Biology:Connectomics
Connectomics is the production and study of connectomes, which are comprehensive maps of connections within an organism's nervous system. Study of neuronal wiring diagrams looks at how they contribute to the health and behavior of an organism. There are two very different types of connectomes; microscale and macroscale. Microscale connectomics maps every neuron and synapse in an organism or chunk of tissue, typically using electron microscopy and histology. This level of detail is only possible for small animals (flies and worms) or tiny portions (less than 1 mm on a side) of large animal brains. Macroscale connectomics, on the other hand, refers to mapping out large fiber tracts and functional gray matter areas within a much larger brain (typically human), typically using forms of MRI to map out structure and function. Both fields simply refer to their maps as "connectomes".
Macroscale connectomics typically concentrates on the human nervous system, a network made of up to billions of connections and responsible for our thoughts, emotions, actions, memories, function and dysfunction. Because these structures are physically large and experiments on humans are routinely non-invasive, typical methods are functional and structural MRI data to measure blood flow (functional) and water diffusivity (structural). Examples include the Human Connectome Project and others.[1][2] Connectomics in this regime aims to advance our understanding of mental health and cognition by understanding how cells in the nervous system are connected and communicate.
In contrast, microscale connectomics looks in much greater detail at much smaller circuits, such as the worm C. elegans, the fruit fly Drosophila,[3] and portions of mammal brains such as the retina[4] and cortex. Connectomics at these scales searches for mechanistic explanations of how the nervous system operates.
Methods
Macroscale Connectomics
Macroscale connectomes are commonly collected using diffusion-weighted magnetic resonance imaging (DW-MRI) and functional magnetic resonance imaging (fMRI). DW-MRI datasets can span the entire brain, imaging white matter between the cortex and subcortex, providing information about the diffusion of water molecules in brain tissue, and allowing researchers to infer the orientation and integrity of white matter pathways.[5] DW-MRI can be used in conjunction with tractography where it enables the reconstruction of white matter tracts in the brain.[5] It does so by measuring the diffusion of water molecules in multiple directions, as DW-MRI can estimate the local fiber orientations and generate a model of the brain's fiber pathways.[5] Meanwhile, tractography algorithms trace the likely trajectories of these pathways, providing a representation of the brain's anatomical connectivity.[5] Metrics such as fractional anisotropy (FA), mean diffusivity (MD), or connectivity strength can be computed from DW-MRI data to assess the microstructural properties of white matter and quantify the strength of (long-range) connections between brain regions.[6]
In contrast to DW-MRI, fMRI measures the blood oxygenation level-dependent (BOLD) signal, which reflects changes in cerebral blood flow and oxygenation associated with neural activity, as regulated by the neurovascular unit.[7] When used together, a resting-state fMRI and a DW-MRI dataset provide a comprehensive view of how regions of the brain are structurally connected, and how closely they are communicating.[8][9][10] Resting-state functional connectivity (RSFC) analysis is a common method to measure connectomes using fMRI that involves acquiring fMRI data while the subject is at rest and not performing any specific tasks or stimuli.[11] RSFC examines the temporal correlation of the BOLD signals between different brain regions (after accounting for the confounding effect of other regions), providing insights into functional connectivity.[7]
Stimulation
Techniques that actively manipulate the brain, often called neuromodulation, can provide insights into the connectome.[12] For example, transcranial magnetic stimulation (TMS) is a non-invasive neuromodulation technique that applies strong magnetic pulses between scalp electrodes which target specific brain regions with electrical currents.[13] This can temporarily disrupt or enhance the activity of specific brain areas and observe changes in connectivity.[13] Transcranial direct current stimulation (tDCS) is another non-invasive neuromodulation technique that applies a constant but relatively weak electrical current for a few minutes, modulating neuronal excitability.[14] It allows researchers to investigate the causal relationship between targeted brain regions and changes in connectivity.[14] tDCS increases the functional connectivity within the brain, with a bias towards specific networks (e.g., cortical processing), and may even cause structural changes to take place in the white matter via myelination and in the gray matter via synaptic plasticity.[14] Another neuromodulation technique is deep brain stimulation (DBS), an invasive technique that involves surgically implanting electrodes into specific brain regions in order to apply localized, high-frequency electrical impulses.[15] This technique modulates brain networks and is often used to alleviate motor symptoms from disorders like Parkinson's, essential tremor, and dystonia.[16] The functional and structural connectivity between electrodes can be used to predict patient outcomes and estimate optimal connectivity profiles.[15]
Electrophysiological Methods
Electrophysiological methods measure the difference in signals from different parts of the brain to estimate the connectivity between them, a process that requires a low signal-to-noise ratio to maintain the accuracy of the measurements and sufficient spatial resolution to support the connectivity between specific regions of the brain.[17] These methods offer insights into real-time neural dynamics and functional connectivity between brain regions. Electroencephalography (EEG) measures the differences in the electrical potential generated by oscillating currents at the surface of the scalp, due to the non-invasive, external placement of the electrodes.[18] Meanwhile, magnetoencephalography (MEG) relies on the magnetic fields generated by the electrical activity of the brain to collect information.[18]
Microscale Connectomics
Microscale connectomes focuses on resolving individual cell-to-cell connectivity within much smaller volumes of nervous system tissue. The most common method for neural circuit reconstruction is chemical brain preservation followed by 3D electron microscopy,[19] which offers single synapse resolution. The first microscale connectome encompassing an entire nervous system was produced for the nematode C. elegans in 1986.[20] This was done by manually annotating printouts of the EM scans.[20] Advances in EM acquisition, image alignment and segmentation, and manipulation of large datasets have since allowed for larger volumes to be imaged and segmented more easily. EM has been used to produce connectomes from a variety of nervous system samples, including publicly available datasets that encompass the full body of a Platynereis dumerilii larva,[21] the entire brain[22][23] and ventral nerve cord[24][25] of adult Drosophila melanogaster, the full central nervous system (connected brain and ventral nerve cord) of larval Drosophila melanogaster,[26] and volumes from mouse[27] and human cortex.[28][29] The National Institutes of Health (NIH) has now invested in creating an EM connectome of an entire mouse brain.[30] EM can be combined with fluorescence in correlative microscopy, which can generate more interpretable data as is it able to automatically detect specific neuron types and can trace them in their entirety using fluorescent markers.[31]
Other imaging modalities are approaching the nanometer-scale resolution necessary for microscale connectomics. X-ray nanotomography using a synchrotron source can now reach <100 nm resolution, and can theoretically continue to improve.[32] Unlike EM, this technique does not require the tissue being imaged to be stained with heavy metals or to be physically sectioned.[32] Conventional light microscopy is constrained by light diffraction. Researchers have recently used stimulated emission depletion (STED) microscopy, a super-resolution light microscopy technique, to image the extracellular space of living human brain organoids and mouse hippocampal slice cultures, allowing for reconstruction of all neurites within this volume by implementing a two-stage machine learning approach.[33] They combined this with fluorescently-tagged synaptic markers to find synaptic connectivity in the sample as well as with calcium imaging to monitor neuronal activity .[33] However, this live-imaging approach was limited to ~130 nm resolution, and was therefore not able to reliably reconstruct thin axons over long distances.[33] In 2024, a new technique called LICONN combined hydrogel expansion with light microscopy (as opposed to electron microscopy) to generate neuron level connectomes.[34] Potential advantages include cheaper equipment (optical vs EM microscopes), faster data acquisition, and multi-color labelling.
Software
In addition to advanced microscopy techniques, connectomics heavily relies on software analysis tools and machine learning pipelines for reconstructing and analyzing neural networks. These tools are designed to process and interpret the vast amounts of data generated by volume electron microscopy and other imaging methods. Key steps in connectomic reconstruction include image segmentation, where individual neurons and their components are identified and annotated, and network mapping, where the connections between these neurons are established.[35]
Several software platforms facilitate these tasks. CATMAID (Collaborative Annotation Toolkit for Massive Amounts of Image Data) is a decentralized web interface allowing seamless navigation of large image stacks. It is designed to facilitate the collaborative exploration, annotation, and efficient sharing of regions of interests by bookmarking.[36] Another example is WEBKNOSSOS, an online platform used for viewing, annotating, and sharing large 3D images, aiding in the detailed analysis of neural structures by allowing efficient navigation and annotation of 3D datasets.[37] Neuroglancer, a web-based tool designed for visualizing and navigating large-scale neuroscience data, offers features like 3D rendering and interactive exploration of brain datasets.
Examples
To see one of the first micro-connectomes at full-resolution, visit the Open Connectome Project, which is hosting several connectome datasets, including the 12TB dataset from Bock et al. (2011).
Comparative connectomics
Comparative connectomics is a subfield in neuroscience that focuses on comparing the connectomes, or neural network maps, across different species, developmental stages, or pathological states.[38] This comparative approach aims to uncover fundamental principles of brain organization and function by identifying conserved and divergent patterns in neural circuitry. By analyzing similarities and differences in the wiring diagrams of various organisms, researchers can gain insights into the evolutionary processes shaping the nervous system, as well as into the neural basis of behavior and cognition.
For example, a 2022 study comparing synaptic connectivity in the mouse and human/macaque cortex revealed that, even though the human cortex contains three times more interneurons than the mouse cortex, the excitation-to-inhibition ratio is similar between the species.[29] Another study explicitly examined the differences between the male and female nervous systems of Drosophila melanogaster.[39]
Plasticity of the connectome
At the beginning of the connectome project, it was thought that the connections between neurons were unchangeable once established and that only individual synapses could be altered.[40] However, recent evidence suggests that connectivity is also subject to change, termed neuroplasticity. There are two ways that the brain can rewire: formation and removal of synapses in an established connection or formation or removal of entire connections between neurons.[41] Both mechanisms of rewiring are useful for learning completely novel tasks that may require entirely new connections between regions of the brain.[42] However, the ability of the brain to gain or lose entire connections poses an issue for mapping a universal species connectome. Although rewiring happens on different scales, from microscale to macroscale, each scale does not occur in isolation. For example, in the C. elegans connectome, the total number of synapses increases 5-fold from birth to adulthood, changing both local and global network properties.[43] Other developmental connectomes, such as the muscle connectome, retain some global network properties even though the number of synapses decreases by 10-fold in early postnatal life.[44]
Macroscale rewiring
Evidence for macroscale rewiring mostly comes from research on grey and white matter density, which could indicate new connections or changes in axon density. Direct evidence for this level of rewiring comes from primate studies, using viral tracing to map the formation of connections. Primates that were taught to use novel tools developed new connections between the interparietal cortex and higher visual areas of the brain.[45] Further viral tracing studies have provided evidence that macroscale rewiring occurs in adult animals during associative learning.[46] However, it is not likely that long-distance neural connections undergo extensive rewiring in adults. Small changes in an already established nerve tract are likely what is observed in macroscale rewiring.
Mesoscale rewiring
Rewiring at the mesoscale involves studying the presence or absence of entire connections between neurons.[42] Evidence for this level of rewiring comes from observations that local circuits form new connections as a result of experience-dependent plasticity in the visual cortex. Additionally, the number of local connections between pyramidal neurons in the primary somatosensory cortex increases following altered whisker sensory experience in rodents.[47]
Microscale rewiring
Microscale rewiring is the formation or removal of synaptic connections between two neurons and can be studied with longitudinal two-photon imaging. Dendritic spines on pyramidal neurons can be shown forming within days following sensory experience and learning.[48][49][50] Changes can even be seen within five hours on apical tufts of layer five pyramidal neurons in the primary motor cortex after a seed reaching task in primates.[50]
Model systems
For macroscale connectomes, the most common subject is the human. For microscale connectomes, some of the model systems are the mouse,[51] the fruit fly,[52][53] the nematode C. elegans,[54][55] and the barn owl.[56] C. elegans and the fruit fly have the advantage that their nervous systems are small enough to be reconstructed in their entirety, plus the organisms are well studied in other domains, such as genetics and behavior. The mouse, the human, and the owl are vertebrates and mammals, with larger and more capable nervous systems, but their brains are much too large to reconstruct completely as of 2025[update].
A scientific advantage of an optically transparent organism such as Danionella or larval zebrafish is that they allow neural activity recordings (typically calcium imaging)[57] to be performed in the same animal that will later be used for neural circuit reconstruction. This allows researchers to bypass many of the animal-to-animal variability problems caused when trying to correlate the behavior observed in one animal with the reconstructed circuitry of a different animal.
Humans
The Human Connectome Project (HCP) was an initiative launched in 2009 by the National Institutes of Health (NIH) to map the macroscale neural pathways that underlie human brain function.[58] Additional programs within the Connectome Initiative, such as the Lifespan Connectome and Disease Connectome, focus on mapping brain connections across different age groups and studying connectome variations in individuals with specific clinical diagnoses.[58] The Connectome Coordination Facility serves as a centralized repository for HCP data and provides support to researchers.[58]
Caenorhabditis Elegans
The C. elegans roundworm has a simple nervous system of 302 neurons and 5000 synaptic connections, (as compared to the human brain which has 100 billion neurons and more than 100 trillion chemical synapses).[59] It was the first of the very few animals in which a full connectome has been mapped using various imaging techniques, mainly serial-electron microscopy.[60] This has made it a natural target for connectomics.
One project studied the aging process of the C. elegans brain by comparing varying worms from birth to adulthood.[61] Researchers found the biggest change with age is the wiring circuits, and that connectivity between and within brain regions increases with age.[61] Additional findings are likely through comparative connectomics, comparing and contrasting different species' brain networks to pinpoint relations in behavior.[61]
Another study analyzed connections about sensory neurons, interneurons, neck motor neurons, behavior, environmental influences, and more in detail.[62]
Fruit Fly
Within the last decade, largely owing to technological advancements in EM data collection and image processing, multiple synapse-scale connectome datasets have been generated for the fruit fly Drosophila melanogaster in its adult and larval forms. The full fly connectome contains on the order of 150 thousand neurons and 300 million synapses.
The largest current dataset is central nervous system (CNS) of the male, consisting of both the brain and ventral nerve cord.[39] Next in size is the CNS of the female,[63] though not as completely reconstructed. Another large dataset, but not including the nerve cord, is the FlyWire segmentation and annotation of the female adult fly brain (FAFB) volume,[22][23] which encompasses the entire brain of an adult. Another adult brain dataset available is the Hemibrain, generated as a collaboration between the Janelia FlyEM team and Google.[64][65] This dataset is an incomplete but large section of the fly central brain. There are also currently two publicly available datasets of the adult fly ventral nerve cord (VNC). The female adult nerve cord (FANC) was collected using high-throughput SEM by Dr. Wei-Chung Allen Lee's lab at Harvard Medical School.[3] The male adult nerve cord (MANC) was collected at Janelia using FIB-SEM.[25] The connectome of a complete central nervous system (connected brain and VNC) of a 1st instar D. melanogaster larva has been collected as a single volume. This dataset of 3016 neurons was segmented and annotated manually using CATMAID by a team of people mainly led by researchers at Janelia, Cambridge, and the MRC LMB.[26]
Mouse
An online database known as MouseLight displays over 1000 neurons mapped in the mouse brain based on a collective database of sub-micron resolution images of these brains. This platform illustrates the thalamus, hippocampus, cerebral cortex, and hypothalamus based on single-cell projections.[66] Imaging technology to produce this mouse brain does not allow an in-depth look at synapses but can show axonal arborizations which contain many synapses.[67] A limiting factor to studying mouse connectomes, much like with humans, is the complexity of labeling all the cell types of the mouse brain; This is a process that would require the reconstruction of 100,000+ neurons and the imaging technology is advanced enough to do so.[67]
Mice models in the lab have provided insight into genetic brain disorders, one study manipulated mice with a deletion of 22q11.2 (chromosome 22, a likely known genetic risk factor that leads to schizophrenia).[68] The findings of this study showed that this impaired neural activity in mice's working memory is similar to what it does in humans.[68]
Applications
Macroscale and microscale connectomics have very different applications. Macroscale connectomics has furthered our understanding of various brain networks including visual,[69][70] brainstem,[71][72] and language networks,[73][74] among others. Microscale connectomics, on the other hand, concentrates on mechanistic explanations of how the neural circuits of the brain perform specific functions. Examples include motion vision,[75] olfactory learning,[76] navigation,[77] and escape responses,[78] all in Drosophila.
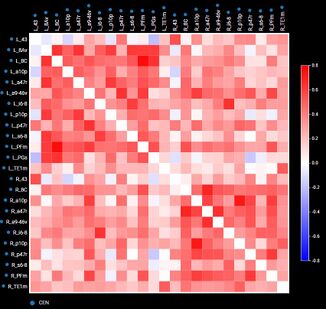
By comparing diseased and healthy connectomes, we can gain insight into certain psychopathologies, such as neuropathic pain, and potential therapies for them. Generally, the field of neuroscience would benefit from standardization and raw data. For example, connectome maps can be used to inform computational models of whole-brain dynamics.[79][self-published source?] Current neural networks mostly rely on probabilistic representations of connectivity patterns.[80] Connectivity matrices (checkerboard diagrams of connectomics) have been used in stroke recovery to evaluate the response to treatment via Transcranial Magnetic Stimulation.[81] Similarly, connectograms (circular diagrams of connectomics) have been used in traumatic brain injury cases to document the extent of damage to neural networks.[82][83]
Looking into these methods of research, they can reveal information about different mental illnesses and brain disorders. The tracking of brain networks in alignment with diseases and illnesses would be enhanced by these advanced technologies that can produce complex images of neural networks.[84] With this in mind, diseases can not only be tracked, but predicted based on behavior of previous cases, a process that would take an extensive period of time to collect and record.[84] Specifically, studies on different brain disorders such as schizophrenia and bipolar disorder with a focus on the connectomics involved reveal information. Both of these disorders have a similar genetic origin,[84][85] and research found that those with higher polygenic scores for schizophrenia and bipolar disorder have lower amounts of connectivity shown through neuroimaging.[86] This method of research tackles real-world applications of connectomics, combining methods of imaging with genetics to dig deeper into the origins and outcomes of genetically related disorders.[84] Another study supports the finding that there is relation between connectivity and likelihood of disease, as researchers found those diagnosed with schizophrenia have less structurally complete brain networks.[87] The main drawback in this area of connectomics is not being able to achieve images of whole-brain networks, therefore it is hard to make complete and accurate assumptions about cause and effect of diseases' neural pathways.[87] Connectomics has been used to study patients with strokes using MRI imaging, however because such little research is done in this specific area, conclusions cannot be drawn regarding the relation between strokes and connectivity.[88] The research did find results that highlight an association between poor connectivity in the language system and poor motor coordination, however the results were not substantial enough to make a bold claim.[88] For behavioral disorders, it can be difficult to diagnose and treat because most situations revolve on a symptoms-based approach. However, this can be difficult because many disorders have overlapping symptoms. Connectomics has been used to find neuromarkers associated with social anxiety disorder (SAD) at a high precision rate in improving related symptoms.[89] This is an expanding field and there is room for greater application to mental health disorders and brain malfunction, in which current research is building on neural networks and the psychopathology involved.[90]
Human connectomes have an individual variability, which can be measured with the cumulative distribution function, as it was shown in.[91] By analyzing the individual variability of the human connectomes in distinct cerebral areas, it was found that the frontal and the limbic lobes are more conservative, and the edges in the temporal and occipital lobes are more diverse. A "hybrid" conservative/diverse distribution was detected in the paracentral lobule and the fusiform gyrus. Smaller cortical areas were also evaluated: precentral gyri were found to be more conservative, and the postcentral and the superior temporal gyri to be very diverse.
Comparison to genomics
Both connectomics and genomics involve hugely detailed datasets of complex biological systems, derived by large and expensive research efforts. Genomics, the earlier of the two, focuses on the genetic blueprint of an organism while connectomics reveals the structural and functional connectivity of the brain. Comparisons of the two efforts often concentrate on what connectomics can learn from the more mature field of genomics.[92]
By using results from both connectomics and genomics, researchers can explore how genetic variations and gene expression patterns influence the wiring and organization of neural circuits.[93] This interdisciplinary approach helps uncover the relationship between genes, neural connectivity, and brain function. Understanding the genetic basis of neural connectivity can enhance our understanding of brain development, neural plasticity, and the mechanisms underlying various neurological disorders. Additionally, biologists can augment their analysis of circuits found by connectomics by using genetic tools and techniques to manipulate specific genes or neuronal populations to study their impact on neural circuitry and behavior.[92]
The Human Genome Project initially faced many of the same initial accusations of expense and impracticality leveled at connectomics. In the end, however, it was completed ahead of schedule and has led to many advances in genetics. Given the analogies between the two efforts, some have argued that we should be at least slightly more optimistic about the prospects in connectomics.[94] Others, however, have criticized attempts towards a microscale connectome, arguing that we do not have enough knowledge about where to look for insights, or that it cannot be completed within a realistic time frame.[95]
Mapping functional connectivity
Using fMRI in the resting state and during tasks, functions of the connectome circuits are being studied.[96] Just as detailed road maps of the Earth's surface do not tell us much about the kind of vehicles that travel those roads or what cargo they are hauling, to understand how neural structures result in specific functional behavior such as consciousness, it is necessary to build theories that relate functions to anatomical connectivity.[97] However, the bond between structural and functional connectivity is not straightforward. Computational models of whole-brain network dynamics are valuable tools to investigate the role of the anatomical network in shaping functional connectivity.[98][99] In particular, computational models can be used to predict the dynamic effect of lesions in the connectome.[100][101]
As a network or graph
A connectome can be viewed as a graph, and the rich tools, definitions and algorithms of graph theory and network science can be applied to these graphs. In case of a micro-scale connectome, the nodes of this network (or graph) are the neurons, and the edges correspond to the synapses between those neurons. For the macro-scale connectome, the nodes correspond to the ROIs (regions of interest), while the edges of the graph are derived from the axons interconnecting those areas. Thus connectomes are sometimes referred to as brain graphs, as they are indeed graphs in a mathematical sense which describe the connections in the brain (or, in a broader sense, the whole nervous system).
One group of researchers (Iturria-Medina et al., 2008)[102] has constructed connectome data sets using diffusion tensor imaging (DTI)[103][104] followed by the derivation of average connection probabilities between 70 and 90 cortical and basal brain gray matter areas. All networks were found to have small-world attributes and "broad-scale" degree distributions. An analysis of betweenness centrality in these networks demonstrated high centrality for the precuneus, the insula, the superior parietal and the superior frontal cortex. Another group (Gong et al. 2008)[105] has applied DTI to map a network of anatomical connections between 78 cortical regions. This study also identified several hub regions in the human brain, including the precuneus and the superior frontal gyrus.
Hagmann et al. (2007)[106] constructed a connection matrix from fiber densities measured between homogeneously distributed and equal-sized ROIs numbering between 500 and 4000. A quantitative analysis of connection matrices obtained for approximately 1,000 ROIs and approximately 50,000 fiber pathways from two subjects demonstrated an exponential (one-scale) degree distribution as well as robust small-world attributes for the network. The data sets were derived from diffusion spectrum imaging (DSI) (Wedeen, 2005),[107] a variant of diffusion-weighted imaging[108][109] that is sensitive to intra-voxel heterogeneities in diffusion directions caused by crossing fiber tracts and thus allows more accurate mapping of axonal trajectories than other diffusion imaging approaches (Wedeen, 2008).[110] The combination of whole-head DSI datasets acquired and processed according to the approach developed by Hagmann et al. (2007)[106] with the graph analysis tools conceived initially for animal tracing studies (Sporns, 2006; Sporns, 2007)[111][112] allow a detailed study of the network structure of human cortical connectivity (Hagmann et al., 2008).[113] The human brain network was characterized using a broad array of network analysis methods including core decomposition, modularity analysis, hub classification and centrality. Hagmann et al. presented evidence for the existence of a structural core of highly and mutually interconnected brain regions, located primarily in posterior medial and parietal cortex. The core comprises portions of the posterior cingulate cortex, the precuneus, the cuneus, the paracentral lobule, the isthmus of the cingulate, the banks of the superior temporal sulcus, and the inferior and superior parietal cortex, all located in both cerebral hemispheres.
A subfield of connectomics deals with the comparison of the brain graphs of multiple subjects. It is possible to build a consensus graph such the Budapest Reference Connectome by allowing only edges that are present in at least connectomes, for a selectable parameter. The Budapest Reference Connectome has led the researchers to the discovery of the Consensus Connectome Dynamics of the human brain graphs. The edges appeared in all of the brain graphs form a connected subgraph around the brainstem. By allowing gradually less frequent edges, this core subgraph grows continuously, as a shrub. The growth dynamics may reflect the individual brain development and provide an opportunity to direct some edges of the human consensus brain graph.[114]
Alternatively, local difference which are statistically significantly different among groups have attracted more attention as they highlight specific connections and therefore shed more light on specific brain traits or pathology. Hence, algorithms to find local difference between graph populations have also been introduced (e.g. to compare case versus control groups).[115] Those can be found by using either an adjusted t-test,[116] or a sparsity model,[115] with the aim of finding statistically significant connections which are different among those groups.
Comparisons between the connectomes (or braingraphs) of healthy women and men[117][118][119] have shown that in several deep graph-theoretical parameters, the structural connectome of women is significantly better connected than that of men. For example, women's connectome has more edges, higher minimum bipartition width, larger eigengap, greater minimum vertex cover than that of men. The minimum bipartition width (or, in other words, the minimum balanced cut) is a well-known measure of quality of computer multistage interconnection networks, it describes the possible bottlenecks in network communication: The higher this value is, the better is the network. The larger eigengap shows that the female connectome is better expander graph than the connectome of males. The better expanding property, the higher minimum bipartition width and the greater minimum vertex cover show deep advantages in network connectivity in the case of female braingraph.
Connectomes generally exhibit a small-world character, with overall cortical connectivity decreasing with age.[120] The aim of the as of 2015 ongoing HCP Lifespan Pilot Project is to identify connectome differences between 6 age groups (4–6, 8–9, 14–15, 25–35, 45–55, 65–75).
More recently, connectograms have been used to visualize full-brain data by placing cortical areas around a circle, organized by lobe.[121][122] Inner circles then depict cortical metrics on a color scale. White matter fiber connections in DTI data are then drawn between these cortical regions and weighted by fractional anisotropy and strength of the connection. Such graphs have even been used to analyze the damage done to the famous traumatic brain injury patient Phineas Gage.[123]
Statistical graph theory is an emerging discipline which is developing sophisticated pattern recognition and inference tools to parse these brain graphs (Goldenberg et al., 2009).
Origin and usage of the term
In 2005, Dr. Olaf Sporns at Indiana University and Dr. Patric Hagmann at Lausanne University Hospital independently and simultaneously suggested the term "connectome" to refer to a map of the neural connections within the brain. This term was directly inspired by the ongoing effort to sequence the human genetic code—to build a genome.
"Connectomics" has been defined as the science concerned with assembling and analyzing connectome data sets.[124]
In their 2005 paper, "The Human Connectome, a structural description of the human brain", Sporns et al. wrote:
To understand the functioning of a network, one must know its elements and their interconnections. The purpose of this article is to discuss research strategies aimed at a comprehensive structural description of the network of elements and connections forming the human brain. We propose to call this dataset the human "connectome," and we argue that it is fundamentally important in cognitive neuroscience and neuropsychology. The connectome will significantly increase our understanding of how functional brain states emerge from their underlying structural substrate, and will provide new mechanistic insights into how brain function is affected if this structural substrate is disrupted.[40]
In his 2005 Ph.D. thesis, From diffusion MRI to brain connectomics, Hagmann wrote:
It is clear that, like the genome, which is much more than just a juxtaposition of genes, the set of all neuronal connections in the brain is much more than the sum of their individual components. The genome is an entity it-self, as it is from the subtle gene interaction that [life] emerges. In a similar manner, one could consider the brain connectome, set of all neuronal connections, as one single entity, thus emphasizing the fact that the huge brain neuronal communication capacity and computational power critically relies on this subtle and incredibly complex connectivity architecture.[124]
The term "connectome" was more recently popularized by Sebastian Seung's I am my Connectome speech given at the 2010 TED conference, which discusses the high-level goals of mapping the human connectome, as well as ongoing efforts to build a three-dimensional neural map of brain tissue at the microscale.[125] In 2012, Seung published the book Connectome: How the Brain's Wiring Makes Us Who We Are.
Public datasets
Websites to explore publicly available connectomics datasets:
Macroscale Connectomics (Healthy Young Adult Datasets)
- Human Connectome Project Young Adult
- Amsterdam Open MRI Collection
- Harvard Brain Genomic Superstruct Project
For a more comprehensive list of open macroscale datasets, check out this article
Microscale Connectomics
- Whole C. elegans connectome
- Whole Platynereis dumerilii larva connectome
- NeuPRINT Fly Hemibrain, Male VNC, Male Complete CNS
- Flywire (whole fly brain)
- MICrONS Explorer (mouse cortical data)
- H01 Browser Release (human cortical data)
- Connectomic comparison of mouse and human cortex (mouse, macaque, and human cortical data)
See also
- Dynamic Functional Connectivity
- List of Functional Connectivity Software
- Human Connectome Project
- Budapest Reference Connectome
- Drosophila connectome
- History of microscale connectomics
- https://eyewire.org/explore
References
- ↑ "New insights into cortico-basal-cerebellar connectome: clinical and physiological considerations". Brain 143 (2): 396–406. February 2020. doi:10.1093/brain/awz310. PMID 31628799.
- ↑ Nguyen, Tri M.; Thomas, Logan A.; Rhoades, Jeff L.; Ricchi, Ilaria; Yuan, Xintong Cindy; Sheridan, Arlo; Hildebrand, David G. C.; Funke, Jan et al. (2023-01-19). "Structured cerebellar connectivity supports resilient pattern separation". Nature 613 (7944): 543–549. doi:10.1038/s41586-022-05471-w. ISSN 0028-0836. PMID 36418404. Bibcode: 2023Natur.613..543N.
- ↑ 3.0 3.1 "Reconstruction of motor control circuits in adult Drosophila using automated transmission electron microscopy". Cell 184 (3): 759–774.e18. February 2021. doi:10.1016/j.cell.2020.12.013. PMID 33400916.
- ↑ "Connectomic reconstruction of the inner plexiform layer in the mouse retina". Nature 500 (7461): 168–174. August 2013. doi:10.1038/nature12346. PMID 23925239. Bibcode: 2013Natur.500..168H.
- ↑ 5.0 5.1 5.2 5.3 Sotiropoulous, Statmotios; Zalesky, Andrew (June 27, 2017). "Building connectomes using diffusion MRI: why, how and but". NMR in Biomedicine 32 (4). doi:10.1002/nbm.3752. PMID 28654718.
- ↑ "Strong long-range connections can delocalize a subset of eigenvectors.". eLife. doi:10.7554/elife.01239.010. Figure 7.
- ↑ 7.0 7.1 Deligianni, Fani; Centeno, Maria; Carmichael, David W.; Clayden, Jonathan D. (2014). "Relating resting-state fMRI and EEG whole-brain connectomes across frequency bands". Frontiers in Neuroscience 8: 258. doi:10.3389/fnins.2014.00258. ISSN 1662-453X. PMID 25221467.
- ↑ Soyuhos, Orhan; Scarpa, Aurelia; Baldauf, Daniel (2025-07-15). "Distinct resting-state connectomes for face and scene perception predict individual task performance". bioRxiv 10.1101/2025.07.09.663812.
- ↑ "Greater than the sum of its parts: a review of studies combining structural connectivity and resting-state functional connectivity". Brain Structure & Function 213 (6): 525–533. October 2009. doi:10.1007/s00429-009-0208-6. PMID 19565262.
- ↑ "Predicting human resting-state functional connectivity from structural connectivity". Proceedings of the National Academy of Sciences of the United States of America 106 (6): 2035–2040. February 2009. doi:10.1073/pnas.0811168106. PMID 19188601. Bibcode: 2009PNAS..106.2035H.
- ↑ Zhu, Feiyin; Tang, Liying (July 5, 2019). "Resting-state functional magnetic resonance imaging (fMRI) and functional connectivity density mapping in patients with corneal ulcer". Neuropsychiatric Disease and Treatment 15: 1833–1844. doi:10.2147/NDT.S210658. PMID 31308676.
- ↑ Horn, Andreas; Fox, Michael D. (2020-11-01). "Opportunities of connectomic neuromodulation" (in en). NeuroImage 221. doi:10.1016/j.neuroimage.2020.117180. ISSN 1053-8119. PMID 32702488.
- ↑ 13.0 13.1 Xia, Mingrui; He, Yong (2022-10-01). "Connectome-guided transcranial magnetic stimulation treatment in depression" (in en). European Child & Adolescent Psychiatry 31 (10): 1481–1483. doi:10.1007/s00787-022-02089-1. ISSN 1435-165X. PMID 36151354.
- ↑ 14.0 14.1 14.2 Kunze, Tim; Hunold, Alexander; Haueisen, Jens; Jirsa, Viktor; Spiegler, Andreas (2016-10-15). "Transcranial direct current stimulation changes resting state functional connectivity: A large-scale brain network modeling study" (in en). NeuroImage. Transcranial electric stimulation (tES) and Neuroimaging 140: 174–187. doi:10.1016/j.neuroimage.2016.02.015. ISSN 1053-8119. PMID 26883068.
- ↑ 15.0 15.1 Wang, Qiang; Akram, Harith; Muthuraman, Muthuraman; Gonzalez-Escamilla, Gabriel; Sheth, Sameer A.; Oxenford, Simón; Yeh, Fang-Cheng; Groppa, Sergiu et al. (2021-01-01). "Normative vs. patient-specific brain connectivity in deep brain stimulation" (in en). NeuroImage 224. doi:10.1016/j.neuroimage.2020.117307. ISSN 1053-8119. PMID 32861787.
- ↑ Benabid, Alim Louis (2003-12-01). "Deep brain stimulation for Parkinson's disease" (in en). Current Opinion in Neurobiology 13 (6): 696–706. doi:10.1016/j.conb.2003.11.001. ISSN 0959-4388. PMID 14662371.
- ↑ Sadaghiani, Sepideh; Brookes, Matthew J; Baillet, Sylvain (2022-02-15). "Connectomics of human electrophysiology" (in en). NeuroImage 247. doi:10.1016/j.neuroimage.2021.118788. ISSN 1053-8119. PMID 34906715.
- ↑ 18.0 18.1 Sadaghiani, Sepideh; Brookes, Matthew J.; Baillet, Sylvain (2022-02-15). "Connectomics of human electrophysiology" (in en). NeuroImage 247. doi:10.1016/j.neuroimage.2021.118788. ISSN 1053-8119. PMID 34906715.
- ↑ "Exploring the retinal connectome". Molecular Vision 17: 355–379. February 2011. PMID 21311605.
- ↑ 20.0 20.1 White, J. G.; Southgate, E.; Thomson, J. N.; Brenner, S. (1986-11-12). "The structure of the nervous system of the nematode Caenorhabditis elegans" (in en). Philosophical Transactions of the Royal Society of London. B, Biological Sciences 314 (1165): 1–340. doi:10.1098/rstb.1986.0056. ISSN 0080-4622. PMID 22462104. Bibcode: 1986RSPTB.314....1W. https://royalsocietypublishing.org/doi/10.1098/rstb.1986.0056.
- ↑ Verasztó, Csaba; Jasek, Sanja; Gühmann, Martin; Bezares-Calderón, Luis Alberto; Williams, Elizabeth A.; Shahidi, Réza; Jékely, Gáspár (3 March 2025). "Whole-body connectome of a segmented annelid larva". eLife 13. doi:10.7554/eLife.97964.2. PMID 40862480.
- ↑ 22.0 22.1 Dorkenwald, Sven et al. (2024-10-02). "Neuronal wiring diagram of an adult brain". Nature 634 (8032): 124–138. doi:10.1038/s41586-024-07558-y. ISSN 1476-4687. PMID 39358518.
- ↑ 23.0 23.1 Schlegel, Philipp et al. (2024-10-02). "Whole-brain annotation and multi-connectome cell typing of Drosophila". Nature 634 (8032): 139–152. doi:10.1038/s41586-024-07686-5. ISSN 1476-4687. PMID 39358521.
- ↑ Phelps, Jasper S.; Hildebrand, David Grant Colburn; Graham, Brett J.; Kuan, Aaron T.; Thomas, Logan A.; Nguyen, Tri M.; Buhmann, Julia; Azevedo, Anthony W. et al. (February 2021). "Reconstruction of motor control circuits in adult Drosophila using automated transmission electron microscopy". Cell 184 (3): 759–774.e18. doi:10.1016/j.cell.2020.12.013. ISSN 0092-8674. PMID 33400916.
- ↑ 25.0 25.1 Takemura, Shin-ya; Hayworth, Kenneth J; Huang, Gary B; Januszewski, Michal; Lu, Zhiyuan; Marin, Elizabeth C; Preibisch, Stephan; Xu, C Shan; Bogovic, John (2023-06-06). "A Connectome of the Male Drosophila Ventral Nerve Cord". bioRxiv 10.1101/2023.06.05.543757.
- ↑ 26.0 26.1 Winding, Michael; Pedigo, Benjamin D.; Barnes, Christopher L.; Patsolic, Heather G.; Park, Youngser; Kazimiers, Tom; Fushiki, Akira; Andrade, Ingrid V. et al. (2023-03-10). "The connectome of an insect brain" (in en). Science 379 (6636). doi:10.1126/science.add9330. ISSN 0036-8075. PMID 36893230. Bibcode: 2023Sci...379d9330W.
- ↑ The MICrONS Consortium; Bae, J. Alexander; Baptiste, Mahaly; Bishop, Caitlyn A.; Bodor, Agnes L.; Brittain, Derrick; Buchanan, JoAnn; Bumbarger, Daniel J. et al. (2025). "Functional connectomics spanning multiple areas of mouse visual cortex" (in en). Nature 640 (8058): 435–447. doi:10.1038/s41586-025-08790-w. PMID 40205214.
- ↑ Shapson-Coe, Alexander; Januszewski, Michał; Berger, Daniel R.; Pope, Art; Wu, Yuelong; Blakely, Tim; Schalek, Richard L.; Li, Peter H.; Wang, Shuohong (2021-05-30). "A connectomic study of a petascale fragment of human cerebral cortex". bioRxiv 10.1101/2021.05.29.446289.
- ↑ 29.0 29.1 Loomba, Sahil; Straehle, Jakob; Gangadharan, Vijayan; Heike, Natalie; Khalifa, Abdelrahman; Motta, Alessandro; Ju, Niansheng; Sievers, Meike et al. (2022-07-08). "Connectomic comparison of mouse and human cortex" (in en). Science 377 (6602). doi:10.1126/science.abo0924. ISSN 0036-8075. PMID 35737810.
- ↑ "BRAIN CONNECTS: A Center for High-throughput Integrative Mouse Connectomics". https://reporter.nih.gov/search/OZaPZ82vPUynldKK6St1Pg/project-details/10665380.
- ↑ BV, DELMIC. "Neuroscience: Synaptic connectivity in the songbird brain - Application Note | DELMIC" (in en). http://request.delmic.com/neuroscience.
- ↑ 32.0 32.1 Kuan, Aaron T.; Phelps, Jasper S.; Thomas, Logan A.; Nguyen, Tri M.; Han, Julie; Chen, Chiao-Lin; Azevedo, Anthony W.; Tuthill, John C. et al. (December 2020). "Dense neuronal reconstruction through X-ray holographic nano-tomography" (in en). Nature Neuroscience 23 (12): 1637–1643. doi:10.1038/s41593-020-0704-9. ISSN 1097-6256. PMID 32929244.
- ↑ 33.0 33.1 33.2 Velicky, Philipp; Miguel, Eder; Michalska, Julia M.; Lyudchik, Julia; Wei, Donglai; Lin, Zudi; Watson, Jake F.; Troidl, Jakob et al. (August 2023). "Dense 4D nanoscale reconstruction of living brain tissue" (in en). Nature Methods 20 (8): 1256–1265. doi:10.1038/s41592-023-01936-6. ISSN 1548-7091. PMID 37429995.
- ↑ Tavakoli, Mojtaba R.; Lyudchik, Julia; Januszewski, Michał; Vistunou, Vitali; Agudelo, Nathalie; Vorlaufer, Jakob; Sommer, Christoph; Kreuzinger, Caroline et al. (2025). "Light-microscopy-based connectomic reconstruction of mammalian brain tissue". Nature 642 (8067): 398–410. doi:10.1038/s41586-025-08985-1. PMID 40335689. Bibcode: 2025Natur.642..398T.
- ↑ Motta, Alessandro; Berning, Manuel; Boergens, Kevin M.; Staffler, Benedikt; Beining, Marcel; Loomba, Sahil; Hennig, Philipp; Wissler, Heiko et al. (2019-11-29). "Dense connectomic reconstruction in layer 4 of the somatosensory cortex" (in en). Science 366 (6469). doi:10.1126/science.aay3134. ISSN 0036-8075. PMID 31649140. https://www.science.org/doi/10.1126/science.aay3134.
- ↑ Saalfeld, Stephan; Cardona, Albert; Hartenstein, Volker; Tomančák, Pavel (2009-04-17). "CATMAID: collaborative annotation toolkit for massive amounts of image data". Bioinformatics 25 (15): 1984–1986. doi:10.1093/bioinformatics/btp266. ISSN 1367-4811. PMID 19376822.
- ↑ Boergens, Kevin M.; Berning, Manuel; Bocklisch, Tom; Bräunlein, Dominic; Drawitsch, Florian; Frohnhofen, Johannes; Herold, Tom; Otto, Philipp et al. (July 2017). "webKnossos: efficient online 3D data annotation for connectomics" (in en). Nature Methods 14 (7): 691–694. doi:10.1038/nmeth.4331. ISSN 1548-7105. PMID 28604722. https://www.nature.com/articles/nmeth.4331.
- ↑ van den Heuvel, Martijn P.; Bullmore, Edward T.; Sporns, Olaf (May 2016). "Comparative Connectomics". Trends in Cognitive Sciences 20 (5): 345–361. doi:10.1016/j.tics.2016.03.001. ISSN 1364-6613. PMID 27026480. https://www.repository.cam.ac.uk/handle/1810/254415.
- ↑ 39.0 39.1 Berg, Stuart; et al. (2025). "Sexual dimorphism in the complete connectome of the Drosophila male central nervous system". pp. 2025–10. bioRxiv 10.1101/2025.10.09.680999.
- ↑ 40.0 40.1 "The human connectome: A structural description of the human brain". PLOS Computational Biology 1 (4). September 2005. doi:10.1371/journal.pcbi.0010042. PMID 16201007. Bibcode: 2005PLSCB...1...42S.

- ↑ Greenough, William T.; Bailey, Craig H. (January 1988). "The anatomy of a memory: convergence of results across a diversity of tests". Trends in Neurosciences 11 (4): 142–147. doi:10.1016/0166-2236(88)90139-7.
- ↑ 42.0 42.1 "Rewiring the connectome: Evidence and effects". Neuroscience and Biobehavioral Reviews 88: 51–62. May 2018. doi:10.1016/j.neubiorev.2018.03.001. PMID 29540321.
- ↑ Witvliet, Daniel; Mulcahy, Ben; Mitchell, James K.; Meirovitch, Yaron; Berger, Daniel R.; Wu, Yuelong; Liu, Yufang; Koh, Wan Xian et al. (August 2021). "Connectomes across development reveal principles of brain maturation" (in en). Nature 596 (7871): 257–261. doi:10.1038/s41586-021-03778-8. ISSN 1476-4687. PMID 34349261. Bibcode: 2021Natur.596..257W.
- ↑ Meirovitch, Yaron; Kang, Kai; Draft, Ryan W.; Pavarino, Elisa C.; Henao E., Maria F.; Yang, Fuming; Turney, Stephen G.; Berger, Daniel R.; Peleg, Adi; Schalek, Richard L.; Lu, Ju L.; Tapia, Juan-Carlos; Lichtman, Jeff W. (September 2021). "Neuromuscular connectomes across development reveal synaptic ordering rules". bioRxiv 10.1101/2021.09.20.460480.
- ↑ "Extension of corticocortical afferents into the anterior bank of the intraparietal sulcus by tool-use training in adult monkeys". Neuropsychologia 44 (13): 2636–46. 2006. doi:10.1016/j.neuropsychologia.2005.11.020. PMID 16427666.
- ↑ "Axonal sprouting and formation of terminals in the adult cerebellum during associative motor learning". The Journal of Neuroscience 33 (45): 17897–907. November 2013. doi:10.1523/JNEUROSCI.0511-13.2013. PMID 24198378.
- ↑ "The emergence of functional microcircuits in visual cortex". Nature 496 (7443): 96–100. April 2013. doi:10.1038/nature12015. PMID 23552948. Bibcode: 2013Natur.496...96K.
- ↑ "Experience-dependent and cell-type-specific spine growth in the neocortex" (in En). Nature 441 (7096): 979–83. June 2006. doi:10.1038/nature04783. PMID 16791195. Bibcode: 2006Natur.441..979H.
- ↑ "Spine growth precedes synapse formation in the adult neocortex in vivo" (in En). Nature Neuroscience 9 (9): 1117–24. September 2006. doi:10.1038/nn1747. PMID 16892056.
- ↑ 50.0 50.1 "Rapid formation and selective stabilization of synapses for enduring motor memories" (in En). Nature 462 (7275): 915–9. December 2009. doi:10.1038/nature08389. PMID 19946267. Bibcode: 2009Natur.462..915X.
- ↑ "Network anatomy and in vivo physiology of visual cortical neurons". Nature 471 (7337): 177–182. March 2011. doi:10.1038/nature09802. PMID 21390124. Bibcode: 2011Natur.471..177B.
- ↑ "Semi-automated reconstruction of neural circuits using electron microscopy". Current Opinion in Neurobiology 20 (5): 667–675. October 2010. doi:10.1016/j.conb.2010.08.002. PMID 20833533.
- ↑ "A Complete Electron Microscopy Volume of the Brain of Adult Drosophila melanogaster". Cell 174 (3): 730–743.e22. July 2018. doi:10.1016/j.cell.2018.06.019. PMID 30033368. Bibcode: 2018Cell..174..730Z.
- ↑ "Wiring optimization can relate neuronal structure and function". Proceedings of the National Academy of Sciences of the United States of America 103 (12): 4723–4728. March 2006. doi:10.1073/pnas.0506806103. PMID 16537428. Bibcode: 2006PNAS..103.4723C.
- ↑ "Structure of deviations from optimality in biological systems". Proceedings of the National Academy of Sciences of the United States of America 106 (48): 20544–20549. December 2009. doi:10.1073/pnas.0905336106. PMID 19918070. Bibcode: 2009PNAS..10620544P.
- ↑ "Auditory processing, plasticity, and learning in the barn owl". ILAR Journal 51 (4): 338–352. 2010. doi:10.1093/ilar.51.4.338. PMID 21131711.
- ↑ Fouke, Kaitlyn E.; He, Zichen; Loring, Matthew D.; Naumann, Eva A. (2025). "Neural circuits underlying divergent visuomotor strategies of zebrafish and Danionella cerebrum". Current Biology. doi:10.1016/j.cub.2025.04.027. PMID 40318635. https://doi.org/10.1016/j.cub.2025.04.027.
- ↑ 58.0 58.1 58.2 "Connectome Programs | Blueprint". https://www.ninds.nih.gov:443/current-research/trans-agency-activities/nih-blueprint-neuroscience-research.
- ↑ "Overview" (in en). https://medicine.yale.edu/lab/colon_ramos/overview/.
- ↑ "WormWiring". https://www.wormwiring.org/.
- ↑ 61.0 61.1 61.2 "Connectomes across development reveal principles of brain maturation". Nature 596 (7871): 257–261. August 2021. doi:10.1038/s41586-021-03778-8. PMID 34349261. Bibcode: 2021Natur.596..257W.
- ↑ "Connecting a connectome to behavior: an ensemble of neuroanatomical models of C. elegans klinotaxis". PLOS Computational Biology 9 (2). 2013-02-07. doi:10.1371/journal.pcbi.1002890. PMID 23408877. Bibcode: 2013PLSCB...9E2890I.
- ↑ Bates, Alexander Shakeel; Phelps, Jasper S.; Kim, Minsu; Yang, Helen H.; Matsliah, Arie; Ajabi, Zaki; Perlman, Eric; Delgado, Kevin M.; Osman, Mohammed Abdal Monium (2025-08-02). "Distributed control circuits across a brain-and-cord connectome". pp. 2025.07.31.667571. bioRxiv 10.1101/2025.07.31.667571.
- ↑ Scheffer, Louis K; Xu, C Shan; Januszewski, Michal; Lu, Zhiyuan; Takemura, Shin-ya; Hayworth, Kenneth J; Huang, Gary B; Shinomiya, Kazunori et al. (2020-09-03). Marder, Eve; Eisen, Michael B; Pipkin, Jason et al.. eds. "A connectome and analysis of the adult Drosophila central brain". eLife 9. doi:10.7554/eLife.57443. ISSN 2050-084X. PMID 32880371. Bibcode: 2020eLife...957443S.
- ↑ "Hemibrain" (in en). https://www.janelia.org/project-team/flyem/hemibrain.
- ↑ "MouseLight". http://www.mouselight.janelia.org/.
- ↑ 67.0 67.1 "Reconstruction of 1,000 Projection Neurons Reveals New Cell Types and Organization of Long-Range Connectivity in the Mouse Brain". Cell 179 (1): 268–281.e13. September 2019. doi:10.1016/j.cell.2019.07.042. PMID 31495573.
- ↑ 68.0 68.1 "Impaired hippocampal-prefrontal synchrony in a genetic mouse model of schizophrenia". Nature 464 (7289): 763–767. April 2010. doi:10.1038/nature08855. PMID 20360742. Bibcode: 2010Natur.464..763S.
- ↑ "Automated retinofugal visual pathway reconstruction with multi-shell HARDI and FOD-based analysis". NeuroImage 125: 767–779. January 2016. doi:10.1016/j.neuroimage.2015.11.005. PMID 26551261.
- ↑ "Defining Meyer's loop-temporal lobe resections, visual field deficits and diffusion tensor tractography". Brain 132 (Pt 6): 1656–1668. June 2009. doi:10.1093/brain/awp114. PMID 19460796.
- ↑ The Human Central Nervous System. 2008. doi:10.1007/978-3-540-34686-9. ISBN 978-3-540-34684-5.
- ↑ "Organization of Brainstem Nuclei". The Human Nervous System. Elsevier. 2012. pp. 260–327. doi:10.1016/b978-0-12-374236-0.10008-2. ISBN 978-0-12-374236-0. https://ro.uow.edu.au/cgi/viewcontent.cgi?article=4108&context=hbspapers.
- ↑ "DTI tractography of the human brain's language pathways". Cerebral Cortex 18 (11): 2471–2482. November 2008. doi:10.1093/cercor/bhn011. PMID 18281301.
- ↑ "Perisylvian language networks of the human brain". Annals of Neurology 57 (1): 8–16. January 2005. doi:10.1002/ana.20319. PMID 15597383.
- ↑ Borst, Alexander; Groschner, Lukas N (2023). "How flies see motion". Annual Review of Neuroscience (Annual Reviews) 46 (1): 17–37. doi:10.1146/annurev-neuro-080422-111929. PMID 37428604.
- ↑ Li, Feng; Lindsey, Jack W; Marin, Elizabeth C; Otto, Nils; Dreher, Marisa; Dempsey, Georgia; Stark, Ildiko; Bates, Alexander S et al. (2020). "The connectome of the adult Drosophila mushroom body provides insights into function". eLife (eLife Sciences Publications, Ltd) 9. doi:10.7554/eLife.62576. PMID 33315010. PMC 7909955. Bibcode: 2020eLife...962576L. https://elifesciences.org/articles/62576.pdf.
- ↑ Hulse, Brad K; Haberkern, Hannah; Franconville, Romain; Evans, Daniel Turner-; Takemura, Shin- ya; Wolff, Tanya; Noorman, Marcella; Dreher, Marisa et al. (2021). "A connectome of the Drosophila central complex reveals network motifs suitable for flexible navigation and context-dependent action selection". eLife (eLife Sciences Publications, Ltd) 10. doi:10.7554/eLife.66039. PMID 34696823.
- ↑ Ache, Jan M.; Polsky, Jason; Alghailani, Shada; Parekh, Ruchi; Breads, Patrick; Peek, Martin Y.; Bock, Davi D.; Reyn, Catherine R. von et al. (2019). "Neural basis for looming size and velocity encoding in the Drosophila giant fiber escape pathway". Current Biology (Elsevier) 29 (6): 1073–1081. doi:10.1016/j.cub.2019.01.079. PMID 30827912. Bibcode: 2019CBio...29E1073A.
- ↑ http://www.scholarpedia.org/article/Connectome[unreliable medical source?]
- ↑ Friston, Karl J., ed (August 2009). "Towards reproducible descriptions of neuronal network models". PLOS Computational Biology 5 (8). doi:10.1371/journal.pcbi.1000456. PMID 19662159. Bibcode: 2009PLSCB...5E0456N.
- ↑ "Changes in the Brain Connectome Following Repetitive Transcranial Magnetic Stimulation for Stroke Rehabilitation". Cureus 13 (10). October 2021. doi:10.7759/cureus.19105. PMID 34858752.
- ↑ "Mapping connectivity damage in the case of Phineas Gage". PLOS ONE 7 (5). 2012. doi:10.1371/journal.pone.0037454. PMID 22616011. Bibcode: 2012PLoSO...737454V.
- ↑ "Patient-tailored connectomics visualization for the assessment of white matter atrophy in traumatic brain injury". Frontiers in Neurology 3: 10. 6 February 2012. doi:10.3389/fneur.2012.00010. PMID 22363313.
- ↑ 84.0 84.1 84.2 84.3 Fornito, Alex; Zalesky, Andrew; Breakspear, Michael (March 2015). "The connectomics of brain disorders" (in en). Nature Reviews Neuroscience 16 (3): 159–172. doi:10.1038/nrn3901. ISSN 1471-0048. PMID 25697159. https://www.nature.com/articles/nrn3901.
- ↑ Stahl, Eli A.; Breen, Gerome; Forstner, Andreas J.; McQuillin, Andrew; Ripke, Stephan; Trubetskoy, Vassily; Mattheisen, Manuel; Wang, Yunpeng et al. (May 2019). "Genome-wide association study identifies 30 loci associated with bipolar disorder" (in en). Nature Genetics 51 (5): 793–803. doi:10.1038/s41588-019-0397-8. ISSN 1546-1718. PMID 31043756.
- ↑ Wei, Yongbin; De Lange, Siemon C.; Savage, Jeanne E.; Tissink, Elleke; Qi, Ting; Repple, Jonathan; Gruber, Marius; Kircher, Tilo et al. (2022-11-09). "Associated Genetics and Connectomic Circuitry in Schizophrenia and Bipolar Disorder" (in en). Biological Psychiatry 94 (2): 174–183. doi:10.1016/j.biopsych.2022.11.006. ISSN 0006-3223. PMID 36803976.
- ↑ 87.0 87.1 Heuvel, Martijn P. van den; Mandl, René C. W.; Stam, Cornelis J.; Kahn, René S.; Pol, Hilleke E. Hulshoff (2010-11-24). "Aberrant Frontal and Temporal Complex Network Structure in Schizophrenia: A Graph Theoretical Analysis" (in en). Journal of Neuroscience 30 (47): 15915–15926. doi:10.1523/JNEUROSCI.2874-10.2010. ISSN 0270-6474. PMID 21106830. PMC 6633761. https://www.jneurosci.org/content/30/47/15915.
- ↑ 88.0 88.1 Bian, Rong; Huo, Ming; Liu, Wan; Mansouri, Negar; Tanglay, Onur; Young, Isabella; Osipowicz, Karol; Hu, Xiaorong et al. (2023). "Connectomics underlying motor functional outcomes in the acute period following stroke". Frontiers in Aging Neuroscience 15. doi:10.3389/fnagi.2023.1131415. ISSN 1663-4365. PMID 36875697.
- ↑ Whitfield-Gabrieli, S.; Ghosh, S. S.; Nieto-Castanon, A.; Saygin, Z.; Doehrmann, O.; Chai, X. J.; Reynolds, G. O.; Hofmann, S. G. et al. (May 2016). "Brain connectomics predict response to treatment in social anxiety disorder" (in en). Molecular Psychiatry 21 (5): 680–685. doi:10.1038/mp.2015.109. ISSN 1476-5578. PMID 26260493.
- ↑ Menon, Vinod (2011-10-01). "Large-scale brain networks and psychopathology: a unifying triple network model" (in en). Trends in Cognitive Sciences 15 (10): 483–506. doi:10.1016/j.tics.2011.08.003. ISSN 1364-6613. PMID 21908230.
- ↑ "Comparative connectomics: Mapping the inter-individual variability of connections within the regions of the human brain". Neuroscience Letters 662 (1): 17–21. January 2018. doi:10.1016/j.neulet.2017.10.003. PMID 28988973.
- ↑ 92.0 92.1 "What connectomics can learn from genomics". PLOS Genetics 17 (7). July 2021. doi:10.1371/journal.pgen.1009692. PMID 34270560.
- ↑ "Where the genome meets the connectome: Understanding how genes shape human brain connectivity". NeuroImage 244. December 2021. doi:10.1016/j.neuroimage.2021.118570. PMID 34508898.
- ↑ "Ome sweet ome: what can the genome tell us about the connectome?". Current Opinion in Neurobiology 18 (3): 346–353. June 2008. doi:10.1016/j.conb.2008.08.010. PMID 18801435.
- ↑ Vance, Ashlee (27 December 2010). "Seeking the Connectome, a Mental Map, Slice by Slice". The New York Times. https://www.nytimes.com/2010/12/28/science/28brain.html.
- ↑ "Intrinsic functional connectivity as a tool for human connectomics: theory, properties, and optimization". Journal of Neurophysiology 103 (1): 297–321. January 2010. doi:10.1152/jn.00783.2009. PMID 19889849.
- ↑ "Consciousness, plasticity, and connectomics: the role of intersubjectivity in human cognition". Frontiers in Psychology 2: 20. 2011. doi:10.3389/fpsyg.2011.00020. PMID 21687435.
- ↑ "Exploring the network dynamics underlying brain activity during rest". Progress in Neurobiology 114: 102–31. March 2014. doi:10.1016/j.pneurobio.2013.12.005. PMID 24389385.
- ↑ "Network structure of cerebral cortex shapes functional connectivity on multiple time scales". Proceedings of the National Academy of Sciences of the United States of America 104 (24): 10240–5. June 2007. doi:10.1073/pnas.0701519104. PMID 17548818. Bibcode: 2007PNAS..10410240H.
- ↑ "Modeling the outcome of structural disconnection on resting-state functional connectivity". NeuroImage 62 (3): 1342–53. September 2012. doi:10.1016/j.neuroimage.2012.06.007. PMID 22705375.
- ↑ "Dynamical consequences of lesions in cortical networks". Human Brain Mapping 29 (7): 802–9. July 2008. doi:10.1002/hbm.20579. PMID 18438885.
- ↑ "Studying the human brain anatomical network via diffusion-weighted MRI and Graph Theory". NeuroImage 40 (3): 1064–76. April 2008. doi:10.1016/j.neuroimage.2007.10.060. PMID 18272400.
- ↑ "MR diffusion tensor spectroscopy and imaging". Biophysical Journal 66 (1): 259–67. January 1994. doi:10.1016/S0006-3495(94)80775-1. PMID 8130344. Bibcode: 1994BpJ....66..259B.
- ↑ "Estimation of the effective self-diffusion tensor from the NMR spin echo". Journal of Magnetic Resonance, Series B 103 (3): 247–54. March 1994. doi:10.1006/jmrb.1994.1037. PMID 8019776. Bibcode: 1994JMagR.103..247B. https://zenodo.org/record/1229906.
- ↑ "Mapping anatomical connectivity patterns of human cerebral cortex using in vivo diffusion tensor imaging tractography". Cerebral Cortex 19 (3): 524–36. March 2009. doi:10.1093/cercor/bhn102. PMID 18567609.
- ↑ 106.0 106.1 "Mapping human whole-brain structural networks with diffusion MRI". PLOS ONE 2 (7). July 2007. doi:10.1371/journal.pone.0000597. PMID 17611629. Bibcode: 2007PLoSO...2..597H.

- ↑ "Mapping complex tissue architecture with diffusion spectrum magnetic resonance imaging". Magnetic Resonance in Medicine 54 (6): 1377–86. December 2005. doi:10.1002/mrm.20642. PMID 16247738.
- ↑ "Imagerie de diffusion in vivo par résonance magnétique nucléaire" (in fr). Comptes Rendus de l'Académie des Sciences 93 (5): 27–34. 1985.
- ↑ "MR imaging of intravoxel incoherent motions: application to diffusion and perfusion in neurologic disorders". Radiology 161 (2): 401–7. November 1986. doi:10.1148/radiology.161.2.3763909. PMID 3763909.
- ↑ "Diffusion spectrum magnetic resonance imaging (DSI) tractography of crossing fibers". NeuroImage 41 (4): 1267–77. July 2008. doi:10.1016/j.neuroimage.2008.03.036. PMID 18495497. http://infoscience.epfl.ch/record/160877.
- ↑ "Small-world connectivity, motif composition, and complexity of fractal neuronal connections". Bio Systems 85 (1): 55–64. July 2006. doi:10.1016/j.biosystems.2006.02.008. PMID 16757100. Bibcode: 2006BiSys..85...55S.
- ↑ "Identification and classification of hubs in brain networks". PLOS ONE 2 (10). October 2007. doi:10.1371/journal.pone.0001049. PMID 17940613. Bibcode: 2007PLoSO...2.1049S.

- ↑ "Mapping the structural core of human cerebral cortex". PLOS Biology 6 (7). July 2008. doi:10.1371/journal.pbio.0060159. PMID 18597554.

- ↑ "How to Direct the Edges of the Connectomes: Dynamics of the Consensus Connectomes and the Development of the Connections in the Human Brain". PLOS ONE 11 (6). 2016. doi:10.1371/journal.pone.0158680. PMID 27362431. Bibcode: 2016PLoSO..1158680K.
- ↑ 115.0 115.1 Crimi, Alessandro; Giancardo, Luca; Sambataro, Fabio; Diego, Sona (2019). "MultiLink analysis: brain network comparison via sparse connectivity analysis". Scientific Reports 9 (1): 1–13. doi:10.1038/s41598-018-37300-4. PMID 30635604. Bibcode: 2019NatSR...9...65C.
- ↑ Zalesky, Andrew; Fornito, Alex; Bullmore, Edward (2010). "Network-based statistic: identifying differences in brain networks.". NeuroImage 53 (4): 1197–1207. doi:10.1016/j.neuroimage.2010.06.041. PMID 20600983.
- ↑ "Graph Theoretical Analysis Reveals: Women's Brains Are Better Connected than Men's". PLOS ONE 10 (7). 2015. doi:10.1371/journal.pone.0130045. PMID 26132764. Bibcode: 2015PLoSO..1030045S.
- ↑ "Brain size bias compensated graph-theoretical parameters are also better in women's structural connectomes". Brain Imaging and Behavior 12 (3): 663–673. June 2018. doi:10.1007/s11682-017-9720-0. PMID 28447246.
- ↑ "Sex differences in the structural connectome of the human brain". Proceedings of the National Academy of Sciences of the United States of America 111 (2): 823–8. January 2014. doi:10.1073/pnas.1316909110. PMID 24297904. Bibcode: 2014PNAS..111..823I.
- ↑ "Age- and gender-related differences in the cortical anatomical network". The Journal of Neuroscience 29 (50): 15684–93. December 2009. doi:10.1523/JNEUROSCI.2308-09.2009. PMID 20016083.
- ↑ "Circular representation of human cortical networks for subject and population-level connectomic visualization". NeuroImage 60 (2): 1340–51. April 2012. doi:10.1016/j.neuroimage.2012.01.107. PMID 22305988.
- ↑ "Patient-tailored connectomics visualization for the assessment of white matter atrophy in traumatic brain injury". Frontiers in Neurology 3: 10. 2012. doi:10.3389/fneur.2012.00010. PMID 22363313.
- ↑ "Mapping connectivity damage in the case of Phineas Gage". PLOS ONE 7 (5). 2012. doi:10.1371/journal.pone.0037454. PMID 22616011. Bibcode: 2012PLoSO...737454V.

- ↑ 124.0 124.1 Hagmann, Patric (2005). From diffusion MRI to brain connectomics (Thesis). Lausanne: EPFL. doi:10.5075/epfl-thesis-3230. Retrieved 2014-01-16.
- ↑ Seung, Sebastian (September 2010). "Sebastian Seung: I am my connectome". TEDTalks. https://www.ted.com/talks/sebastian_seung_i_am_my_connectome.
Further reading
- Friston, Karl J., ed (July 2008). "Mapping the structural core of human cerebral cortex". PLOS Biology 6 (7). doi:10.1371/journal.pbio.0060159. PMID 18597554.
- "New map IDs the core of the human brain". 2008-07-02. http://www.brainmysteries.com/Research/New_map_IDs_the_core_of_the_human_brain.asp.
External links
- The NIH Blueprint for Neuroscience Research
- TED talk by Sebastian Seung: "I am my connectome".
- braingraph.org: a database of hundreds of braingraphs, computed from the data of the Human Connectome Project
 |