlueJay Genome Browser: A stand-alone visualization tool for the multi-scale viewing of annotated genomes and other genomic elements.[64]
- SyRI: SyRI stands for Synteny and Rearrangement Identifier and is a versatile tool for comparative genomics, offering functionalities for synteny analysis and visualization, aiding in the prediction of genomic differences between related genomes using whole-genome assemblies (WGA).[65]
- Synmap2: Specifically designed for synteny mapping, Synmap2 efficiently compares genetic maps or assemblies, providing insights into genome evolution and rearrangements among related organisms.[66]
- GSAlign: GSAlign facilitates accurate alignment of genomic sequences, particularly useful for large-scale comparative genomics studies, enabling researchers to identify similarities and differences across genomes.[67]
- IGV (Integrative Genomics Viewer): A widely used tool for visualizing and analyzing genomic data, IGV supports comparative genomics by enabling users to explore alignments, variants, and annotations across multiple genomes.[68]
- Manta: Manta is a rapid structural variant caller, crucial for comparative genomics as it detects genomic rearrangements such as insertions, deletions, inversions, and duplications, aiding in understanding genetic variation among populations or species.[69]
- CNVNatar: CNVNatar specializes in detecting copy number variations (CNVs), which are crucial in understanding genome evolution and population genetics, providing insights into genomic structural changes across different organisms.[70]
- PIPMaker: PIPMaker facilitates the alignment and comparison of two genomic sequences, enabling the identification of conserved regions, duplications, and evolutionary breakpoints, aiding in comparative genomics analyses.[71]
- GLASS (Genome-wide Location and Sequence Searcher): GLASS is a tool for identifying conserved regulatory elements across genomes, crucial for comparative genomics studies focusing on understanding gene regulation and evolution.[72]
- PatternHunter: PatternHunter is a versatile tool for sequence analysis, offering functionalities for identifying conserved patterns, motifs, and repeats across genomic sequences, aiding in comparative genomics studies of gene families and regulatory elements.
- Mummer: Mummer is a suite of tools for whole-genome alignment and comparison, widely used in comparative genomics for identifying similarities, differences, and evolutionary events among genomes at various scales.[73]
An advantage of using online tools is that these websites are being developed and updated constantly. There are many new settings and content can be used online to improve efficiency.[59]
Selected applications
Agriculture
Agriculture is a field that reaps the benefits of comparative genomics. Identifying the loci of advantageous genes is a key step in breeding crops that are optimized for greater yield, cost-efficiency, quality, and disease resistance. For example, one genome wide association study conducted on 517 rice landraces revealed 80 loci associated with several categories of agronomic performance, such as grain weight, amylose content, and drought tolerance. Many of the loci were previously uncharacterized.[74] Not only is this methodology powerful, it is also quick. Previous methods of identifying loci associated with agronomic performance required several generations of carefully monitored breeding of parent strains, a time-consuming effort that is unnecessary for comparative genomic studies.[75]
Medicine
Vaccine development
The medical field also benefits from the study of comparative genomics. In an approach known as reverse vaccinology, researchers can discover candidate antigens for vaccine development by analyzing the genome of a pathogen or a family of pathogens.[76] Applying a comparative genomics approach by analyzing the genomes of several related pathogens can lead to the development of vaccines that are multi-protective. A team of researchers employed such an approach to create a universal vaccine for Group B Streptococcus, a group of bacteria responsible for severe neonatal infection.[77] Comparative genomics can also be used to generate specificity for vaccines against pathogens that are closely related to commensal microorganisms. For example, researchers used comparative genomic analysis of commensal and pathogenic strains of E. coli to identify pathogen-specific genes as a basis for finding antigens that result in immune response against pathogenic strains but not commensal ones.[78] In May 2019, using the Global Genome Set, a team in the UK and Australia sequenced thousands of globally collected isolates of Group A Streptococcus, providing potential targets for developing a vaccine against the pathogen, also known as S. pyogenes.[79]
Personalized Medicine
Personalized Medicine, enabled by Comparative Genomics, represents a revolutionary approach in healthcare, tailoring medical treatment and disease prevention to the individual patient's genetic makeup.[80] By analyzing genetic variations across populations and comparing them with an individual's genome, clinicians can identify specific genetic markers associated with disease susceptibility, drug metabolism, and treatment response. By identifying genetic variants associated with drug metabolism pathways, drug targets, and adverse reactions, personalized medicine can optimize medication selection, dosage, and treatment regimens for individual patients. This approach minimizes the risk of adverse drug reactions, enhances treatment efficacy, and improves patient outcomes.
Cancer
Cancer Genomics represents a cutting-edge field within oncology that leverages comparative genomics to revolutionize cancer diagnosis, treatment, and prevention strategies. Comparative genomics plays a crucial role in cancer research by identifying driver mutations, and providing comprehensive analyses of mutations, copy number alterations, structural variants, gene expression, and DNA methylation profiles in large-scale studies across different cancer types. By analyzing the genomes of cancer cells and comparing them with healthy cells, researchers can uncover key genetic alterations driving tumorigenesis, tumor progression, and metastasis. This deep understanding of the genomic landscape of cancer has profound implications for precision oncology. Moreover, Comparative Genomics is instrumental in elucidating mechanisms of drug resistance—a major challenge in cancer treatment.

Mouse models in immunology
T cells (also known as a T lymphocytes or a thymocytes) are immune cells that grow from stem cells in the bone marrow. They assist to defend the body from infection and may aid in the fight against cancer. Because of their morphological, physiological, and genetic resemblance to humans, mice and rats have long been the preferred species for biomedical research animal models. Comparative Medicine Research is built on the ability to use information from one species to understand the same processes in another. We can get new insights into molecular pathways by comparing human and mouse T cells and their effects on the immune system utilizing comparative genomics. In order to comprehend its TCRs and their genes, Glusman conducted research on the sequencing of the human and mouse T cell receptor loci. TCR genes are well-known and serve as a significant resource for supporting functional genomics and understanding how genes and intergenic regions of the genome contribute to biological processes.[81]
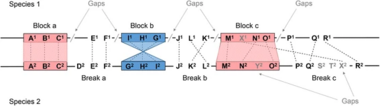
T-cell immune receptors are important in seeing the world of pathogens in the cellular immune system. One of the reasons for sequencing the human and mouse TCR loci was to match the orthologous gene family sequences and discover conserved areas using comparative genomics. These, it was thought, would reflect two sorts of biological information: (1) exons and (2) regulatory sequences. In fact, the majority of V, D, J, and C exons could be identified in this method. The variable regions are encoded by multiple unique DNA elements that are rearranged and connected during T cell (TCR) differentiation: variable (V), diversity (D), and joining (J) elements for the and polypeptides; and V and J elements for the and polypeptides.[Figure 1] However, several short noncoding conserved blocks of the genome had been shown. Both human and mouse motifs are largely clustered in the 200 bp [Figure 2], the known 3′ enhancers in the TCR/ were identified, and a conserved region of 100 bp in the mouse J intron was subsequently shown to have a regulatory function.

Comparisons of the genomic sequences within each physical site or location of a specific gene on a chromosome (locs) and across species allow for research on other mechanisms and other regulatory signals. Some suggest new hypotheses about the evolution of TCRs, to be tested (and improved) by comparison to the TCR gene complement of other vertebrate species. A comparative genomic investigation of humans and mice will obviously allow for the discovery and annotation of many other genes, as well as identifying in other species for regulatory sequences.[81]
Research
Comparative genomics also opens up new avenues in other areas of research. As DNA sequencing technology has become more accessible, the number of sequenced genomes has grown. With the increasing reservoir of available genomic data, the potency of comparative genomic inference has grown as well.
A notable case of this increased potency is found in recent primate research. Comparative genomic methods have allowed researchers to gather information about genetic variation, differential gene expression, and evolutionary dynamics in primates that were indiscernible using previous data and methods.[82]
Comparative genomics has also become increasingly important in microbial ecology and plant–microbe interaction research. By comparing large collections of bacterial genomes from different environments, researchers have been able to identify genes and functional pathways associated with adaptation to specific ecological niches, such as plant-associated versus free-living lifestyles. When combined with transcriptomic or metagenomic data, comparative genomic approaches can link gene conservation and abundance to ecological relevance, revealing bacterial genes that are consistently enriched and transcriptionally active in plant-associated environments. These studies have shown that bacterial adaptation to plants often involves numerous, previously uncharacterized genes rather than a small number of well-known determinants, highlighting the role of comparative genomics in uncovering hidden functional diversity and evolutionary specialization in microbial genomes.[83][84][85]
Great Ape Genome Project
The Great Ape Genome Project used comparative genomic methods to investigate genetic variation with reference to the six great ape species, finding healthy levels of variation in their gene pool despite shrinking population size.[86] Another study showed that patterns of DNA methylation, which are a known regulation mechanism for gene expression, differ in the prefrontal cortex of humans versus chimps, and implicated this difference in the evolutionary divergence of the two species.[87]
See also
- Data mining
- Molecular evolution
- Comparative anatomy
- Homology
- Sequence mining
- Alignment-free sequence analysis
References
- ↑ "Dynamics of genome rearrangement in bacterial populations". PLOS Genetics 4 (7). July 2008. doi:10.1371/journal.pgen.1000128. PMID 18650965.
- ↑ 2.0 2.1 2.2 2.3 "Comparative Genomics". Nature Education Knowledge 3 (10): 13. 2010. http://www.nature.com/scitable/knowledge/library/comparative-genomics-13239404.
- ↑ Comparative Genomics. SpringerBriefs in Genetics. Heidelberg: Springer. 2013. doi:10.1007/978-3-642-37146-2. ISBN 978-3-642-37145-5.
- ↑ 4.0 4.1 "Comparative genomics approaches to study organism similarities and differences". Journal of Biomedical Informatics 35 (2): 142–150. April 2002. doi:10.1016/s1532-0464(02)00506-3. PMID 12474427.
- ↑ "Comparisons with Caenorhabditis (approximately 100 Mb) and Drosophila (approximately 175 Mb) using flow cytometry show genome size in Arabidopsis to be approximately 157 Mb and thus approximately 25% larger than the Arabidopsis genome initiative estimate of approximately 125 Mb". Annals of Botany 91 (5): 547–557. April 2003. doi:10.1093/aob/mcg057. PMID 12646499.
- ↑ "A whole-genome assembly of the domestic cow, Bos taurus". Genome Biology 10 (4). 2009. doi:10.1186/gb-2009-10-4-r42. PMID 19393038. Bibcode: 2009GenBi..10..R42Z.
- ↑ "Chromosomal Aberrations in Cattle". Genes 12 (9): 1330. August 2021. doi:10.3390/genes12091330. PMID 34573313.
- ↑ "The genome sequence of taurine cattle: a window to ruminant biology and evolution". Science 324 (5926): 522–528. April 2009. doi:10.1126/science.1169588. PMID 19390049. Bibcode: 2009Sci...324..522A.
- ↑ "Inferring synteny between genome assemblies: a systematic evaluation". BMC Bioinformatics 19 (1). January 2018. doi:10.1186/s12859-018-2026-4. PMID 29382321. Bibcode: 2018BMCBi..19...26L.
- ↑ "Large synteny blocks revealed between Caenorhabditis elegans and Caenorhabditis briggsae genomes using OrthoCluster". BMC Genomics 11: 516. September 2010. doi:10.1186/1471-2164-11-516. PMID 20868500.
- ↑ "Screening synteny blocks in pairwise genome comparisons through integer programming". BMC Bioinformatics 12. April 2011. doi:10.1186/1471-2105-12-102. PMID 21501495.
- ↑ "Synteny conservation and chromosome rearrangements during mammalian evolution". Genetics 147 (1): 289–296. September 1997. doi:10.1093/genetics/147.1.289. PMID 9286688.
- ↑ "Comparative genomic data of the Avian Phylogenomics Project". GigaScience 3 (1). 2014-12-11. doi:10.1186/2047-217X-3-26. PMID 25671091.
- ↑ "WormBase 2016: expanding to enable helminth genomic research". Nucleic Acids Research 44 (D1): D774–D780. January 2016. doi:10.1093/nar/gkv1217. PMID 26578572.
- ↑ 15.0 15.1 The C. elegans Sequencing Consortium (December 1998). "Genome sequence of the nematode C. elegans: a platform for investigating biology". Science 282 (5396): 2012–2018. doi:10.1126/science.282.5396.2012. PMID 9851916.
- ↑ "Birth of a metabolic gene cluster in yeast by adaptive gene relocation". Nature Genetics 37 (7): 777–782. July 2005. doi:10.1038/ng1584. PMID 15951822. Bibcode: 2005NaGen..37..777W.
- ↑ "Cancer: Genomic evolution of metastasis". Nature 467 (7319): 1053–1055. October 2010. doi:10.1038/4671053a. PMID 20981088. Bibcode: 2010Natur.467.1053L.
- ↑ "FLOWERING LOCUS C in monocots and the tandem origin of angiosperm-specific MADS-box genes". Nature Communications 4. 2013. doi:10.1038/ncomms3280. PMID 23955420. Bibcode: 2013NatCo...4.2280R.
- ↑ "Gene synteny comparisons between different vertebrates provide new insights into breakage and fusion events during mammalian karyotype evolution". BMC Evolutionary Biology 9 (1). April 2009. doi:10.1186/1471-2148-9-84. PMID 19393055. Bibcode: 2009BMCEE...9...84K.
- ↑ 20.0 20.1 20.2 20.3 Sequence - Evolution - Function: Computational approaches in comparative genomics. Dordrecht: Springer Science+Business Media. 2003.
- ↑ 21.0 21.1 "Pathogen comparative genomics in the next-generation sequencing era: genome alignments, pangenomics and metagenomics". Briefings in Functional Genomics 10 (6): 322–333. November 2011. doi:10.1093/bfgp/elr042. PMID 22199376.
- ↑ Biology: The Dynamic Science (2nd ed.). Belmont, CA: Brooks/Cole. 2011. pp. 409–410.
- ↑ "Similarity in gene organization and homology between proteins of animal picornaviruses and a plant comovirus suggest common ancestry of these virus families". Nucleic Acids Research 12 (18): 7251–7267. September 1984. doi:10.1093/nar/12.18.7251. PMID 6384934.
- ↑ "DNA sequence of the herpes simplex virus type 1 gene encoding glycoprotein gH, and identification of homologues in the genomes of varicella-zoster virus and Epstein-Barr virus". Nucleic Acids Research 14 (10): 4281–4292. May 1986. doi:10.1093/nar/14.10.4281. PMID 3012465.
- ↑ "Whole-genome random sequencing and assembly of Haemophilus influenzae Rd". Science 269 (5223): 496–512. July 1995. doi:10.1126/science.7542800. PMID 7542800. Bibcode: 1995Sci...269..496F.
- ↑ "The minimal gene complement of Mycoplasma genitalium". Science 270 (5235): 397–403. October 1995. doi:10.1126/science.270.5235.397. PMID 7569993. Bibcode: 1995Sci...270..397F.
- ↑ "Life with 6000 genes". Science 274 (5287): 546, 563-546, 567. October 1996. doi:10.1126/science.274.5287.546. PMID 8849441. Bibcode: 1996Sci...274..546G.
- ↑ "The genome sequence of Drosophila melanogaster". Science 287 (5461): 2185–2195. March 2000. doi:10.1126/science.287.5461.2185. PMID 10731132. Bibcode: 2000Sci...287.2185..
- ↑ "Comparative genomics of the eukaryotes". Science 287 (5461): 2204–2215. March 2000. doi:10.1126/science.287.5461.2204. PMID 10731134. Bibcode: 2000Sci...287.2204..
- ↑ "Human and mouse gene structure: comparative analysis and application to exon prediction". Genome Research 10 (7): 950–958. July 2000. doi:10.1101/gr.10.7.950. PMID 10899144.
- ↑ "Comparative genomics: genome-wide analysis in metazoan eukaryotes". Nature Reviews. Genetics 4 (4): 251–262. April 2003. doi:10.1038/nrg1043. PMID 12671656.
- ↑ "The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics". PLOS Biology 1 (2). November 2003. doi:10.1371/journal.pbio.0000045. PMID 14624247.
- ↑ "Newly Sequenced Worm a Boon for Worm Biologists". PLOS Biology 1 (2). 2003. doi:10.1371/journal.pbio.0000044.
- ↑ "Analysis of copy number variations among diverse cattle breeds". Genome Research 20 (5): 693–703. May 2010. doi:10.1101/gr.105403.110. PMID 20212021.
- ↑ "Analysis of genomic copy number variations through whole-genome scan in Chinese Qaidam cattle". Frontiers in Veterinary Science 10. 2023. doi:10.3389/fvets.2023.1148070. PMID 37065216.
- ↑ "Copy Number Variation | Learn Science at Scitable" (in en). http://www.nature.com/scitable/topicpage/copy-number-variation-445.
- ↑ "The NIH Comparative Genomics Resource: addressing the promises and challenges of comparative genomics on human health". BMC Genomics 24 (1). September 2023. doi:10.1186/s12864-023-09643-4. PMID 37759191.
- ↑ "A comparative genomics multitool for scientific discovery and conservation". Nature 587 (7833): 240–245. November 2020. doi:10.1038/s41586-020-2876-6. PMID 33177664. Bibcode: 2020Natur.587..240Z.
- ↑ "Genomic Analysis in the Age of Human Genome Sequencing". Cell 177 (1): 70–84. March 2019. doi:10.1016/j.cell.2019.02.032. PMID 30901550.
- ↑ "A general framework for estimating the relative pathogenicity of human genetic variants". Nature Genetics 46 (3): 310–315. March 2014. doi:10.1038/ng.2892. PMID 24487276. Bibcode: 2014NaGen..46..310K.
- ↑ "Genomic Signature in Evolutionary Biology: A Review". Biology 12 (2): 322. February 2023. doi:10.3390/biology12020322. PMID 36829597.
- ↑ "Differences in innate and adaptive immune response traits of Pahari (Indian non-descript indigenous breed) and Jersey crossbred cattle". Veterinary Immunology and Immunopathology 192: 20–27. October 2017. doi:10.1016/j.vetimm.2017.09.003. PMID 29042011.
- ↑ Introduction to Computational Genomics. Cambridge University Press. 2006. ISBN 978-0-521-67191-0. http://www.computational-genomics.net.
- ↑ "An alignment-free method to find and visualise rearrangements between pairs of DNA sequences". Scientific Reports 5. May 2015. doi:10.1038/srep10203. PMID 25984837. Bibcode: 2015NatSR...510203P.
- ↑ "Ten simple rules for developing a short bioinformatics training course". PLOS Computational Biology 7 (10). October 2011. doi:10.1371/journal.pcbi.1002245. PMID 22046119. Bibcode: 2011PLSCB...7E2245V.
- ↑ 46.0 46.1 "Evolution of the ancestral mammalian karyotype and syntenic regions". Proceedings of the National Academy of Sciences of the United States of America 119 (40). October 2022. doi:10.1073/pnas.2209139119. PMID 36161960. Bibcode: 2022PNAS..11909139D.
- ↑ "Sequence Alignment". Handbook of Discrete and Combinatorial Mathematics (2nd ed.). Boca Raton (FL): CRC Press/Taylor & Francis. 2017. ISBN 978-1-58488-780-5. https://www.ncbi.nlm.nih.gov/books/NBK464187/. Retrieved 2022-12-18.
- ↑ "Sequence Analysis" (in en). Encyclopedia of Bioinformatics and Computational Biology. Oxford: Academic Press. 2019-01-01. pp. 292–322. doi:10.1016/b978-0-12-809633-8.20106-4. ISBN 978-0-12-811432-2.
- ↑ "Comparative genomics: methods and applications". Die Naturwissenschaften 91 (9): 405–421. September 2004. doi:10.1007/s00114-004-0542-8. PMID 15278216. Bibcode: 2004NW.....91..405H.
- ↑ "Pairwise comparisons across species are problematic when analyzing functional genomic data". Proceedings of the National Academy of Sciences of the United States of America 115 (3): E409–E417. January 2018. doi:10.1073/pnas.1707515115. PMID 29301966. Bibcode: 2018PNAS..115E.409D.
- ↑ "Long human-mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome". Genome Research 7 (10): 959–966. October 1997. doi:10.1101/gr.7.10.959. PMID 9331366.
- ↑ "Small is beautiful: comparative genomics with the pufferfish (Fugu rubripes)". Trends in Genetics 12 (4): 145–150. April 1996. doi:10.1016/0168-9525(96)10018-4. PMID 8901419. http://scholarbank.nus.edu.sg/handle/10635/112196.
- ↑ "Phylogenetic shadowing of primate sequences to find functional regions of the human genome". Science 299 (5611): 1391–1394. February 2003. doi:10.1126/science.1081331. PMID 12610304. Bibcode: 2003Sci...299.1391B. https://digital.library.unt.edu/ark:/67531/metadc779156/.
- ↑ "Whole-Genome Alignment". Evolutionary Genomics. Methods in Molecular Biology. 855. Totowa, NJ: Humana Press. 2012. pp. 237–257. doi:10.1007/978-1-61779-582-4_8. ISBN 978-1-61779-581-7.
- ↑ "SyRI: Finding genomic rearrangements and local sequence differences from whole-genome assemblies". Genome Biology 20 (1). 2019. doi:10.1186/s13059-019-1911-0. PMID 31842948.
- ↑ "Comparative genomics: methods and applications". Die Naturwissenschaften 91 (9): 405–421. September 2004. doi:10.1007/s00114-004-0542-8. PMID 15278216. Bibcode: 2004NW.....91..405H.
- ↑ "Inferring synteny between genome assemblies: a systematic evaluation". BMC Bioinformatics 19 (1). January 2018. doi:10.1186/s12859-018-2026-4. PMID 29382321. Bibcode: 2018BMCBi..19...26L.
- ↑ "Genetic Maps and the Use of Synteny". Plant Genomics. Methods in Molecular Biology. 513. 2009. pp. 41–55. doi:10.1007/978-1-59745-427-8_3. ISBN 978-1-58829-997-0.
- ↑ 59.0 59.1 59.2 Comparative Genomics: Volumes 1 and 2. Totowa, New Jersey: Humana Press. 2007. ISBN 978-193411-537-4. https://www.ncbi.nlm.nih.gov/books/NBK1732/.
- ↑ "UCSC Browser". http://genome.ucsc.edu/.
- ↑ "Ensembl Genome Browser". http://asia.ensembl.org/index.html.
- ↑ "Map Viewer". https://www.ncbi.nlm.nih.gov/mapview/.
- ↑ "VISTA tools". http://genome.lbl.gov/vista/index.shtml.
- ↑ "The Bluejay genome browser". Current Protocols in Bioinformatics (John Wiley & Sons, Inc.) 37: Chapter 10, Unit 10.9. March 2012. doi:10.1002/0471250953.bi1009s37. ISBN 978-0-471-25095-1. PMID 22389011.
- ↑ "SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies". Genome Biology 20 (1). December 2019. doi:10.1186/s13059-019-1911-0. PMID 31842948.
- ↑ "SynMap2 and SynMap3D: web-based whole-genome synteny browsers". Bioinformatics 33 (14): 2197–2198. July 2017. doi:10.1093/bioinformatics/btx144. PMID 28334338.
- ↑ "GSAlign: an efficient sequence alignment tool for intra-species genomes". BMC Genomics 21 (1). February 2020. doi:10.1186/s12864-020-6569-1. PMID 32093618.
- ↑ "Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration". Briefings in Bioinformatics 14 (2): 178–192. March 2013. doi:10.1093/bib/bbs017. PMID 22517427.
- ↑ "Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications". Bioinformatics 32 (8): 1220–1222. April 2016. doi:10.1093/bioinformatics/btv710. PMID 26647377.
- ↑ "CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing". Genome Research 21 (6): 974–984. June 2011. doi:10.1101/gr.114876.110. PMID 21324876.
- ↑ "PipMaker: a World Wide Web server for genomic sequence alignments". Current Protocols in Bioinformatics Chapter 10: Chapter 10, Unit 10.2. February 2003. doi:10.1002/0471250953.bi1002s00. PMID 18428692.
- ↑ "GLASS: assisted and standardized assessment of gene variations from Sanger sequence trace data". Bioinformatics 33 (23): 3802–3804. December 2017. doi:10.1093/bioinformatics/btx423. PMID 29036643.
- ↑ "MUMmer4: A fast and versatile genome alignment system". PLOS Computational Biology 14 (1). January 2018. doi:10.1371/journal.pcbi.1005944. PMID 29373581. Bibcode: 2018PLSCB..14E5944M.
- ↑ "Genome-wide association studies of 14 agronomic traits in rice landraces". Nature Genetics 42 (11): 961–967. November 2010. doi:10.1038/ng.695. PMID 20972439.
- ↑ "Crop genomics: advances and applications". Nature Reviews. Genetics 13 (2): 85–96. December 2011. doi:10.1038/nrg3097. PMID 22207165.
- ↑ "Developing vaccines in the era of genomics: a decade of reverse vaccinology". Clinical Microbiology and Infection 18 (Suppl 5): 109–116. October 2012. doi:10.1111/j.1469-0691.2012.03939.x. PMID 22882709.
- ↑ "Identification of a universal Group B streptococcus vaccine by multiple genome screen". Science 309 (5731): 148–150. July 2005. doi:10.1126/science.1109869. PMID 15994562. Bibcode: 2005Sci...309..148M.
- ↑ "The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates". Journal of Bacteriology 190 (20): 6881–6893. October 2008. doi:10.1128/JB.00619-08. PMID 18676672. Bibcode: 2008JBact.190.6881R.
- ↑ "Group a Streptococcus Vaccine Target Candidates Identified from Global Genome Set". 28 May 2019. https://www.genomeweb.com/sequencing/group-streptococcus-vaccine-target-candidates-identified-global-genome-set#.XRKFu_ZFxPY.
- ↑ "Genomics and personalized medicine". International Journal of Pharmaceutics 415 (1–2): 2–4. August 2011. doi:10.1016/j.ijpharm.2011.04.048. PMID 21539903.
- ↑ 81.0 81.1 81.2 81.3 "Comparative genomics of the human and mouse T cell receptor loci". Immunity 15 (3): 337–349. September 2001. doi:10.1016/s1074-7613(01)00200-x. PMID 11567625.
- ↑ "Comparative primate genomics: emerging patterns of genome content and dynamics". Nature Reviews. Genetics 15 (5): 347–359. May 2014. doi:10.1038/nrg3707. PMID 24709753.
- ↑ Saati-Santamaría, Zaki; Baroncelli, Riccardo; Rivas, Raúl; García-Fraile, Paula (2022-11-10). "Comparative Genomics of the Genus Pseudomonas Reveals Host- and Environment-Specific Evolution". Microbiology Spectrum 10 (6): e02370–22. doi:10.1128/spectrum.02370-22. PMID 36354324.
- ↑ Levy, Asaf; Salas Gonzalez, Isai; Mittelviefhaus, Maximilian; Clingenpeel, Scott; Herrera Paredes, Sur; Miao, Jiamin; Wang, Kunru; Devescovi, Giulia et al. (2017-12-18). "Genomic features of bacterial adaptation to plants". Nature Genetics 50 (1): 138–150. doi:10.1038/s41588-017-0012-9. ISSN 1061-4036. PMID 29255260.
- ↑ Saati-Santamaría, Zaki; González-Dominici, Lihuén Iraí; Jiménez-Gómez, Alejandro; Morais, Daniel; Tláskal, Vojtěch; Frontela, Irene; Benada, Oldřich; Qi, Li et al. (2025-12-05). "Transcriptome-guided discovery of novel plant-associated genes in a rhizosphere Pseudomonas". Microbiome 14 (1). doi:10.1186/s40168-025-02277-6. ISSN 2049-2618.
- ↑ "Great ape genetic diversity and population history". Nature 499 (7459): 471–475. July 2013. doi:10.1038/nature12228. PMID 23823723. Bibcode: 2013Natur.499..471P.
- ↑ "Divergent whole-genome methylation maps of human and chimpanzee brains reveal epigenetic basis of human regulatory evolution". American Journal of Human Genetics 91 (3): 455–465. September 2012. doi:10.1016/j.ajhg.2012.07.024. PMID 22922032.
Further reading
- "Sequencing and comparison of yeast species to identify genes and regulatory elements". Nature 423 (6937): 241–254. May 2003. doi:10.1038/nature01644. PMID 12748633. Bibcode: 2003Natur.423..241K.
- "Finding functional features in Saccharomyces genomes by phylogenetic footprinting". Science 301 (5629): 71–76. July 2003. doi:10.1126/science.1084337. PMID 12775844. Bibcode: 2003Sci...301...71C.
- "Phylogenetic shadowing of primate sequences to find functional regions of the human genome". Science 299 (5611): 1391–1394. February 2003. doi:10.1126/science.1081331. PMID 12610304. Bibcode: 2003Sci...299.1391B. https://digital.library.unt.edu/ark:/67531/metadc779156/.
- "Genome evolution in yeasts". Nature 430 (6995): 35–44. July 2004. doi:10.1038/nature02579. PMID 15229592. Bibcode: 2004Natur.430...35D.
- "Comparative genomics in eukaryotes". The Evolution of the Genome. San Diego: Elsevier. 2005. pp. 521–583.
- "Comparative genomics in prokaryotes". The Evolution of the Genome. San Diego: Elsevier. 2005. pp. 585–675.
- "Systematic discovery of regulatory motifs in human promoters and 3' UTRs by comparison of several mammals". Nature 434 (7031): 338–345. March 2005. doi:10.1038/nature03441. PMID 15735639. Bibcode: 2005Natur.434..338X.
- "Genome update: purine strand bias in 280 bacterial genomes". Microbiology 152 (Pt 3): 579–583. March 2006. doi:10.1099/mic.0.28637-0. PMID 16514138.
- "Systematic discovery of regulatory motifs in Fusarium graminearum by comparing four Fusarium genomes". BMC Genomics 11. March 2010. doi:10.1186/1471-2164-11-208. PMID 20346147.
- "Human and mouse gene structure: comparative analysis and application to exon prediction". Genome Research 10 (7): 950–958. July 2000. doi:10.1101/gr.10.7.950. PMID 10899144.
External links
 |
