Biology:Plant microbiome
The plant microbiome, also known as the phytomicrobiome, plays roles in plant health and productivity and has received significant attention in recent years.[1][2] The microbiome has been defined as "a characteristic microbial community occupying a reasonably well-defined habitat which has distinct physio-chemical properties. The term thus not only refers to the microorganisms involved but also encompasses their theatre of activity".[3][4]
Plants live in association with diverse microbial consortia. These microbes, referred to as the plant's microbiota, live both inside (the endosphere) and outside (the episphere) of plant tissues, and play important roles in the ecology and physiology of plants.[5] "The core plant microbiome is thought to comprise keystone microbial taxa that are important for plant fitness and established through evolutionary mechanisms of selection and enrichment of microbial taxa containing essential functions genes for the fitness of the plant holobiont."[6]
Plant microbiomes are shaped by both factors related to the plant itself, such as genotype, organ, species and health status, as well as factors related to the plant's environment, such as management, land use and climate.[7] The health status of a plant has been reported in some studies to be reflected by or linked to its microbiome.[8][1][9][2]
Overview
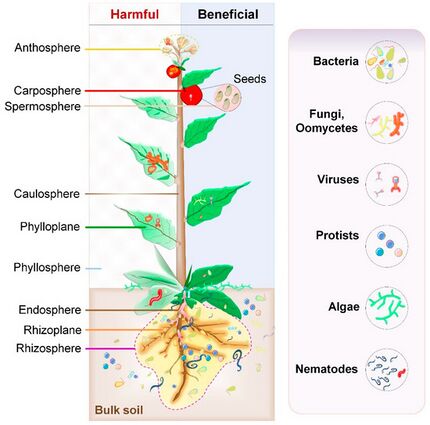
The study of the association of plants with microorganisms precedes that of the animal and human microbiomes, notably the roles of microbes in nitrogen and phosphorus uptake. The most notable examples are plant root-arbuscular mycorrhizal (AM) and legume-rhizobial symbioses, both of which greatly influence the ability of roots to uptake various nutrients from the soil. Some of these microbes cannot survive in the absence of the plant host (obligate symbionts include viruses and some bacteria and fungi), which provides space, oxygen, proteins, and carbohydrates to the microorganisms. The association of AM fungi with plants has been known since 1842, and over 80% of land plants are found associated with them.[11] It is thought AM fungi helped in the domestication of plants.[5]


Traditionally, plant-microbe interaction studies have been confined to culturable microbes. The numerous microbes that could not be cultured have remained uninvestigated, so knowledge of their roles is largely unknown.[5] The possibilities of unraveling the types and outcomes of these plant-microbe interactions has generated considerable interest among ecologists, evolutionary biologists, plant biologists, and agronomists.[8][13][1] Recent developments in multiomics and the establishment of large collections of microorganisms have dramatically increased knowledge of the plant microbiome composition and diversity. The sequencing of marker genes of entire microbial communities, referred to as metagenomics, sheds light on the phylogenetic diversity of the microbiomes of plants. It also adds to the knowledge of the major biotic and abiotic factors responsible for shaping plant microbiome community assemblages.[13][5]
The composition of microbial communities associated with different plant species is correlated with the phylogenetic distance between the plant species, that is, closely related plant species tend to have more alike microbial communities than distant species.[14] The composition of these microbiomes is dynamic and can be modulated by the environment and by climatic conditions.[15] The focus of plant microbiome studies has been directed at model plants, such as Arabidopsis thaliana, as well as important economic crop species including barley (Hordeum vulgare), corn (Zea mays), rice (Oryza sativa), soybean (Glycine max), wheat (Triticum aestivum), whereas less attention has been given to fruit crops and tree species.[16][2]
Plant microbiota
Cyanobacteria are an example of a microorganism which widely interacts in a symbiotic manner with land plants.[17][18][19][20] Cyanobacteria can enter the plant through the stomata and colonise the intercellular space, forming loops and intracellular coils.[21] Anabaena spp. colonize the roots of wheat and cotton plants.[22][23][24] Calothrix sp. has also been found on the root system of wheat.[23][24] Monocots, such as wheat and rice, have been colonised by Nostoc spp.,[25][26][27][28] In 1991, Ganther and others isolated diverse heterocystous nitrogen-fixing cyanobacteria, including Nostoc, Anabaena and Cylindrospermum, from plant root and soil. Assessment of wheat seedling roots revealed two types of association patterns: loose colonization of root hair by Anabaena and tight colonization of the root surface within a restricted zone by Nostoc.[25][29]



(2) On the root surface, cyanobacteria exhibit two types of colonization pattern; in the root hair, filaments of Anabaena and Nostoc species form loose colonies, and in the restricted zone on the root surface, specific Nostoc species form cyanobacterial colonies.
(3) Co-inoculation with 2,4-D and Nostoc spp. increases para-nodule formation and nitrogen fixation. A large number of Nostoc spp. isolates colonize the root endosphere and form para-nodules.[29]
Rhizosphere microbiome
The rhizosphere comprises the 1–10 mm zone of soil immediately surrounding the roots that is under the influence of the plant through its deposition of root exudates, mucilage and dead plant cells.[31] A diverse array of organisms specialize in living in the rhizosphere, including bacteria, fungi, oomycetes, nematodes, algae, protozoa, viruses, and archaea.[32]
Mycorrhizal fungi are abundant members of the rhizosphere community, and have been found in over 200,000 plant species, and are estimated to associate with over 80% of all plants.[33] Mycorrhizae–root associations play profound roles in land ecosystems by regulating nutrient and carbon cycles. Mycorrhizae are integral to plant health because they provide up to 80% of the nitrogen and phosphorus requirements. In return, the fungi obtain carbohydrates and lipids from host plants.[34] Recent studies of arbuscular mycorrhizal fungi using sequencing technologies show greater between-species and within-species diversity than previously known.[35][5]
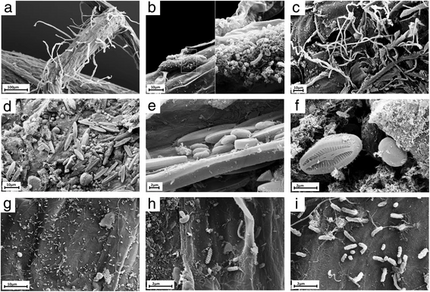
on the roots of Arabidopsis thaliana
a) Overview of an A. thaliana root (primary root) with numerous root hairs. b) Biofilm-forming bacteria. c) Fungal or oomycete hyphae surrounding the root surface. d) Primary root densely covered by spores and protists. e, f) Protists, most likely belonging to the Bacillariophyceae class. g) Bacteria and bacterial filaments. h, i) Different bacterial individuals showing great varieties of shapes and morphological features.[36]

– Berendsen et al, 2012 [8]
The most frequently studied beneficial rhizosphere organisms are mycorrhizae, rhizobium bacteria, plant-growth promoting rhizobacteria (PGPR), and biocontrol microbes. It has been projected that one gram of soil could contain more than one million distinct bacterial genomes,[38] and over 50,000 OTUs (operational taxonomic units) have been found within the potato rhizosphere.[39] Among the prokaryotes in the rhizosphere, the most frequent bacteria are within the Acidobacteriota, Pseudomonadota, Planctomycetota, Actinomycetota, Bacteroidota, and Bacillota.[40][41] In some studies, no significant differences were reported in the microbial community composition between the bulk soil (soil not attached to the plant root) and rhizosphere soil.[42][43] Certain bacterial groups (e.g. Actinomycetota, Xanthomonadaceae) are less abundant in the rhizosphere than in nearby bulk soil .[40][5] Beyond facilitating access to nutrients and to provide protection to biotic and abiotic stresses, certain PGPR modify root architecture.[44]
Endosphere microbiome
Some microorganisms, such as endophytes, penetrate and occupy the plant internal tissues, forming the endospheric microbiome. The arbuscular mycorrhizal and other endophytic fungi are the dominant colonizers of the endosphere.[45] Bacteria, and to some degree archaea, are important members of endosphere communities. Some of these endophytic microbes interact with their host and provide obvious benefits to plants.[40][46][47] Unlike the rhizosphere and the rhizoplane, the endospheres harbor highly specific microbial communities. The root endophytic community can be very distinct from that of the adjacent soil community. In general, diversity of the endophytic community is lower than the diversity of the microbial community outside the plant.[43] The identity and diversity of the endophytic microbiome of above-and below-ground tissues may also differ within the plant.[48][45][5]
In 2025, the "tree microbiome" was presented in an article in the journal Nature. The focus was microbial symbionts regularly found in the wood of trees with even very large trunk diameters. Distinctly different microbes were found not only in diverse tree species but also as they varied between the sapwood and the deep heartwood.[49] As reported in the New York Times, "Sapwood is dominated by microbes that require oxygen, whereas heartwood is dominated by microbes that don't. Much of the methane produced by a tree originates from the heartwood, the study found."[50]
Phyllosphere microbiome

The aerial surface of a plant (stem, leaf, flower, fruit) is called the phyllosphere and is considered comparatively nutrient poor when compared to the rhizosphere and endosphere. The environment in the phyllosphere is more dynamic than the rhizosphere and endosphere environments. Microbial colonizers are subjected to diurnal and seasonal fluctuations of heat, moisture, and radiation. In addition, these environmental elements affect plant physiology (such as photosynthesis, respiration, water uptake etc.) and indirectly influence microbiome composition.[5] Rain and wind also cause temporal variation to the phyllosphere microbiome.[52]
Interactions between plants and their associated microorganisms in many of these microbiomes can play pivotal roles in host plant health, function, and evolution.[53] The leaf surface, or phyllosphere, harbours a microbiome comprising diverse communities of bacteria, fungi, algae, archaea, and viruses.[54][55] Interactions between the host plant and phyllosphere bacteria have the potential to drive various aspects of host plant physiology.[56][57][58] However, as of 2020 knowledge of these bacterial associations in the phyllosphere remains relatively modest, and there is a need to advance fundamental knowledge of phyllosphere microbiome dynamics.[59][60]
Overall, there remains high species richness in phyllosphere communities. Fungal communities are highly variable in the phyllosphere of temperate regions and are more diverse than in tropical regions.[61] There can be up to 107 microbes per square centimetre present on the leaf surfaces of plants, and the bacterial population of the phyllosphere on a global scale is estimated to be 1026 cells.[62] The population size of the fungal phyllosphere is likely to be smaller.[63]
Phyllosphere microbes from different plants appear to be somewhat similar at high levels of taxa, but at the lower levels taxa there remain significant differences. This indicates microorganisms may need finely tuned metabolic adjustment to survive in phyllosphere environment.[61] Pseudomonadota seems to be the dominant colonizers, with Bacteroidota and Actinomycetota also predominant in phyllospheres.[64] Although there are similarities between the rhizosphere and soil microbial communities, very little similarity has been found between phyllosphere communities and microorganisms floating in open air (aeroplankton).[45][5]
The assembly of the phyllosphere microbiome, which can be strictly defined as epiphytic bacterial communities on the leaf surface, can be shaped by the microbial communities present in the surrounding environment (i.e., stochastic colonisation) and the host plant (i.e., biotic selection).[54][62][60] However, although the leaf surface is generally considered a discrete microbial habitat,[65][66] there is no consensus on the dominant driver of community assembly across phyllosphere microbiomes. For example, host-specific bacterial communities have been reported in the phyllosphere of co-occurring plant species, suggesting a dominant role of host selection.[66][45][67][60]
Conversely, microbiomes of the surrounding environment have also been reported to be the primary determinant of phyllosphere community composition.[65][68][61][69] As a result, the processes that drive phyllosphere community assembly are not well understood but unlikely to be universal across plant species. However, the existing evidence does indicate that phyllosphere microbiomes exhibiting host-specific associations are more likely to interact with the host than those primarily recruited from the surrounding environment.[56][70][71][72][60]
The search for a core microbiome in host-associated microbial communities is a useful first step in trying to understand the interactions that may be occurring between a host and its microbiome.[73][74] The prevailing core microbiome concept is built on the notion that the persistence of a taxon across the spatiotemporal boundaries of an ecological niche is directly reflective of its functional importance within the niche it occupies; it therefore provides a framework for identifying functionally critical microorganisms that consistently associate with a host species.[73][75][42][60]
Divergent definitions of "core microbiome" have arisen across scientific literature with researchers variably identifying "core taxa" as those persistent across distinct host microhabitats [76][77] and even different species.[67][70] Given the functional divergence of microorganisms across different host species [67] and microhabitats,[78] defining core taxa sensu stricto as those persistent across broad geographic distances within tissue- and species-specific host microbiomes, represents the most biologically and ecologically appropriate application of this conceptual framework.[79][60] Tissue- and species-specific core microbiomes across host populations separated by broad geographical distances have not been widely reported for the phyllosphere using the stringent definition established by Ruinen.[57][60]
Example: The mānuka phyllosphere
The flowering tea tree commonly known as mānuka is indigenous to New Zealand.[80] Mānuka honey, produced from the nectar of mānuka flowers, is known for its non-peroxide antibacterial properties.[81][82] Microorganisms have been studied in the mānuka rhizosphere and endosphere.[83][84][85] Earlier studies primarily focussed on fungi, and a 2016 study provided the first investigation of endophytic bacterial communities from three geographically and environmentally distinct mānuka populations using fingerprinting techniques and revealed tissue-specific core endomicrobiomes.[86][60]

(B) The chart on the right shows how OTUs in phyllosphere and associated soil communities differed in relative abundances.[60]
A 2020 study identified a habitat-specific and relatively abundant core microbiome in the mānuka phyllosphere, which was persistent across all samples. In contrast, non-core phyllosphere microorganisms exhibited significant variation across individual host trees and populations that was strongly driven by environmental and spatial factors. The results demonstrated the existence of a dominant and ubiquitous core microbiome in the phyllosphere of mānuka.[60]
Seed microbiome
Plant seeds can serve as natural vectors for vertical transmission of their beneficial endophytes, such as those that confer disease resistance. A 2021 research paper explained, "It makes sense that their most important symbionts would be vertically transmitted through seed rather than gambling that all of the correct soil-dwelling microbes might be available at the germination site."[87]
The new paradigm regarding mutualistic fungi and bacterial transmission via the seeds of host plants has been fostered largely by research pertaining to plants of agricultural value.[87][88] Rice seeds were found to entail high microbial diversity, with the greatest diversity inhabiting the embryo rather than the pericarp.[89] Fungi of genus Fusarium transmitted via seeds were found to be dominant members of the microbiome within the stems of maize.[87] This facet of the plant microbiome came to be known as the seed microbiome.[90]
Forestry researchers have also begun to identify members of the seed microbiome pertaining to valuable tree species. Vertical transmission of fungal and bacterial mutualists was confirmed in 2021 for the acorns of oak trees.[48][91] If the research on oaks turns out to apply to other tree species, it will be understood that the above-soil portions of a plant (the phyllosphere) obtain nearly all of their beneficial fungi from those carried in the seed.[48] In contrast, the roots (the rhizosphere) acquire only a small fraction of their mutualists from the seed. Most arrive via the surrounding soil, and this includes their vital associations with arbuscular mycorrhizal fungi.[88]
Microbial species consistently found in plant seeds are known as the "core microbiome."[87][92] Benefits to the host plant include their ability to assist in the production of antimicrobial compounds, detoxification, nutrient uptake, and growth-promoting activities.[48] Discerning the functions of symbiotic microbes in seeds is shifting the agricultural paradigm away from seed breeding and preparation that traditionally sought to minimize the presence of fungal and bacterial propagules. The likelihood that a microbe found within a seed is mutualistic is now a routine presumption. Such partners may contribute to "seed dormancy and germination, environmental adaptation, resistance and tolerance against diseases, and growth promotion."[92]
Application of the new understanding of beneficial microbes inhabiting seeds has been suggested for use beyond agriculture and for biodiversity conservation.[88] A citizen group advocating for northward assisted migration of an endangered tree in the USA has pointed to the seed microbiome paradigm shift as a reason for the official institutions to lift their ban on seed transfer beyond the ex situ conservation plantings in northern Georgia.[93]
Plant holobiont
Since the colonization of land by ancestral plant lineages 450 million years ago, plants and their associated microbes have been interacting with each other, forming an assemblage of species that is often referred to as a holobiont. Selective pressure acting on holobiont components has likely shaped plant-associated microbial communities and selected for host-adapted microorganisms that impact plant fitness. However, the high microbial densities detected on plant tissues, together with the fast generation time of microbes and their more ancient origin compared to their host, suggest that microbe-microbe interactions are also important selective forces sculpting complex microbial assemblages in the phyllosphere, rhizosphere, and plant endosphere compartments.[36]
See also
- Biomass partitioning
- Mangrove microbiome
- Phytobiome
References
- ↑ 1.0 1.1 1.2 Turner, Thomas R.; James, Euan K.; Poole, Philip S. (2013). "The plant microbiome". Genome Biology 14 (6): 209. doi:10.1186/gb-2013-14-6-209. PMID 23805896.
- ↑ 2.0 2.1 2.2 Purahong, Witoon; Orrù, Luigi; Donati, Irene; Perpetuini, Giorgia; Cellini, Antonio; Lamontanara, Antonella; Michelotti, Vania; Tacconi, Gianni et al. (2018). "Plant Microbiome and Its Link to Plant Health: Host Species, Organs and Pseudomonas syringae pv. Actinidiae Infection Shaping Bacterial Phyllosphere Communities of Kiwifruit Plants". Frontiers in Plant Science 9. doi:10.3389/fpls.2018.01563. PMID 30464766. Bibcode: 2018FrPS....9.1563P.. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Whipps, J. M.; Lewis, Karen; Cooke, R. C. (1988). "Mycoparasitism and plant disease control". in Burge, Michael N.. Fungi in biological control systems. Manchester, UK New York: Manchester University Press. pp. 161–187. ISBN 978-0-7190-1979-1.
- ↑ Berg, Gabriele; Rybakova, Daria; Fischer, Doreen; Cernava, Tomislav; Vergès, Marie-Christine Champomier; Charles, Trevor; Chen, Xiaoyulong; Cocolin, Luca et al. (December 2020). "Microbiome definition re-visited: old concepts and new challenges". Microbiome 8 (1): 103. doi:10.1186/s40168-020-00875-0. PMID 32605663.50x50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ 5.00 5.01 5.02 5.03 5.04 5.05 5.06 5.07 5.08 5.09 Dastogeer, K.M., Tumpa, F.H., Sultana, A., Akter, M.A. and Chakraborty, A. (2020) "Plant microbiome–an account of the factors that shape community composition and diversity". Current Plant Biology: 100161. doi:10.1016/j.cpb.2020.100161. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Compant, Stéphane; Samad, Abdul; Faist, Hanna; Sessitsch, Angela (2019). "A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application" (in en). Journal of Advanced Research 19: 29–37. doi:10.1016/j.jare.2019.03.004. PMID 31341667.
- ↑ Bringel, Françoise; Couée, Ivan (2015). "Pivotal roles of phyllosphere microorganisms at the interface between plant functioning and atmospheric trace gas dynamics". Frontiers in Microbiology 06: 486. doi:10.3389/fmicb.2015.00486. PMID 26052316.
- ↑ 8.0 8.1 8.2 Berendsen, Roeland L.; Pieterse, Corné M.J.; Bakker, Peter A.H.M. (2012). "The rhizosphere microbiome and plant health". Trends in Plant Science 17 (8): 478–486. doi:10.1016/j.tplants.2012.04.001. PMID 22564542. Bibcode: 2012TPS....17..478B.
- ↑ Berg, Gabriele; Grube, M.; Schloter, M.; Smalla, K. (2014). "The plant microbiome and its importance for plant and human health". Frontiers in Microbiology 5: 491. doi:10.3389/fmicb.2014.00491. PMID 25278934.
- ↑ Shelake, R.M., Pramanik, D. and Kim, J.Y. (2019) "Exploration of plant-microbe interactions for sustainable agriculture in CRISPR era". Microorganisms, 7(8): 269. doi:10.3390/microorganisms7080269. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Koide, Roger T.; Mosse, Barbara (2004). "A history of research on arbuscular mycorrhiza". Mycorrhiza 14 (3): 145–163. doi:10.1007/s00572-004-0307-4. PMID 15088135. Bibcode: 2004Mycor..14..145K.
- ↑ Backer, Rachel; Rokem, J. Stefan; Ilangumaran, Gayathri; Lamont, John; Praslickova, Dana; Ricci, Emily; Subramanian, Sowmyalakshmi; Smith, Donald L. (2018-10-23). "Plant Growth-Promoting Rhizobacteria: Context, Mechanisms of Action, and Roadmap to Commercialization of Biostimulants for Sustainable Agriculture". Frontiers in Plant Science 9. doi:10.3389/fpls.2018.01473. ISSN 1664-462X. PMID 30405652. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ 13.0 13.1 Bulgarelli, Davide; Schlaeppi, Klaus; Spaepen, Stijn; Van Themaat, Emiel Ver Loren; Schulze-Lefert, Paul (2013). "Structure and Functions of the Bacterial Microbiota of Plants". Annual Review of Plant Biology 64 (1): 807–838. doi:10.1146/annurev-arplant-050312-120106. PMID 23373698. Bibcode: 2013AnRPB..64..807B.
- ↑ Abdelfattah, Ahmed; Tack, Ayco J. M.; Wasserman, Birgit; Liu, Jia; Berg, Gabriele; Norelli, John; Droby, Samir; Wisniewski, Michael (2021). "Evidence for host–microbiome co-evolution in apple" (in en). New Phytologist 234 (6): 2088–2100. doi:10.1111/nph.17820. ISSN 1469-8137. PMID 34823272.
- ↑ Saati-Santamaría, Zaki; Vicentefranqueira, Rocío; Kolařik, Miroslav; Rivas, Raúl; García-Fraile, Paula (2023-07-22). "Microbiome specificity and fluxes between two distant plant taxa in Iberian forests". Environmental Microbiome 18 (1): 64. doi:10.1186/s40793-023-00520-x. ISSN 2524-6372. PMID 37481564. Bibcode: 2023EMicb..18...64S.
- ↑ Busby, Posy E.; Soman, Chinmay; Wagner, Maggie R.; Friesen, Maren L.; Kremer, James; Bennett, Alison; Morsy, Mustafa; Eisen, Jonathan A. et al. (2017). "Research priorities for harnessing plant microbiomes in sustainable agriculture". PLOS Biology 15 (3). doi:10.1371/journal.pbio.2001793. PMID 28350798.
- ↑ Gantar, M.; Elhai, J. (1999). "Colonization of wheatpara-nodules by the N2-fixing cyanobacterium Nostocsp. Strain 2S9B". New Phytologist 141 (3): 373–379. doi:10.1046/j.1469-8137.1999.00352.x.
- ↑ Gantar, M. (2000). "Mechanical damage of roots provides enhanced colonization of the wheat endorhizosphere by the dinitrogen-fixing cyanobacterium Nostoc sp. Strain 2S9B". Biology and Fertility of Soils 32 (3): 250–255. doi:10.1007/s003740000243. Bibcode: 2000BioFS..32..250G.
- ↑ Treves, Haim; Raanan, Hagai; Kedem, Isaac; Murik, Omer; Keren, Nir; Zer, Hagit; Berkowicz, Simon M.; Giordano, Mario et al. (2016). "The mechanisms whereby the green alga Chlorella ohadii , isolated from desert soil crust, exhibits unparalleled photodamage resistance". New Phytologist 210 (4): 1229–1243. doi:10.1111/nph.13870. PMID 26853530. Bibcode: 2016NewPh.210.1229T.
- ↑ Zhu, Huan; Li, Shuyin; Hu, Zhengyu; Liu, Guoxiang (2018). "Molecular characterization of eukaryotic algal communities in the tropical phyllosphere based on real-time sequencing of the 18S rDNA gene". BMC Plant Biology 18 (1): 365. doi:10.1186/s12870-018-1588-7. PMID 30563464. Bibcode: 2018BMCPB..18..365Z.
- ↑ Krings, Michael; Hass, Hagen; Kerp, Hans; Taylor, Thomas N.; Agerer, Reinhard; Dotzler, Nora (2009). "Endophytic cyanobacteria in a 400-million-yr-old land plant: A scenario for the origin of a symbiosis?". Review of Palaeobotany and Palynology 153 (1–2): 62–69. doi:10.1016/j.revpalbo.2008.06.006. Bibcode: 2009RPaPa.153...62K.
- ↑ Karthikeyan, N.; Prasanna, R.; Sood, A.; Jaiswal, P.; Nayak, S.; Kaushik, B. D. (2009). "Physiological characterization and electron microscopic investigation of cyanobacteria associated with wheat rhizosphere". Folia Microbiologica 54 (1): 43–51. doi:10.1007/s12223-009-0007-8. PMID 19330544.
- ↑ 23.0 23.1 Babu, Santosh; Prasanna, Radha; Bidyarani, Ngangom; Singh, Rajendra (2015). "Analysing the colonisation of inoculated cyanobacteria in wheat plants using biochemical and molecular tools". Journal of Applied Phycology 27 (1): 327–338. doi:10.1007/s10811-014-0322-6. Bibcode: 2015JAPco..27..327B.
- ↑ 24.0 24.1 Bidyarani, Ngangom; Prasanna, Radha; Chawla, Gautam; Babu, Santosh; Singh, Rajendra (2015). "Deciphering the factors associated with the colonization of rice plants by cyanobacteria". Journal of Basic Microbiology 55 (4): 407–419. doi:10.1002/jobm.201400591. PMID 25515189.
- ↑ 25.0 25.1 Gantar, M.; Kerby, N. W.; Rowell, P. (1991). "Colonization of wheat (Triticum vulgare L.) by N2-fixing cyanobacteria: II. An ultrastructural study". New Phytologist 118 (3): 485–492. doi:10.1111/j.1469-8137.1991.tb00031.x.
- ↑ Ahmed, Mehboob; Stal, Lucas J.; Hasnain, Shahida (2010). "Association of non-heterocystous cyanobacteria with crop plants". Plant and Soil 336 (1–2): 363–375. doi:10.1007/s11104-010-0488-x. Bibcode: 2010PlSoi.336..363A. http://dare.uva.nl/personal/pure/en/publications/association-of-nonheterocystous-cyanobacteria-with-crop-plants(a90f9b12-6bd1-47cc-87c4-389ab649d969).html.
- ↑ Hussain, Anwar; Hamayun, Muhammad; Shah, Syed Tariq (2013). "Root Colonization and Phytostimulation by Phytohormones Producing Entophytic Nostoc sp. AH-12". Current Microbiology 67 (5): 624–630. doi:10.1007/s00284-013-0408-4. PMID 23794014.
- ↑ Hussain, Anwar; Shah, Syed T.; Rahman, Hazir; Irshad, Muhammad; Iqbal, Amjad (2015). "Effect of IAA on in vitro growth and colonization of Nostoc in plant roots". Frontiers in Plant Science 6: 46. doi:10.3389/fpls.2015.00046. PMID 25699072. Bibcode: 2015FrPS....6...46H.
- ↑ 29.0 29.1 29.2 Lee, Sang-Moo; Ryu, Choong-Min (4 Feb 2021). "Algae as New Kids in the Beneficial Plant Microbiome". Frontiers in Plant Science (Frontiers Media SA) 12. doi:10.3389/fpls.2021.599742. ISSN 1664-462X. PMID 33613596. Bibcode: 2021FrPS...1299742L. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ 30.0 30.1 Chaudhry, Vasvi; Runge, Paul; Sengupta, Priyamedha; Doehlemann, Gunther; Parker, Jane E; Kemen, Eric (10 September 2020). Kopriva, Stanislav. ed. "Shaping the leaf microbiota: plant–microbe–microbe interactions". Journal of Experimental Botany (Oxford University Press (OUP)) 72 (1): 36–56. doi:10.1093/jxb/eraa417. ISSN 0022-0957. PMID 32910810. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Hinsinger, Philippe; Bengough, A. Glyn; Vetterlein, Doris; Young, Iain M. (2009). "Rhizosphere: Biophysics, biogeochemistry and ecological relevance". Plant and Soil 321 (1–2): 117–152. doi:10.1007/s11104-008-9885-9. Bibcode: 2009PlSoi.321..117H.
- ↑ Bonkowski, Michael; Villenave, Cécile; Griffiths, Bryan (2009). "Rhizosphere fauna: The functional and structural diversity of intimate interactions of soil fauna with plant roots". Plant and Soil 321 (1–2): 213–233. doi:10.1007/s11104-009-0013-2. Bibcode: 2009PlSoi.321..213B.
- ↑ Van Der Heijden, Marcel G. A.; Martin, Francis M.; Selosse, Marc-André; Sanders, Ian R. (2015). "Mycorrhizal ecology and evolution: The past, the present, and the future". New Phytologist 205 (4): 1406–1423. doi:10.1111/nph.13288. PMID 25639293. Bibcode: 2015NewPh.205.1406V.
- ↑ Rich, Mélanie K.; Nouri, Eva; Courty, Pierre-Emmanuel; Reinhardt, Didier (2017). "Diet of Arbuscular Mycorrhizal Fungi: Bread and Butter?". Trends in Plant Science 22 (8): 652–660. doi:10.1016/j.tplants.2017.05.008. PMID 28622919. Bibcode: 2017TPS....22..652R. http://doc.rero.ch/record/305034/files/rei_dam.pdf.
- ↑ Lee, Eun-Hwa; Eo, Ju-Kyeong; Ka, Kang-Hyeon; Eom, Ahn-Heum (2013). "Diversity of Arbuscular Mycorrhizal Fungi and Their Roles in Ecosystems". Mycobiology 41 (3): 121–125. doi:10.5941/MYCO.2013.41.3.121. PMID 24198665. Bibcode: 2013Mycob..41..121L.
- ↑ 36.0 36.1 Hassani, M. Amine; Durán, Paloma; Hacquard, Stéphane (March 2018). "Microbial interactions within the plant holobiont". Microbiome 6 (1): 58. doi:10.1186/s40168-018-0445-0. PMID 29587885. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Vishwakarma, Kanchan; Kumar, Nitin; Shandilya, Chitrakshi; Mohapatra, Swati; Bhayana, Sahil; Varma, Ajit (2020). "Revisiting Plant–Microbe Interactions and Microbial Consortia Application for Enhancing Sustainable Agriculture: A Review". Frontiers in Microbiology 11. doi:10.3389/fmicb.2020.560406. PMID 33408698. Bibcode: 2020FrMic..1160406V. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ Gans, J.; Wolinsky, M.; Dunbar, J. (2005). "Computational Improvements Reveal Great Bacterial Diversity and High Metal Toxicity in Soil". Science 309 (5739): 1387–1390. doi:10.1126/science.1112665. PMID 16123304. Bibcode: 2005Sci...309.1387G.
- ↑ i̇Nceoğlu, Özgül; Al-Soud, Waleed Abu; Salles, Joana Falcão; Semenov, Alexander V.; Van Elsas, Jan Dirk (2011). "Comparative Analysis of Bacterial Communities in a Potato Field as Determined by Pyrosequencing". PLOS ONE 6 (8). doi:10.1371/journal.pone.0023321. PMID 21886785. Bibcode: 2011PLoSO...623321I.
- ↑ 40.0 40.1 40.2 Bulgarelli, Davide; Rott, Matthias; Schlaeppi, Klaus; Ver Loren Van Themaat, Emiel; Ahmadinejad, Nahal; Assenza, Federica; Rauf, Philipp; Huettel, Bruno et al. (2012). "Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota". Nature 488 (7409): 91–95. doi:10.1038/nature11336. PMID 22859207. Bibcode: 2012Natur.488...91B.
- ↑ Uroz, Stéphane; Buée, Marc; Murat, Claude; Frey-Klett, Pascale; Martin, Francis (2010). "Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil". Environmental Microbiology Reports 2 (2): 281–288. doi:10.1111/j.1758-2229.2009.00117.x. PMID 23766079. Bibcode: 2010EnvMR...2..281U.
- ↑ 42.0 42.1 Lundberg, Derek S.; Lebeis, Sarah L.; Paredes, Sur Herrera; Yourstone, Scott; Gehring, Jase; Malfatti, Stephanie; Tremblay, Julien; Engelbrektson, Anna et al. (2012). "Defining the core Arabidopsis thaliana root microbiome". Nature 488 (7409): 86–90. doi:10.1038/nature11237. PMID 22859206. Bibcode: 2012Natur.488...86L.
- ↑ 43.0 43.1 Schlaeppi, K.; Dombrowski, N.; Oter, R. G.; Ver Loren Van Themaat, E.; Schulze-Lefert, P. (2014). "Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives". Proceedings of the National Academy of Sciences 111 (2): 585–592. doi:10.1073/pnas.1321597111. PMID 24379374. Bibcode: 2014PNAS..111..585S.
- ↑ Saati-Santamaría, Zaki; González-Dominici, Lihuén Iraí; Jiménez-Gómez, Alejandro; Morais, Daniel; Tláskal, Vojtěch; Frontela, Irene; Benada, Oldřich; Qi, Li et al. (2025-12-05). "Transcriptome-guided discovery of novel plant-associated genes in a rhizosphere Pseudomonas" (in en). Microbiome 14 (1). doi:10.1186/s40168-025-02277-6. ISSN 2049-2618. PMID 41345977.
- ↑ 45.0 45.1 45.2 45.3 Vokou, Despoina; Vareli, Katerina; Zarali, Ekaterini; Karamanoli, Katerina; Constantinidou, Helen-Isis A.; Monokrousos, Nikolaos; Halley, John M.; Sainis, Ioannis (2012). "Exploring Biodiversity in the Bacterial Community of the Mediterranean Phyllosphere and its Relationship with Airborne Bacteria". Microbial Ecology 64 (3): 714–724. doi:10.1007/s00248-012-0053-7. PMID 22544345. Bibcode: 2012MicEc..64..714V.
- ↑ Dastogeer, Khondoker M.G.; Li, Hua; Sivasithamparam, Krishnapillai; Jones, Michael G.K.; Du, Xin; Ren, Yonglin; Wylie, Stephen J. (2017). "Metabolic responses of endophytic Nicotiana benthamiana plants experiencing water stress". Environmental and Experimental Botany 143: 59–71. doi:10.1016/j.envexpbot.2017.08.008. Bibcode: 2017EnvEB.143...59D. https://researchrepository.murdoch.edu.au/id/eprint/38407/.
- ↑ Rodriguez, R. J.; White Jr, J. F.; Arnold, A. E.; Redman, R. S. (2009). "Fungal endophytes: Diversity and functional roles". New Phytologist 182 (2): 314–330. doi:10.1111/j.1469-8137.2009.02773.x. PMID 19236579. Bibcode: 2009NewPh.182..314R.
- ↑ 48.0 48.1 48.2 48.3 Abdelfattah, Ahmed; Wisniewski, Michael; Schena, Leonardo; Tack, Ayco J. M. (2021). "Experimental evidence of microbial inheritance in plants and transmission routes from seed to phyllosphere and root" (in en). Environmental Microbiology 23 (4): 2199–2214. doi:10.1111/1462-2920.15392. ISSN 1462-2920. PMID 33427409. Bibcode: 2021EnvMi..23.2199A.
- ↑ Arnold, Wyatt (28 August 2025). "A diverse and distinct microbiome inside living trees". Nature 644 (8078): 1039–1048. doi:10.1038/s41586-025-09316-0. PMID 40770104. Bibcode: 2025Natur.644.1039A. https://www.nature.com/articles/s41586-025-09316-0.
- ↑ Robles-Gil, Alexa (27 August 2025). "In Every Tree, a Trillion Tiny Lives". New York Times. https://www.nytimes.com/2025/08/27/science/biology-trees-microbiomes.html?unlocked_article_code=1.hU8.9NSf.R-cZ1_pEtZMs.
- ↑ He, Sheng-Yang (28 August 2020). "When plants and their microbes are not in sync, the results can be disastrous". The Conversation. https://theconversation.com/when-plants-and-their-microbes-are-not-in-sync-the-results-can-be-disastrous-143802.
- ↑ Lindow, Steven E. (1996). "Role of Immigration and Other Processes in Determining Epiphytic Bacterial Populations". Aerial Plant Surface Microbiology. pp. 155–168. doi:10.1007/978-0-585-34164-4_10. ISBN 978-0-306-45382-3.
- ↑ Friesen, Maren L.; Porter, Stephanie S.; Stark, Scott C.; von Wettberg, Eric J.; Sachs, Joel L.; Martinez-Romero, Esperanza (2011). "Microbially Mediated Plant Functional Traits". Annual Review of Ecology, Evolution, and Systematics 42: 23–46. doi:10.1146/annurev-ecolsys-102710-145039.
- ↑ 54.0 54.1 Leveau, Johan HJ (2019). "A brief from the leaf: Latest research to inform our understanding of the phyllosphere microbiome". Current Opinion in Microbiology 49: 41–49. doi:10.1016/j.mib.2019.10.002. PMID 31707206. Bibcode: 2019COMb...49...41L.
- ↑ Ruinen, J. (February 1956). "Occurrence of Beijerinckia Species in the 'Phyllosphere'" (in en). Nature 177 (4501): 220–221. doi:10.1038/177220a0. ISSN 1476-4687. Bibcode: 1956Natur.177..220R.
- ↑ 56.0 56.1 Vogel, Christine; Bodenhausen, Natacha; Gruissem, Wilhelm; Vorholt, Julia A. (2016). "The Arabidopsis leaf transcriptome reveals distinct but also overlapping responses to colonization by phyllosphere commensals and pathogen infection with impact on plant health". New Phytologist 212 (1): 192–207. doi:10.1111/nph.14036. PMID 27306148. Bibcode: 2016NewPh.212..192V. https://www.zora.uzh.ch/id/eprint/131259/1/PMID27306148.pdf.
- ↑ 57.0 57.1 Cid, Fernanda P.; Maruyama, Fumito; Murase, Kazunori; Graether, Steffen P.; Larama, Giovanni; Bravo, Leon A.; Jorquera, Milko A. (2018). "Draft genome sequences of bacteria isolated from the Deschampsia antarctica phyllosphere". Extremophiles 22 (3): 537–552. doi:10.1007/s00792-018-1015-x. PMID 29492666.
- ↑ Kumaravel, Sowmya; Thankappan, Sugitha; Raghupathi, Sridar; Uthandi, Sivakumar (2018). "Draft Genome Sequence of Plant Growth-Promoting and Drought-Tolerant Bacillus altitudinis FD48, Isolated from Rice Phylloplane". Genome Announcements 6 (9). doi:10.1128/genomeA.00019-18. PMID 29496824.
- ↑ Laforest-Lapointe, Isabelle; Whitaker, Briana K. (2019). "Decrypting the phyllosphere microbiota: Progress and challenges". American Journal of Botany 106 (2): 171–173. doi:10.1002/ajb2.1229. PMID 30726571. Bibcode: 2019AmJB..106..171L.
- ↑ 60.00 60.01 60.02 60.03 60.04 60.05 60.06 60.07 60.08 60.09 60.10 Noble, Anya S.; Noe, Stevie; Clearwater, Michael J.; Lee, Charles K. (2020). "A core phyllosphere microbiome exists across distant populations of a tree species indigenous to New Zealand". PLOS ONE 15 (8). doi:10.1371/journal.pone.0237079. PMID 32790769. Bibcode: 2020PLoSO..1537079N. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. 50px Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ↑ 61.0 61.1 61.2 Finkel, Omri M.; Burch, Adrien Y.; Lindow, Steven E.; Post, Anton F.; Belkin, Shimshon (2011). "Geographical Location Determines the Population Structure in Phyllosphere Microbial Communities of a Salt-Excreting Desert Tree". Applied and Environmental Microbiology 77 (21): 7647–7655. doi:10.1128/AEM.05565-11. PMID 21926212. Bibcode: 2011ApEnM..77.7647F.
- ↑ 62.0 62.1 Vorholt, Julia A. (2012). "Microbial life in the phyllosphere". Nature Reviews Microbiology 10 (12): 828–840. doi:10.1038/nrmicro2910. PMID 23154261.
- ↑ Lindow, Steven E.; Brandl, Maria T. (2003). "Microbiology of the Phyllosphere". Applied and Environmental Microbiology 69 (4): 1875–1883. doi:10.1128/AEM.69.4.1875-1883.2003. PMID 12676659. Bibcode: 2003ApEnM..69.1875L.
- ↑ Bodenhausen, Natacha; Horton, Matthew W.; Bergelson, Joy (2013). "Bacterial Communities Associated with the Leaves and the Roots of Arabidopsis thaliana". PLOS ONE 8 (2). doi:10.1371/journal.pone.0056329. PMID 23457551. Bibcode: 2013PLoSO...856329B.
- ↑ 65.0 65.1 Stone, Bram W. G.; Jackson, Colin R. (2016). "Biogeographic Patterns Between Bacterial Phyllosphere Communities of the Southern Magnolia (Magnolia grandiflora) in a Small Forest". Microbial Ecology 71 (4): 954–961. doi:10.1007/s00248-016-0738-4. PMID 26883131. Bibcode: 2016MicEc..71..954S.
- ↑ 66.0 66.1 Redford, Amanda J.; Bowers, Robert M.; Knight, Rob; Linhart, Yan; Fierer, Noah (2010). "The ecology of the phyllosphere: Geographic and phylogenetic variability in the distribution of bacteria on tree leaves". Environmental Microbiology 12 (11): 2885–2893. doi:10.1111/j.1462-2920.2010.02258.x. PMID 20545741. Bibcode: 2010EnvMi..12.2885R.
- ↑ 67.0 67.1 67.2 Laforest-Lapointe, Isabelle; Messier, Christian; Kembel, Steven W. (2016). "Host species identity, site and time drive temperate tree phyllosphere bacterial community structure". Microbiome 4 (1): 27. doi:10.1186/s40168-016-0174-1. PMID 27316353.
- ↑ Zarraonaindia, Iratxe; Owens, Sarah M.; Weisenhorn, Pamela; West, Kristin; Hampton-Marcell, Jarrad; Lax, Simon; Bokulich, Nicholas A.; Mills, David A. et al. (2015). "The Soil Microbiome Influences Grapevine-Associated Microbiota". mBio 6 (2). doi:10.1128/mBio.02527-14. PMID 25805735. Bibcode: 2015mBio....652714Z.
- ↑ Finkel, Omri M.; Burch, Adrien Y.; Elad, Tal; Huse, Susan M.; Lindow, Steven E.; Post, Anton F.; Belkin, Shimshon (2012). "Distance-Decay Relationships Partially Determine Diversity Patterns of Phyllosphere Bacteria on Tamrix Trees across the Sonoran Desert". Applied and Environmental Microbiology 78 (17): 6187–6193. doi:10.1128/AEM.00888-12. PMID 22752165. Bibcode: 2012ApEnM..78.6187F.
- ↑ 70.0 70.1 Kembel, S. W.; O'Connor, T. K.; Arnold, H. K.; Hubbell, S. P.; Wright, S. J.; Green, J. L. (2014). "Relationships between phyllosphere bacterial communities and plant functional traits in a neotropical forest". Proceedings of the National Academy of Sciences 111 (38): 13715–13720. doi:10.1073/pnas.1216057111. PMID 25225376. Bibcode: 2014PNAS..11113715K.
- ↑ Innerebner, Gerd; Knief, Claudia; Vorholt, Julia A. (2011). "Protection of Arabidopsis thaliana against Leaf-Pathogenic Pseudomonas syringae by Sphingomonas Strains in a Controlled Model System". Applied and Environmental Microbiology 77 (10): 3202–3210. doi:10.1128/AEM.00133-11. PMID 21421777. Bibcode: 2011ApEnM..77.3202I.
- ↑ Lajoie, Geneviève; Maglione, Rémi; Kembel, Steven W. (2020). "Adaptive matching between phyllosphere bacteria and their tree hosts in a neotropical forest". Microbiome 8 (1): 70. doi:10.1186/s40168-020-00844-7. PMID 32438916. Bibcode: 2020Micb....8...70L.
- ↑ 73.0 73.1 Shade, Ashley; Handelsman, Jo (2012). "Beyond the Venn diagram: The hunt for a core microbiome". Environmental Microbiology 14 (1): 4–12. doi:10.1111/j.1462-2920.2011.02585.x. PMID 22004523. Bibcode: 2012EnvMi..14....4S.
- ↑ Berg, Gabriele; Rybakova, Daria; Fischer, Doreen; Cernava, Tomislav; Vergès, Marie-Christine Champomier; Charles, Trevor; Chen, Xiaoyulong; Cocolin, Luca et al. (2020). "Microbiome definition re-visited: Old concepts and new challenges". Microbiome 8 (1): 103. doi:10.1186/s40168-020-00875-0. PMID 32605663.
- ↑ Turnbaugh, Peter J.; Hamady, Micah; Yatsunenko, Tanya; Cantarel, Brandi L.; Duncan, Alexis; Ley, Ruth E.; Sogin, Mitchell L.; Jones, William J. et al. (2009). "A core gut microbiome in obese and lean twins". Nature 457 (7228): 480–484. doi:10.1038/nature07540. PMID 19043404. Bibcode: 2009Natur.457..480T.
- ↑ Hamonts, Kelly; Trivedi, Pankaj; Garg, Anshu; Janitz, Caroline; Grinyer, Jasmine; Holford, Paul; Botha, Frederik C.; Anderson, Ian C. et al. (2018). "Field study reveals core plant microbiota and relative importance of their drivers". Environmental Microbiology 20 (1): 124–140. doi:10.1111/1462-2920.14031. PMID 29266641. Bibcode: 2018EnvMi..20..124H.
- ↑ Cernava, Tomislav; Erlacher, Armin; Soh, Jung; Sensen, Christoph W.; Grube, Martin; Berg, Gabriele (2019). "Enterobacteriaceae dominate the core microbiome and contribute to the resistome of arugula (Eruca sativa Mill.)". Microbiome 7 (1): 13. doi:10.1186/s40168-019-0624-7. PMID 30696492.
- ↑ Leff, Jonathan W.; Del Tredici, Peter; Friedman, William E.; Fierer, Noah (2015). "Spatial structuring of bacterial communities within individual Ginkgo bilobatrees". Environmental Microbiology 17 (7): 2352–2361. doi:10.1111/1462-2920.12695. PMID 25367625. Bibcode: 2015EnvMi..17.2352L.
- ↑ Hernandez-Agreda, Alejandra; Gates, Ruth D.; Ainsworth, Tracy D. (2017). "Defining the Core Microbiome in Corals' Microbial Soup". Trends in Microbiology 25 (2): 125–140. doi:10.1016/j.tim.2016.11.003. PMID 27919551.
- ↑ Stephens, J. M. C.; Molan, P. C.; Clarkson, B. D. (2005). "A review of Leptospermum scoparium(Myrtaceae) in New Zealand". New Zealand Journal of Botany 43 (2): 431–449. doi:10.1080/0028825X.2005.9512966. Bibcode: 2005NZJB...43..431S.
- ↑ Cooper, R.A.; Molan, P.C.; Harding, K.G. (2002). "The sensitivity to honey of Gram-positive cocci of clinical significance isolated from wounds". Journal of Applied Microbiology 93 (5): 857–863. doi:10.1046/j.1365-2672.2002.01761.x. PMID 12392533. Bibcode: 2002JApMb..93..857C.
- ↑ Rabie, Erika; Serem, June Cheptoo; Oberholzer, Hester Magdalena; Gaspar, Anabella Regina Marques; Bester, Megan Jean (2016). "How methylglyoxal kills bacteria: An ultrastructural study". Ultrastructural Pathology 40 (2): 107–111. doi:10.3109/01913123.2016.1154914. PMID 26986806.
- ↑ Johnston, Peter R. (1998). "Leaf endophytes of manuka (Leptospermum scoparium)". Mycological Research 102 (8): 1009–1016. doi:10.1017/S0953756297005765.
- ↑ McKenzie, E. H. C.; Johnston, P. R.; Buchanan, P. K. (2006). "Checklist of fungi on teatree (Kunzeaand Leptospermumspecies) in New Zealand". New Zealand Journal of Botany 44 (3): 293–335. doi:10.1080/0028825X.2006.9513025. Bibcode: 2006NZJB...44..293M.
- ↑ Wicaksono, Wisnu Adi; Sansom, Catherine E.; Eirian Jones, E.; Perry, Nigel B.; Monk, Jana; Ridgway, Hayley J. (2018). "Arbuscular mycorrhizal fungi associated with Leptospermum scoparium (Mānuka): Effects on plant growth and essential oil content". Symbiosis 75 (1): 39–50. doi:10.1007/s13199-017-0506-3. Bibcode: 2018Symbi..75...39W.
- ↑ Wicaksono, Wisnu Adi; Jones, E. Eirian; Monk, Jana; Ridgway, Hayley J. (2016). "The Bacterial Signature of Leptospermum scoparium (Mānuka) Reveals Core and Accessory Communities with Bioactive Properties". PLOS ONE 11 (9). doi:10.1371/journal.pone.0163717. PMID 27676607. Bibcode: 2016PLoSO..1163717W.
- ↑ 87.0 87.1 87.2 87.3 Johnston-Monje, David; Gutierrez, Janneth P; Lopez-Lavalle, Luis A B (2021). "Seed-Transmitted Bacteria and Fungi Dominate Juvenile Plant Microbiomes". Frontiers in Microbiology 12. doi:10.3389/fmicb.2021.737616. PMID 34745040. Bibcode: 2021FrMic..1237616J.
- ↑ 88.0 88.1 88.2 War, Aadil Farooq; Bashir, Iqra; Reshi, Zafar A.; Kardol, Paul; Rashid, Irfan (April 2023). "Insights into the seed microbiome and its ecological significance in plant life". Microbiological Research 269. doi:10.1016/j.micres.2023.127318. PMID 36753851.
- ↑ Matsumoto, Haruna; Fan, Xiaoyan; Wang, Yue; Kusstatscher, Peter; Duan, Jie; Wu, Sanling; Chen, Sunlu; Qiao, Kun et al. (4 January 2021). "Bacterial seed endophyte shapes disease resistance in rice". Nature Plants 7 (1): 60–72. doi:10.1038/s41477-020-00826-5. PMID 33398157. Bibcode: 2021NatPl...7...60M.
- ↑ Nelson, Eric B (2018). "REVIEW: The seed microbiome: Origins, interactions, and impacts". Plant and Soil 422: 7–34. doi:10.1007/s11104-017-3289-7.
- ↑ "Seeds transfer their microbes to the next generation" (in en). https://www.eurekalert.org/pub_releases/2021-01/su-stt012121.php.
- ↑ 92.0 92.1 Abdelfattah, Ahmed; Tack, Ayco J.M.; Lobato, Carolina; Wassermann, Birgit; Berg, Gabriele (April 2023). "From seed to seed: the role of microbial inheritance in the assembly of the plant microbiome". Trends in Microbiology 31 (4): 346–355. doi:10.1016/j.tim.2022.10.009. PMID 36481186. Bibcode: 2023TrMic..31..346A.
- ↑ Barlow, Connie. "Published Documents on Endangerment Causes of Torreya taxifolia in Florida". https://www.torreyaguardians.org/extinction-papers.html.
Reference books
- Saleem M (2015) Microbiome Community Ecology: Fundamentals and Applications Springer. ISBN 9783319116655.
- Kumar V, Prasad R, Kumar M and Choudhary DK (2019) Microbiome in Plant Health and Disease: Challenges and Opportunities Springer. ISBN 9789811384950.
- Kumar, Vivek; Prasad, Ram; Kumar, Manoj; Choudhary, Devendra K. (10 August 2019). Microbiome in Plant Health and Disease: Challenges and Opportunities. Springer. ISBN 978-981-13-8495-0. https://books.google.com/books?id=dY2oDwAAQBAJ&q=%22Microbiome+in+Plant+Health+and+Disease%22.
- Grube, Martin; Schloter, Michael; Smalla, Kornelia; Berg, Gabriele (22 January 2015). The plant microbiome and its importance for plant and human health. 5. 491. doi:10.3389/fmicb.2014.00491. ISBN 978-2-88919-378-3. https://books.google.com/books?id=NtlpBgAAQBAJ&q=plant+microbiome.
- Egamberdieva, Dilfuza; Ahmad, Parvaiz (6 February 2018). Plant Microbiome: Stress Response. Springer. ISBN 978-981-10-5514-0. https://books.google.com/books?id=-cNKDwAAQBAJ&q=plant+microbiome.
- Castiglione, Stefano; Cicatelli, Angela; Ferrol, Nuria; Rozpadek, Piotr (22 August 2019). Effects of Plant-Microbiome Interactions on Phyto- and Bio-Remediation Capacity. Frontiers Media SA. ISBN 978-2-88945-932-2. https://books.google.com/books?id=XniqDwAAQBAJ&q=plant+microbiome.
- Castiglione, Stefano; Cicatelli, Angela; Ferrol, Nuria; Rozpadek, Piotr (22 August 2019). Effects of Plant-Microbiome Interactions on Phyto- and Bio-Remediation Capacity. Frontiers Media SA. ISBN 978-2-88945-932-2. https://books.google.com/books?id=XniqDwAAQBAJ&q=plant+microbiome.
- Carvalhais, Lilia C.; Dennis, Paul G. (30 December 2020). The Plant Microbiome: Methods and Protocols. Springer. ISBN 978-1-0716-1039-8. https://books.google.com/books?id=-L6WzQEACAAJ&q=plant+microbiome.
- Kumar, Vivek; Prasad, Ram; Kumar, Manoj; Choudhary, Devendra K. (10 August 2019). Microbiome in Plant Health and Disease: Challenges and Opportunities. Springer. ISBN 978-981-13-8495-0. https://books.google.com/books?id=dY2oDwAAQBAJ&q=plant+microbiome.
- Solanki, Manoj Kumar; Kashyap, Prem Lal; Ansari, Rizwan Ali; Kumari, Baby (28 August 2020). Microbiomes and Plant Health: Panoply and Their Applications. Academic Press. ISBN 978-0-12-822601-8. https://books.google.com/books?id=uujeDwAAQBAJ&q=plant+microbiome.
- Yadav, Ajar (2020). Advances in plant microbiome and sustainable agriculture functional annotation and future challenges. Singapore: Springer. ISBN 978-981-15-3204-7. OCLC 1193118045. https://books.google.com/books?id=_6L0DwAAQBAJ&q=plant+microbiome.
- Varma, Ajit (28 August 2020). Plant Microbiome Paradigm. Springer. ISBN 978-3-030-50395-6. https://books.google.com/books?id=h_75DwAAQBAJ&q=plant+microbiome&pg=PA35.
- Doty, Sharon Lafferty (21 September 2017). Functional Importance of the Plant Microbiome: Implications for Agriculture, Forestry and Bioenergy. Springer. ISBN 978-3-319-65897-1. https://books.google.com/books?id=Mvo2DwAAQBAJ&q=plant+microbiome&pg=PA1.
- Antwis, Rachael E.; Harrison, Xavier A.; Cox, Michael J. (12 March 2020). Microbiomes of Soils, Plants and Animals: An Integrated Approach. Cambridge University Press. ISBN 978-1-108-47371-2. https://books.google.com/books?id=GljPDwAAQBAJ&q=plant+microbiome.
- Varma, Ajit; Tripathi, Swati; Prasad, Ram (20 October 2020). Plant Microbiome Paradigm. Springer. ISBN 978-3-030-50394-9. https://books.google.com/books?id=d6WQzQEACAAJ&q=plant+microbiome.
- Rosenberg, Eugene; Zilber-Rosenberg, Ilana (31 January 2014). The Hologenome Concept: Human, Animal and Plant Microbiota. Springer. ISBN 978-3-319-04241-1. https://books.google.com/books?id=YY24BAAAQBAJ&q=plant+microbiome.
- Yadav, Ajar Nath; Singh, Joginder; Rastegari, Ali Asghar; Yadav, Neelam (6 March 2020). Plant Microbiomes for Sustainable Agriculture. Springer. ISBN 978-3-030-38453-1. https://books.google.com/books?id=HEHVDwAAQBAJ&q=plant+microbiome.
 |