Biology:Protist
A protist (/ˈproʊtɪst/ PROH-tist) or protoctist is any eukaryotic organism that is not an animal, land plant, or fungus. Protists do not form a natural group, or clade, but are a paraphyletic group encompassing the entire eukaryote tree of life, from which land plants, animals, and fungi evolved. They are primarily single-celled, exhibiting a wide range of forms such as amoebae, ciliates, thick-walled microalgae and, more commonly, flagellates. Several transitions to multicellularity have occurred among protists, from colonies with alternating cell types to giant slime molds, fungus-like organisms, and seaweeds with differentiated tissues.
Protists were historically regarded as a separate taxonomic kingdom known as Protista or Protoctista, or were lumped as part of the traditional plant and animal kingdoms as algae and protozoa, respectively. With the advent of molecular phylogenetics and electron microscopy studies, some protists were shown to be more closely related to animals or plants than to other protists, and algae were found to be intermixed with protozoa. The classification suffered major revisions, as seemingly unrelated forms were found to be evolutionarily related, and viceversa.
In modern classifications, protists are spread across several large clades known as supergroups, many of them containing disparate forms. For example, the Archaeplastida includes mostly phototrophs like red and green algae, from which land plants evolved. Opisthokonta groups fungi, animals, and their single-celled relatives. Amoebozoa and Rhizaria harbor the majority of amoeboid organisms, such as testate amoebae, foraminifers and radiolarians. Stramenopiles and Alveolata are diverse groups of flagellates, many of which have evolved into major parasites (e.g., oomycetes, apicomplexans) or phototrophs (diatoms, brown algae, dinoflagellates). The earliest diverging groups, collectively known as Excavata (e.g., euglenids, metamonads), are flagellates that represent the ancestral traits of the last eukaryotic common ancestor (LECA). Despite the comparatively low number of described species, protists compose the majority of eukaryotic diversity as indicated by environmental DNA studies. Most protists are yet undescribed.
Protists encompass almost all of the biological traits seen in eukaryotes, and many exhibit unique adaptations. These include a range of nutritional modes through specialized feeding structures (phagotrophy, osmotrophy, myzocytosis) or chloroplasts (phototrophy), often mixing both as mixotrophy. Cellular respiration also varies due to modifications of their mitochondria. Almost all protists have a complex cytoskeleton composed of relatively conserved structures across evolution, namely a flagellar apparatus with basal bodies from which microtubules emerge and support the remaining cellular structures. Many protists have unique organelles that serve other functions, such as contractile vacuoles for homeostasis, or eyespots for light perception. Protist cells tend to host symbionts such as bacteria and archaea, usually to support their metabolism and nutrition. Although traditionally presumed to be asexual, protists are capable of sexual reproduction, and can exhibit diverse and complex life cycles with different generations and life stages.
Protists are abundantly present in all ecosystems, including extreme habitats, as important components of the biogeochemical cycles and trophic webs. As producers, they are responsible for a large portion of global primary production and carbon fixation. As consumers and decomposers, they regulate fungal and bacterial populations, and release nutrients to other trophic levels. Some form mutualistic relationships with other protists or animals such as corals and termites. Others are important parasites. Pathogenic protists cause many well-known human and animal diseases such as malaria and toxoplasmosis, or significant plant diseases like clubroot and potato blight. Free-living protists can also negatively impact aquatic life as harmful algal blooms.
The early evolution of protists corresponds with the evolution of eukaryotes, which split from archaea around 3 billion years ago and eventually gave rise to a common ancestor (LECA) with essential traits such as mitochondria and a complex endomembrane system, some time during the Paleo- or Mesoproterozoic eras. In the gap between these two events, fossils are often interpreted as stem-group eukaryotes, with intermediate traits. Following the appearance of LECA, its descendants (crown-group eukaryotes) experienced a rapid diversification in the span of 300 million years that originated the modern supergroups. Still, their abundance in the fossil record remained low until the Neoproterozoic, when the first fossils of opisthokonts, amoebae, and multicellular algae appear. Throughout the Phanerozoic, protists evolved into the forms that dominate ecosystems today, leaving an extensive fossil record of primarily siliceous and calcareous shells.
Definition
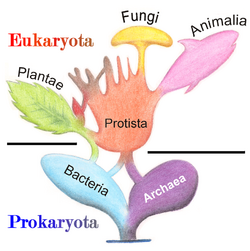
Protists are defined as all eukaryotes that are not animals, land plants or fungi, the three traditional "higher" kingdoms of eukaryotes. Because of this definition by exclusion, protists compose a paraphyletic group from which the ancestors of those three kingdoms evolved.[1][2] As such, there is no unique trait that unifies all protists yet excludes non-protists. Still, together they exhibit a remarkable diversity of life cycles, trophic levels, modes of locomotion, and cellular structures that dwarfs those seen in "higher" eukaryotes.[3] A less popular view is that protists are defined as exclusively single-celled eukaryotes,[4] but this disregards the various transitions to multicellularity among protists.[1]
The distinction between protists and other kingdoms was blurry before genetic analysis. Organisms that are unquestionably known as protists include a wide range of photosynthetic species, known as "algae", including various types of macroalgae that have a similar complexity to plants. Other protists have a fungus-like nutrition or appearance, such as the oomycetes. The remaining heterotrophic protists are often called "protozoa".[1][5] Some minuscule animals (the myxozoans)[6] and the "lower" fungi (namely the aphelids, rozellids and microsporidians, collectively the opisthosporidians)[7] were traditionally classified as protists,[8][9] and some algae (particularly red and green algae) were lumped with plants.[10]
According to the current consensus, the label 'protist' specifically excludes animals, embryophytes (land plants) —meaning that all eukaryotic algae fall under this label— and all fungi. Opisthosporidians are considered part of a larger fungal kingdom, although they are studied by protistologists and mycologists alike.[11][12][13]
Morphology
Protists exist in diverse shapes and sizes.[14]: 27 While most are single-celled, others have evolved various forms of multicellularity, ranging from simple colonies of cells to large, complex organisms like giant kelp and slime molds.[15] Single-celled protists are traditionally grouped by their shape and how they move, into four broad categories.[16]: 5
- Flagellates are the most common protists, and very likely the most abundant eukaryotes on Earth.[17] They move using one or more whip-like structures called flagella.[lower-alpha 1] Most are heterotrophic (known as zooflagellates), feeding on bacteria or other organisms, ranging from filter feeders like choanoflagellates to active predators like provorans. Many are photo- or mixotrophic (known as phytoflagellates) and are studied as algae, like the dinoflagellates.[21][22]
- Amoebae are known for their often flexible shape and ability to form extensions of the cytoplasm known as pseudopodia. These extensions come in various forms, such as lobose (blunt, rounded, as in Amoeba), filose (thin, tapering, as in cercozoans), or reticulose (branching networks, as in foraminifers). Some, called axopodia, take the shape of radiating projections supported by microtubules, characteristic of heliozoa and radiolaria.[14]: 23 Some amoebae can grow to sizes visible to the naked eye, reaching up to 20 cm.[23] Amoeboflagellates can produce both pseudopodia and flagella within the same life cycle.[24]
- Ciliates have larger cells with two types of nuclei and rows of small flagella, known as cilia.[lower-alpha 1] They are often at the top of the microbial food web. Although ciliates compose a single lineage,[25] some protists have evolved unusual large ciliate-like cells, such as the opalines.[26]
- Certain parasitic protists traditionally described as sporozoa[27] are immobile in their adult stage and reproduce through thick-walled spores, such as haplosporidians and apicomplexans. This term also included fungi and animals that have evolved a similar lifestyle—microsporidians and myxozoans, respectively. Other than their lifestyle, they have very little in common, and have evolved in distantly related groups.[28][16]: 8–9
Other single-celled algae exist in forms beyond the motile flagellates. Some are non-motile and encased in hard cell walls (coccoid, like diatoms) or embedded in a mucilage matrix (capsalean, like glaucophytes);[29] others are amoeboid, like the reticulose chlorarachniophytes.[30][31]
Multicellularity has evolved numerous times to various degrees among protists, resulting in organisms built either by cells aggregating together (aggregative) or by cells dividing without separating (clonal).[15] For example:
- Fungus-like protists, traditionally studied by mycologists, are osmotrophic and produce macroscopic fruiting bodies for dispersal (sorocarps if aggregative, sporocarps if clonal). They may have hyphae-like bodies, or they may be slime molds, composed of individual amoebae until they form fruiting bodies.[32][33][34] Some can reach lengths of several meters.[14]: 29
- Multicellular algae range from simple filaments and coenocytes to the highly complex brown algae, which have fully differentiated tissues (parenchymatous) resembling those of plants, or the red algae, with only partial differentiation (pseudoparenchymatous).[31][35] In green algae alone multicellularity is thought to have evolved over 20 separate times, with some groups like the stoneworts developing specialized reproductive organs.[36][15]
Other multicellular protists include amoebae that fuse into large networks, and colonial heliozoa and ciliates with new features not seen in solitary cells. The ciliate Haplozoon is interpreted to have animal-like embryonic development and cell type differentiation.[15] Choanoflagellates, the closest living relatives of animals, include alternating cell types that are interpreted as early stages of animal multicellularity.[37]
Classification
Brief history
Starting in the 17th and 18th centuries, after the discovery of microscopic life by Antonie van Leeuwenhoek, the classification of single-celled protists was largely based on observations under light microscopy. Protists were incorporated into the traditional dichotomy that defined all life as either plant or animal: non-motile algae were considered part of the plant kingdom, and all other protists joined the animal kingdom.[5][38] They were popularly known as "infusion animals" or infusoria, together with bacteria and small invertebrates. Otto Friedrich Müller was the first to introduce microbial protists to the Linnean system of binomial nomenclature.[39][40]
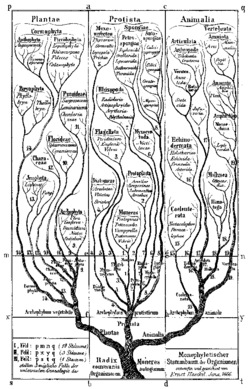
During the 19th century, after several waves of naturalist studies,[40] it became clear that these microorganisms were distinct from animals and plants. John Hogg and Ernst Haeckel proposed a separate kingdom of life, named Protoctista[41] or Protista,[42] respectively, to accommodate the predominantly unicellular eukaryotes, and initially bacteria, which were later excluded.[5] The classical framework of protist classification was established, as exemplified by the works of Otto Bütschli, where they were grouped according to morphological and locomotive features, such as Mastigophora (flagellates), Rhizopoda (amoebae), Sporozoa (spore-forming parasites), and Infusoria (ciliates).[38] However, Bütschli retained a division between the Protozoa (animal-like protists) and Protophyta (plant-like). This dogma remained dominant throughout the early 20th century.[5]
From the mid-20th century, eukaryotes were firmly split from bacteria (prokaryotes) due to the presence of the cell nucleus,[43] and protists (or protoctists) were more popularly accepted as a separate kingdom of eukaryotes.[44][45][46] The advent of electron microscopy shifted the methods of classification, as it revealed previously unrecognized cellular characteristics (i.e., ultrastructure, particularly of the flagellar apparatus and the cytoskeleton) that suggested evolutionary affinities between superficially disparate lineages. For example, the tripartite flagellar mastigonemes were used to group heterokont algae, oomycetes and opalines into the Stramenopiles; the discovery of cortical alveoli showed affinities between dinoflagellates and ciliates, which now belong to the Alveolata; and disc-shaped mitochondrial cristae were shared by kinetoplastids and euglenids, now united as Euglenozoa. The algae-protozoa dichotomy became obsolete.[47][48][38]

Since the 1990s, molecular phylogenetic analyses based primarily on the SSU rRNA gene demonstrated that protists were a paraphyletic assemblage of clades spanning the entire eukaryotic tree of life, from which the other three "kingdoms" (animals, plants, and fungi) had evolved.[49][38] Beginning in the 2000s, single-cell sequencing and phylogenomics technologies progressively improved the resolution of deeper evolutionary relationships. Altogether, these innovations led to successive revisions of protist classification,[38] such as the ones published by the International Society of Protistologists.[50][51][11] Eukaryotes could no longer be divided into four monophyletic kingdoms,[lower-alpha 2] and instead are arranged in "supergroups", each often encompassing an unexpected variety of morphologies and lifestyles that do not resemble one another. New deep-branching groups are added to the tree at a rate of nearly one per year.[55][56][57][58]
Modern classification and diversity
Protists are currently divided among a number of clades informally named supergroups. Most of these supergroups fall under either of two large clades of eukaryotes: Amorphea and Diaphoretickes. The animals and fungi belong to the Opisthokonta supergroup in the Amorphea clade, along with several other groups of protists (e.g., Amoebozoa).[11] Diaphoretickes contains the diverse supergroups Archaeplastida (including plants), Stramenopiles, Alveolata, Rhizaria (combined as the SAR supergroup), and the less species-rich Cryptista, Haptista, Telonemia, and Disparia.[59][60][61] Outside of these larger clades, various groups of protists with primitive cell architecture (Discoba, Metamonada, and Malawimonadida) are collectively known as the excavates or "Excavata". The name 'excavate' refers to the shared characteristic of a ventral groove in the cell used for feeding, which is considered an ancestral trait present in the last eukaryotic common ancestor.[62][63]: 597 [64]
The following table lists estimated numbers of described extant species for all known protist supergroups, and provides an overview of their diversity in terms of morphologies, habitats, and nutritional modes. For large groups, the overview is not exhaustive and only mentions the most characteristic members.
| Clade | Supergroup | Example | Brief description of morphology, lifestyle and habitat | Living species | |
|---|---|---|---|---|---|
| Diaph. | SAR | Stramenopiles | 
|
Ancestrally flagellates distinguished by two 'heterokont' (unequal) flagella, one with tripartite mastigonemes. Present in virtually all habitats. The most species-rich lineage, the ochrophytes, are algae of diverse morphologies, ranging from flagellates (like golden algae) to walled ornamented cells (like diatoms, pictured) to truly multicellular macroalgae with differentiated tissues (brown algae such as kelp). All other lineages are composed of heterotrophs: bacterivorous flagellates (e.g., bicosoecids, bigyromonads), fungus-like osmotrophs (oomycetes, hyphochytrids, and labyrinthulomycetes), heliozoan amoebae (actinophryids), and ciliate-like obligate symbionts of animals (opalinids).[65] | over 100,000[66] |
| Alveolata | 
|
Ancestrally flagellated predators with cortical alveoli. The colponemids represent these ancestral characteristics.[67] The most diverse group are the ciliates (pictured), with large cells covered in rows of cilia, usually at the top of the microbial food chain.[68] The remaining alveolates belong to the clade Myzozoa and are ancestrally photosynthetic; some have retained their photosynthetic ability (chromerids and many dinoflagellates), while others have evolved into parasites of animals and algae (apicomplexans, perkinsozoans, and some dinoflagellates).[69][11][70][71] | over 10,000[67] | ||
| Rhizaria | Error creating thumbnail: | Amoebae with filose or reticulose pseudopodia.[72] The most species-rich group is Retaria, home to conspicuous marine amoebae encased in hard skeletons (radiolarians) or multichambered tests (foraminifers, pictured). Secondly is Cercozoa, with an extreme diversity of morphologies: small flagellates, amoeboflagellates, aggregative slime molds,[73] testate amoebae, heliozoa, and massive radiolarian-like cells (phaeodarians);[74] some are capable of photosynthesis (e.g., chlorarachniophytes). Lastly, Endomyxa contains both free-living predatory amoebae (e.g., vampyrellids) and obligate parasites of animals, plants, and algae (e.g., phytomyxeans and ascetosporeans).[75][11] | over 11,000[76] | ||
| Telonemia | File:Rsob210325f05h Telonema subtile.png | Free-living flagellates with a unique cytoskeleton and a combination of cell structures. Present in all marine and freshwater environments feeding on bacteria.[77] | 10[78][79] | ||
| Haptista | File:9Calcidiscus leptoporus, diploid, SEM, showing coccoliths.tif | Two groups of different free-living single-celled protists: centrohelids—predatory heliozoan amoebae, widespread in aquatic and soil environments[80]—and haptophytes—coccoid or flagellated photosynthetic algae, mostly marine (e.g., coccolithophores, pictured).[81] Both can produce an outer coat of complex mineralized scales.[11] | over 600 | ||
| Pancryptista | File:Cryptomonas uralensis EurJTaxonom 2020 fig2.png | Free-living flagellates, except one species of heliozoan amoebae, Microheliella maris.[82] Almost all of the flagellates are distinguished by specialized ribbon-shaped extrusomes known as ejectisomes. Many are photosynthetic, known as cryptomonads (pictured), while the rest are phagotrophs, consumers of bacteria. Present in aquatic environments worldwide.[83][11] | over 100 | ||
| Archaeplastida* | File:Volvox aureus.jpg | Algae with chloroplasts derived from primary endosymbiosis with a cyanobacterium. Found in all environments. Almost entirely photosynthetic, with the exception of two small groups of phagotrophic flagellates, rhodelphids[84] and picozoans.[85] The two major groups, red algae and green algae (pictured), exhibit diverse morphologies, ranging from single cells—coccoid, palmelloid, sarcinoid, flagellated—to colonies, simple filaments, and macroscopic thalli with varying degrees of complexity (e.g., coralline algae, sea lettuce, stoneworts). Also included are glaucophytes, rare blue-green algae found in surface waters.[86] | over 20,500[87]* | ||
| Disparia | File:Solarion arienae (morphology and ultrastructure) crop.png | Three lineages of free-living predatory flagellates with unique cytoskeletons. These are: Hemimastigophora, with two rows of flagella, present in soils and aquatic sediments;[55] Provora, fast-swimming predators of other protists through a strong feeding apparatus resembling jaws, found in low abundance in marine environments globally;[88][89] and Caelestes (pictured), rare inhabitants of the marine benthos whose cells protrude arms or stalks used for movement or prey capture.[61] | 20 | ||
| Amorph. | Amoebozoa | File:Stemonitopsis typhina 449034989.png | Amoebae of diverse morphologies, with lobose or filose pseudopodia, and sometimes with flagella. Most are free-living phagotrophs found across terrestrial and aquatic environments, such as the archetypal genus Amoeba itself,[90][91][92] or the testate amoebae Arcellinida, one of the most conspicuous groups of protists.[93] Numerous groups have independently evolved fungus-like fruiting bodies,[94] such as myxomycetes (pictured).[95] Some of the free-living amoebae are important vectors of pathogenic bacteria or are pathogenic themselves (e.g., Acanthamoeba).[96] Others are anaerobic intestinal symbionts (e.g., Entamoeba).[11] | over 2,400[97] | |
| Breviatea | File:Undescribed breviates 2025 Aguilera-Campos-et-al.jpg | Anaerobic free-living amoeboflagellates with fine pseudopodia and modified mitochondria. Present only in low-oxygen marine and brackish sediments, their growth depends on mutualistic interactions with prokaryotes.[98] | 4[99] | ||
| Apusomonadida | File:Podomonas kaiyoae C.jpg | Free-living flagellates distinguished by a proboscis, a sleeve-like structure that envelops one of their two flagella.[99] Found gliding on wet soil and aquatic sediments worldwide.[100] | 28[100] | ||
| Opisthokonta** | File:PhysRevLett.116.038102-Fig1a.jpg | Flagellates distinguished by a single posterior flagellum, many with complex life cycles and varying degrees of multicellularity.[95] Some are entirely amoeboid, with fine pseudopodia (e.g., filastereans and nucleariids, including slime molds),[101][102] while others become amoeboid temporarily (e.g., choanoflagellates, pictured).[103] Most species are free-living filter-feeders or predators,[104][102][105][106] but some lineages (e.g., ichthyosporids) evolved into osmotrophic parasites of animals.[107][108] | approx. 300[76]** | ||
| Excavat. | Discoba | 
|
Flagellates with very different lifestyles, present in aquatic and terrestrial environments, ranging from aerobes to anaerobes. The most diverse group, Euglenozoa, includes free-living osmotrophs, phagotrophs, phototrophs (euglenophytes, pictured), and pathogens (kinetoplastids).[63][109] The less diverse Heterolobosea are primarily amoeboflagellates, and include some slime molds (acrasids) and well-known opportunistic parasites (e.g., Naegleria fowleri).[24] The smallest group, Jakobida, consume bacteria by suspension feeding.[110] | ||
| Metamonada | File:Giardia intestinalis - trophozoite.jpg | Anaerobic or microaerophilic flagellates, amoebae, or amoeboflagellates,[111] with reduced or completely lost[112] mitochondria. A few are free-living, found in aquatic hypoxic sediments, but most species are obligate parasites (e.g., Giardia, pictured) or commensals in animal intestines (e.g., parabasalids). Many have a high number of flagella.[113][111][11] | approx. 800[76] | ||
| Malawimonadida | 
|
Free-living bacterivorous flagellates that feed by suspension feeding, present in marine or fresh waters.[114] | 3[114] | ||
| Other | Ancyromonadida | Tiny free-living aquatic flagellates composed of flattened cells with an inflexible pellicle and a lateral rostrum with extrusomes. Found in most aquatic habitats.[115] | over 20[99][115] | ||
| CRuMs | 
|
Free-living flagellates and filose amoebae with a pellicle underneath the cell membrane. Almost all flagellated members can produce filose pseudopodia. Found in aquatic environments.[38] | 14[38] | ||
| *Excluding plants. **Excluding animals and fungi. | |||||
There are also many protists of uncertain position because their DNA has not been sequenced, and consequently their phylogenetic affinities are unknown.[11]
Predicted diversity

The total species diversity of protists is severely underestimated by traditional methods that differentiate species based on morphological characteristics. The number of catalogued protist species is very low (ranging from 26,000[116] to over 76,000)[lower-alpha 3] in comparison to the diversity of land plants, animals and fungi, which are historically and biologically well-known and studied. The predicted number of species also varies greatly, ranging from 140,000 to 1,600,000, and in several groups the number of predicted species is arbitrarily doubled. Most of these predictions are highly subjective. Molecular techniques such as environmental DNA barcoding have revealed a vast diversity of undescribed protists that accounts for the majority of eukaryotic sequences or operational taxonomic units (OTUs), dwarfing those from land plants, animals and fungi.[97] As such, it is considered that protists dominate eukaryotic diversity.[3]
Biology
In general, protists have typical eukaryotic cells that follow the same principles of biology described for those cells within the "higher" eukaryotes (animals, fungi and land plants).[117] However, many have evolved a variety of unique physiological adaptations that do not appear in the remaining eukaryotes,[118] and in fact protists encompass almost all of the broad spectrum of biological characteristics expected in eukaryotes.[3]
Nutrition
Protists display a wide variety of food preferences and feeding mechanisms.[11][119] According to the nutrient source, they can be divided into autotrophs (or phototrophs,[120] producers, traditionally algae), which photosynthesize their own organic molecules, and heterotrophs (consumers, traditionally protozoa), which obtain organic molecules from the environment, either by passive feeding of small particles (i.e., osmotrophs) or by engulfing whole cells or parts of cells of other organisms (phagotrophs).[121]
Phagotrophy

Phagotrophic protists feed by phagocytosis, a process unique to eukaryotes[122] where food particles or cells are digested into a vacuole, the phagosome.[119] This is the general mode of nutrition for protists, and has resulted in a diverse array of strategies for hunting and digestion.[122] Usually, digestion occurs at a specialized mouth-like region of the cell, the cytostome, which may be followed by the cytopharynx,[123] a tract supported by microtubules.[119] In amoebae, phagocytosis takes place anywhere on the cell surface.[124]
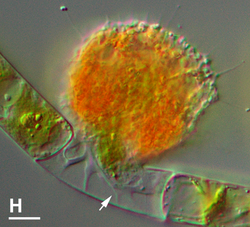
According to the method of digestion, protists can be divided into filter, raptorial, or diffusion feeders. Filter feeders accumulate small suspended particles into the cytostome by filtering them through pseudopodia or rigid tentacles, like choanoflagellates, or by generating water currents around the cytostome, like ciliates.[11] Raptorial feeders capture whole cells, either grazing on surfaces like bacterial lawns or actively preying on larger cells of other organisms.[124][11] Diffusion feeders, like suctorian ciliates and heliozoans, passively engulf prey that happen to collide with their tentacles or pseudopodia and are immobilized.[124] Certain protists exhibit a variation of predation known as myzocytosis, where they perforate the prey cell and suck out its contents or ingest them from the inside, leaving behind an empty shell; this is the case for vampyrellids, viridiraptorids, and many alveolates.[122][11]
Different predatory protists have sophisticated structures for capturing prey, such as the ventral groove of excavates, the hood-like extension or 'pallium' of some dinoflagellates, or the expandable oral pocket of ciliates. Many euglenids have a system of rods and vanes that grab and pull in prey cells, similarly to a Chinese finger trap.[122]
Preys of phagotrophic protists range from prokaryotes (i.e., bacterivores) to other eukaryotes, including single-celled protists, algae,[11] fungi,[125] nematodes,[126] or tissues of larger animals.[124] Prey specificity varies, with some groups specialized in eating only one type of organisms,[125] or only a particular strain.[127] Traditionally, protists were considered primarily bacterivorous due to biases in cultivation techniques, but most are omnivores.[125]
Osmotrophy
Osmotrophic protists absorb soluble[121] or very small (under 0.5 μm) molecules by diffusion, membrane channels and carriers, or pinocytosis, where nutrients are engulfed into small vacuoles or endosomes.[11][119] Some osmotrophs, known as saprotrophs or lysotrophs, perform external digestion by releasing digestive enzymes into the environment and decomposing organic matter[11] into simpler molecules that can be absorbed, allowing finer control over substances that enter the cell and minimizing the risk of harmful substances or infection.[128] Probably all eukaryotes are capable of osmotrophy, but some have no alternative of acquiring nutrients. Obligate osmotrophs include the aphagean euglenids, some green algae, the human parasite Blastocystis, some metamonads,[11] the parasitic trypanosomatids,[129] and the fungus-like oomycetes and hyphochytrids.[128]
Mixotrophy
Most photosynthetic protists are mixotrophs,[130] as they combine photosynthesis with phagocytosis.[lower-alpha 4] While some mixotrophs already have chloroplasts (i.e., algae), others acquire chloroplasts by stealing them from their prey, a process known as kleptoplasty. Kleptoplastic protists may be generalists, able to steal chloroplasts from a variety of prey, like some ciliates, or they may be specialists, only capable of obtaining chloroplasts from very specific prey. Specialists may keep the entire prey inside of their cells, as do many foraminifers and radiolarians, or they may only engulf the plastids and discard the rest.[132][133]
Among exclusively heterotrophic protists, variation of nutritional modes is also observed. The diplonemids, which inhabit deep waters where photosynthesis is absent, can flexibly switch between osmotrophy and bacterivory depending on the environmental conditions.[134]
Homeostasis

Many freshwater protists need to osmoregulate (i.e., remove excess water volume to adjust the ion concentrations) because non-saline water enters in excess from the environment.[135][123] Osmoregulation is done through ion transporters of the cell membrane and through contractile vacuoles, specialized organelles unique to protists that periodically excrete fluid high in potassium and sodium through a cycle of contractions.[118] These vacuoles are surrounded by the spongiome, a system of vesicles or tubes that slowly collect fluid from the cytoplasm into the vacuoles, which then contract and discharge the fluid through a pore. The mechanism, location, and structure of this system vary across protists. For example, ciliates contract the vacuoles by actin and microtubule filaments; dinoflagellates contract it through a sheath formed by a flagellar rootlet, known as the pusule. Marine, parasitic, or thick-walled protists lack these vacuoles.[135]
Certain protists have acidic organelles known as acidocalcisomes, which store high concentrations of phosphorus, calcium, and enzymes related to their metabolism. Among their proposed functions are osmoregulation and maintenance of pH and calcium homeostasis.[136]
Mitochondria and respiration
The last eukaryotic common ancestor was aerobic, bearing mitochondria that synthesize ATP through oxidative respiration, which requires oxygen. Most protists are aerobes, but many lineages of free-living and parasitic protists have independently adapted to inhabit anaerobic or microaerophilic (low-oxygen) habitats by modifying their mitochondria into organelles collectively known as mitochondrion-related organelles (MROs). These exist in a continuum from lower to higher degrees of reduction. For example, hydrogenosomes have lost the electron transport chain used in respiration, as well as other features of classical mitochondria (their DNA, the Krebs cycle, etc.), but can still generate ATP anaerobically through the fermentation of pyruvate, releasing hydrogen gas as a byproduct. Mitosomes have lost both the respiratory chain and the production of ATP. One group of protists, the genus Monocercomonoides, has lost its mitochondria entirely.[118][137][138] In a similar manner, the oxidative peroxisome evolved into a fermentative glycosome in trypanosomatids.[118]
Besides metabolic reduction, the shape, composition, and number of mitochondria varies greatly across protists. Apicomplexans and kinetoplastids have a single large mitochondrion that divides synchronously with the cell, while some amoebae can present hundreds of mitochondria.[139] Mitochondrial genomes (mitogenomes), typically composed of one circular chromosome, can appear as numerous linear chromosomes in many unrelated protists, such as Amoebidium, with hundreds of chromosomes.[140] The large mitogenome of kinetoplastids is condensed into a kinetoplast, which is physically tied to the flagellar apparatus. The smallest known mitogenome belongs to the symbiotic alga Chromera velia.[141]
Mitochondrial cristae, foldings of the inner membrane, have been used to classify protists since the advent of electron microscopy.[47] Flat cristae are the ancestral trait, tubular cristae are present in the SAR supergroup and Amoebozoa, and discoid cristae distinguish the Discoba.[142]
Cytoskeleton
The cytoskeleton of protists generally consists of an array of microtubules and other fibers that radiate from a complex flagellar apparatus. This structure—sometimes known as the mastigont[143]—was present in the ancestor of all eukaryotes, and is fundamental to the structure, movement and division of cells. It is one of the only cellular features that can be compared across all protists, as it is relatively conserved.[144][145]
The interior of the flagellum, the axoneme, consists of a common structure of nine pairs of microtubules surrounding two central microtubules, known as the 9+2 structure. At its base is a basal body or kinetosome, a complex proteic structure that forms the centrioles and behaves as the microtubule organizing center. Microtubules emerge from each basal body in the form of one or two 'roots'.[146] The basic plan of the flagellar apparatus consists of two basal bodies (B1 and B2), one for each flagellum, followed by four primary microtubular 'roots' (named R1 through R4) and a 'singlet root' (SR) formed by a single microtubule and originating from B1. Attached to the R1 is a multilayered structure, also known as C fiber.[145]
Each protist group has modifications or secondary losses of this standard organization.[144] In groups where the standard structure is mostly untouched (e.g., excavates, stramenopiles, and apusomonads), the R1, R2 and SR roots provide reinforcement for the ventral feeding groove, and the R3 supports the dorsal side of the cell.[145] In opisthokonts, one flagellum and all the microtubular roots were lost, but both basal bodies remain.[145][147] In archaeplastids, the SR and R2 supporting the feeding groove were lost, likely due to their shift to autotrophic nutrition.[145] In certain protists the flagellar apparatus is physically linked to the nucleus, forming what is known as the karyomastigont.[148] This connection is often done through different kinds of filamentous structures, variously called rhizoplasts or internal flagellar roots.[149]
Sensory perception

Many flagellates and probably all motile algae exhibit a positive phototaxis (i.e. they swim or glide toward a source of light). For this purpose, they exhibit photoreceptors of varying degrees of complexity, from simple receptors with light antennae (as in the eyespot apparatus of many algae), to receptors with opaque screens, to complex ocelloids with intracellular lenses (as in the dinoflagellate family Warnowiaceae).[118][150] Some ciliates orient themselves in relation to the Earth's gravitational field while moving (geotaxis), and others swim in relation to the concentration of dissolved oxygen in the water.[118]
Symbionts
Protist cells have an accentuated tendency to harbor mutualistic symbionts, which have produced new physiological opportunities. Some associations are permanent, others more transient.[118] Many protists maintain cyanobacteria or other algae as endosymbionts, to benefit from their photosynthesis—especially in radiolarians, foraminifers, and other planktonic marine protists[151]—or their nitrogen fixation.[152] Others maintain only the chloroplasts of algae they ingest, and dispose of the remaining cellular structures (i.e., kleptoplasty).[127] Two species of ciliates have been observed hosting purple bacteria, which perform photosynthesis without oxygen.[153][154]
Several groups of protists host non-photosynthetic prokaryotes, often maintaining an anaerobic lifestyle through the metabolism of their symbionts. Xenosomes are bacterial endosymbionts with a methanogenic role, found in anaerobic ciliates.[118] Symbiontid euglenozoans and select ciliates have sulfur-oxidizing bacteria living as epibionts on their surfaces.[155][156][157] Similarly, breviates have hydrogen-oxidizing epibiotic bacteria.[158] Metamonads, particularly parabasalids and oxymonads found in the hindgut of termites, typically host methanogenic archaea as epi- or endobionts.[159]
Some rare associations involve prokaryotes that defend the protist host against potential predators, namely in symbiontids and in the ciliate Euplotidium, where the epibionts are verrucomicrobia that eject genetic material as a defense mechanism.[160][155] There are also some species of oxymonads whose epibionts function as chemosensors, providing their host with information on the surrounding chemical gradient.[161]
Besides algae, occurrence of mutualistic eukaryotic symbionts is rare among protists. In the genus Neoparamoeba, some species have endosymbionts that resemble Perkinsela amoebae, a species of trypanosomatids. Although no benefits are yet known from this association, their evolution matches almost perfectly, suggesting that the symbionts are inherited.[162]
Life cycle and reproduction
Protists exhibit a large variability of life cycles and strategies involving multiple stages of different morphologies which have allowed them to thrive in most environments. Nevertheless, most research concerning protist life cycles corresponds to model organisms and important parasites; knowledge on the life cycles of the free-living majority remains fragmentary.[163]
Asexual reproduction
Protists typically reproduce asexually under favorable environmental conditions,[164] allowing for rapid exponential population growth with minimal genetic variation. This occurs through mitosis and has historically been considered the main reproductive mode in protists.[163] Unicellular protists often multiply via binary fission, like bacteria;[163] they can also divide through budding, similarly to yeasts, or through multiple fissions, a process known as schizogony.[165] In multicellular protists, this process is often known as vegetative reproduction, only performed by the 'vegetative stage' or individual. It can take the form of fragmentation of body parts, or specialized propagules composed of numerous cells (e.g., in red algae).[166]
Sexual reproduction
Sexual reproduction is a fundamental characteristic of eukaryotes.[167][168] It involves meiosis (a specialized nuclear division enabling genetic recombination) and syngamy (the fusion of nuclei from two parents),[163] two processes thought to have been present in the last eukaryotic common ancestor,[169] which likely had the ability to reproduce sexually on a facultative (non-obligate) basis.[170] Even protists that no longer reproduce sexually still retain a core set of meiosis-related genes, reflecting their descent from sexual ancestors.[171][172] For example, although amoebae are traditionally considered asexual organisms, most asexual amoebae likely arose recently and independently from sexually reproducing amoeboid ancestors.[173] Even in the early 20th century, some researchers interpreted phenomena related to chromidia (chromatin granules free in the cytoplasm) in amoebae as sexual reproduction.[174] Three distinguishable sexual cycles are observed in protists depending on the ploidy[lower-alpha 5] of the individual or vegetative stage:[163]
- Haploid cycle (as in most fungi): the individual is haploid and differentiates through mitosis into haploid gametes, which fuse into a zygote that immediately undergoes meiosis to generate new haploid individuals.[163] This is the case for some green algae (such as volvocine algae), most dinoflagellates, dictyostelids, some metamonads, and apicomplexans.[123]: 26 [175] Zygnematophytes, a group of green algae, fuse vegetative cells directly by conjugation instead of producing gametes.[176]

- Diploid cycle (as in animals): the individual is diploid and undergoes meiosis to generate haploid gametes, which fuse into a zygote that develops as a new individual.[163] This is the case for some metamonads, heliozoans, many green algae, diatoms, and labyrinthulids.[123]: 26 [175] Ciliates are also diploid, but instead of producing gametes they divide their micronucleus into two haploid nuclei, exchange one of them by conjugation with another ciliate, and fuse the two nuclei into a new diploid nucleus.[177]
- Haplo-diploid cycle (as in land plants): there are two alternating generations of individuals. One, the diploid agamont (or sporophyte), undergoes meiosis to generate haploid cells (called spores) that develop into the other generation, the haploid gamont (or gametophyte). The gamont then generates gametes by mitosis, which fuse to form the diploid zygote that develops into the agamont.[163] This is the case for many foraminifera and many algae.[123]: 26 Depending on the relative growth and lifespan of one generation compared to the other, life cycles may be haploid-dominant, diploid-dominant, or with equally dominant generations. Brown algae exhibit the full range of these modes.[178] Many red algae have a three-generational cycle with a carposporophyte, whose spores germinate into a tetrasporophyte, whose spores develop into the gametophyte.[179]
Factors inducing sexual cycles
Free-living protists tend to reproduce sexually under stressful conditions, such as starvation or heat shock. Oxidative stress, which leads to DNA damage, also appears to be an important factor in the induction of sex in protists.[164]
Several protists synchronize their life cycles (namely the formation or release of gametes) according to environmental factors such as nutrient or light levels, resulting in synchronization with the day-night cycle, the lunar cycle, or the seasons. The malaria agent Plasmodium falciparum synchronizes its life cycle with the host's levels of melatonin.[180]
Cycles in pathogenic protists
Pathogenic protists tend to have extremely complex life cycles that involve multiple forms of the organism, some of which reproduce sexually and others asexually.[181] The stages that feed and multiply inside the host are generally known as trophozoites (from gre trophos 'nutrition', and zoia 'animals'), but the names of each stage vary depending on the protist group[165] (e.g., sporozoites and merozoites in apicomplexans;[182][168] primary and secondary zoospores in phytomyxeans).[183]
Some pathogenic protists undergo asexual reproduction in a wide variety of organisms – which act as secondary or intermediate hosts – but can undergo sexual reproduction only in the primary or definitive host (e.g., Toxoplasma gondii in felids such as domestic cats).[184] Others, such as Leishmania, are capable of performing syngamy in the secondary vector.[185] In apicomplexans, sexual reproduction is obligatory for parasite transmission.[186]
Despite undergoing sexual reproduction, it is unclear how frequently there is genetic exchange between different strains of pathogenic protists, as most populations may be clonal lines that rarely exchange genes with other members of their species.[187]
Habitats
Protist diversity, as detected through environmental DNA surveys, is vast in every sampled environment, but it is mostly undescribed.[188] The richest protist communities appear in soils, followed by oceanic and lastly freshwater habitats, mostly as part of the plankton.[189]
Soil-dwelling protist communities are ecologically the richest, possibly due to the complex and highly dynamic distribution of water in the sediment, which creates extremely heterogenous environmental conditions. The constantly changing environment promotes the activity of only one part of the community at a time, while the rest remains inactive; this phenomenon promotes high microbial diversity in prokaryotes as well as protists.[189] Cercozoan amoeboflagellates like the glissomonads and cercomonads are among the most abundant soil protists, their morphological variability well suited for foraging between soil particles. Testate amoebae are also acclimated to the soil environment, as their shells protect against desiccation. Most soil algae are stramenopiles and green algae. Fungus-like protists and slime molds (e.g., oomycetes, myxomycetes, acrasids) are present abundantly as saprotrophs.[190] The major parasites in land are the animal-associated apicomplexans[191] and the plant-associated oomycetes and plasmodiophorids.[190]
Marine protists are present in almost the entire range of oceanic conditions, mostly dominating the photic zone. Their abundance depends mostly on the availability of inorganic nutrients, rather than temperature or sunlight, and may vary seasonally.[192] They are most abundant in coastal waters that receive nutrient-rich run-off from land, and areas where nutrient-rich deep ocean water reaches the surface, namely the upwelling zones in the Arctic Ocean and along continental margins.[193] Radiolarians are widespread as the most dominant marine consumers.[133][132][189] Macroalgae (namely red algae, green algae and brown algae), unlike plankton, generally require a fixation point, which limits their marine distribution to coastal waters, and particularly to rocky substrates.[194] Some communities of seaweeds exist adrift on the ocean surface, serving as a refuge and means of dispersal for associated organisms.[195] Parasitoids such as Syndiniales are abundant pathogens in oceans.[189]
Freshwater protist communities are characterized by a higher beta diversity (highly heterogeneous between samples) than soil and marine plankton. The high diversity can be a result of the hydrological dynamic of recruiting organisms from different habitats through extreme floods.[196] In freshwater phytoplankton, golden algae, cryptophytes and dinoflagellates are the most abundant groups.[189]
Extreme habitats
b) Chlamydomonas pitschmanii, a green alga from hot spring soils.
c) Tetramitus thermacidophilus, an amoeboflagellate from an acidic geothermal lake in California.
d) Galdieria sulphuraria, a thermoacidophilic red alga.
e) Halocafeteria seosinensis, a flagellate from a saltern in Korea.
Protists can survive a broad range of extreme conditions, including extreme temperatures (thermophiles or psychrophiles), salinity (halophiles), and pH (alkaliphiles or acidophiles). Most of the extremophilic eukaryotes are algae, specifically chlorophytes, followed by fungi. Other extremophile-abundant groups are heterolobose amoebae, red algae, stramenopiles, and ciliates.[197]
Eukaryotic algae are well-known to withstand high temperatures; for example, the red alga Cyanidioschyzon merolae persists up to 60°C, similarly to the most extreme thermophilic fungi. Lesser-known thermophilic amoebae and amoeboflagellates (e.g., Echinamoeba thermarum) are repeatedly found in hot environments, including artificially heated systems. While less successful than algae or amoebae, ciliates have also been found in hydrothermal vents up to 52°C. This is still lower than prokaryotes, some of which grow above 80°C.[197] In extremely cold habitats, like snow and the Arctic Ocean, diatoms and green algae are the dominant phototrophs.[198]
In terms of pH and salinity, protists can withstand similar extremes relative to prokaryotes and fungi, and also persist in polyextreme environments (polyextremophiles). The record for acidophily is also C. merolae, with an observed minimum growth of pH 0. Besides red algae, some species of green algae and amoeboflagellates are found in high-temperature, low-pH geothermal springs. Alkaliphilic protists, primarily represented by ciliates, resist up to pH 10.48, higher than the most alkalophilic bacterium.[197]
Protists are remarkably successful in extreme salinity due to their salt-out strategy, which consists of accumulating organic solutes in the cell instead of ions to counterbalance the hypertonic environment. Examples include the alga Dunaliella salina and the flagellate Halocafeteria seosinensis, which is able to tolerate up to 36.3% salinity, higher than the maximum reported in bacteria (35%) and fungi (30%).[197]
Biomass comparison
Protists are abundant in nearly all habitats. They contribute an estimated 4 gigatons (Gt) to Earth's biomass—double that of animals (2 Gt), but less than 1% of the total. Combined, protists, animals, archaea (7 Gt), and fungi (12 Gt) make up less than 10% of global biomass, with plants (450 Gt) and bacteria (70 Gt) dominating.[199]
Ecological roles
Protists are indispensable to modern ecosystems worldwide. They also have been the only eukaryotic component of all ecosystems for much of Earth's history, which allowed them to evolve a vast functional diversity that explains their critical ecological significance. They are essential as primary producers, as intermediates in multiple trophic levels, as key regulating parasites or parasitoids, and as partners in diverse symbioses.[3]
Primary producers
Phototrophic protists are the main contributors to the biomass and primary production in nearly all aquatic environments, as part of the phytoplankton.[lower-alpha 6] Altogether, they are responsible for almost half of the global primary production.[193] They are the main providers of much of the energy and organic matter used by other trophic levels, including essential nutrients such as fatty acids.[200] Macroalgae support numerous herbivorous animals, especially benthic ones, as both food and refuge from predators.[194][195]
Consumers
Phagotrophic protists are the most diverse functional group in all ecosystems.[189] In the trophic webs of soils, protists are the main consumers of both bacteria and fungi, the two main pathways of nutrient flow towards higher trophic levels. As bacterial grazers, they have a significant role in the foodweb: they excrete nitrogen in the form of NH3, making it available to plants and other microbes.[201]
As part of the plant-associated microbiomes (rhizosphere near the roots, phyllosphere on the leaves), predatory protists such as cercomonads regulate the populations of bacteria and fungi, indirectly improving plant health and growth. They can also have a more direct impact by releasing proteins with antimicrobial activity.[202]
Decomposers
Necrophagy (the degradation of dead biomass) among microbes is mainly attributed to bacteria and fungi, but protists have a still poorly recognized role as decomposers with specialized lytic enzymes.[203] In marine and estuarine environments, the well-studied thraustochytrids (a type of labyrinthulomycetes) are relevant saprotrophs that decompose various substrates, including dead plant and animal tissue. Various ciliates and testate amoebae scavenge on dead animals. Some nucleariid amoebae specifically consume the contents of dead or damaged cells, but not healthy cells. However, all these examples are of facultative necrophages that also feed on live prey. In contrast, the cercozoan family Viridiraptoridae, present in shallow bog waters, are obligate necrophages of dead algae, potentially fulfilling an important role in cleaning up the environment and releasing nutrients for other microbes.[203]
Parasites and pathogens
Parasitic protists compose around 15–20% of all environmental DNA samples in marine and soil ecosystems, but only around 5% in freshwater systems, where chytrid fungi likely fill that ecological niche.[189] They are significant parasites of animals, land plants, fungi,[204] and even of other protists.[205] In plants, oomycetes are the most economically important pathogens (e.g., potato blight),[206] but other lesser studied lineages are known to infect plants, such as phytomyxids (e.g., clubroot), labyrinthulids, and trypanosomatids of the genus Phytomonas.[207]
Parasitic protists are among the most well-known human pathogens, causing diseases such as malaria, toxoplasmosis, amoebic meningoencephalitis, sleeping sickness, leishmaniasis, and several diarrheal illnesses like amoebiasis, cryptosporidiosis, and giardiasis.[208] Several amoebae are amphizoic, normally free-living but capable of infection.[209]
While parasitic protists are largely studied as protozoa,[210] some are algae, such as the green alga Cephaleuros virescens which infects plant leaves.[211] Hundreds of red algae species parasitize on other red algae, usually closely related species.[212]
Certain non-parasitic protists can still be toxic to aquatic animals during periods of excessive growth, either by the release of potent toxins, the depletion of oxygen in the water, or mechanical damage to gills from piercing structures (like the skeletons of silicoflagellates).[213] These phenomena are known as harmful algal blooms, sometimes causing water discoloration (red tides). The most common agents are diatoms and dinoflagellates. When toxins are involved, they can reach human consumption, leading to fish or shellfish poisoning like ciguatera.[214]
Mutualists and commensals
Many protists live as endosymbionts in a non-parasitic association, providing their hosts with nutritional advantages. Microalgae, namely zoochlorellae (green algae) and zooxanthellae (dinoflagellates, haptophytes and ochrophytes), are widespread endobionts of other protists, especially foraminifers, radiolarians and ciliates,[215] but also of animals. The association of zooxanthellae with corals is extensively studied and valued for its importance in reef ecosystems.[216][217] Climate warming leads to the loss of zooxanthellae, which manifests as coral bleaching and death of the coral hosts.[218][219] Some molluscs like the giant clams also harbor zooxanthellae.[220] Sloths are the only mammals with green algae as epibionts on their fur; the hypothesized benefits range from camouflage to sunscreen-like protection or nutritional supplements.[221]
Heterotrophic protists are prevalent members of the gut microbiome of animals, although research has focused almost exclusively on gut bacteria.[222] The giant metamonad flagellates found in the hindgut of termites and cockroaches allow them to digest wood. This is an obligate mutualism, as termites will starve if cleaned of these protists.[223] However most gut protists are commensals, such as the ciliates abundantly present in the rumen of ruminants, or the ciliate-like opalinids that inhabit amphibian and reptile guts.[224]
Biogeochemical cycles
Marine protists have a fundamental impact on biogeochemical cycles, particularly the carbon cycle.[225] As phytoplankton, they fix as much carbon as all terrestrial plants combined.[189] Shells of biomineralizing marine protists accumulate as sediment that forms various geological features, such as chalk deposits, made of coccoliths,[226] or sand stars, the remains of star-shaped foraminifers.[75] Soil protists, particularly testate amoebae, contribute to the silica cycle as much as forest trees through the biomineralization of their shells.[190]
Evolution and fossil record
Prior to the existence of animals, land plants and fungi, all eukaryotes were protists. As such, questions regarding their origin and evolution are questions on protists. Because of the scarce fossil record prior to ca. 750 Mya, dating of early events in eukaryotic evolution relies primarily on molecular clock reconstructions.[227][228] Eukaryotic cells first evolved from archaea[229][230] some time during the late Archean eon (ca. 3 billion years ago),[231] forming a lineage that eventually gave rise to the last eukaryotic common ancestor (LECA) with the traits associated with crown-group (modern) eukaryotes, namely mitochondria and a complex endomembrane system.[232] Estimations on the date of LECA range from 2.4 to 1.6 billion years ago, during the Paleoproterozoic,[233] leaving a considerably long gap of stem-group (extinct) eukaryotes with intermediate traits.[232][231][234]
Template:Protist fossil record
First fossils and stem eukaryotes
Fossils before 1 billion years ago are limited and cannot be confidently assigned to modern eukaryotic groups. As such, they are interpreted as potential stem-group eukaryotes, based on their large cell sizes and complex shapes that would still require diagnostic eukaryotic features such as a cytoskeleton and an endomembrane system.[232][234][235] The earliest potential stem eukaryotes reach back to the late Paleoproterozoic (2–1.6 billion years ago).[227] Examples of these are Leiosphaeridia, Tappania, Shuiyousphaeridium, and Grypania.[234][236] There are also two fossils of putative red algae, Ramathallus and Rafatazmia.[233]
Sterols, common to modern eukaryotic membranes (e.g., cholesterol), appear comparatively late in the fossil record in comparison to the molecular dating of LECA, suggesting that crown-group eukaryotes flourished relatively late. Instead, stem eukaryotes may have produced simpler protosterols that require less oxygen during biosynthesis. The advanced sterols of modern eukaryotes, although metabolically expensive, likely provided numerous advantages through increased membrane flexibility, such as resilience to osmotic shock during dessication-rehydration cycles, extreme temperatures, oxidative damage, and UV light exposure, allowing them to colonize diverse and harsh environments (e.g., mudflats, rivers, agitated shorelines and land). In contrast, stem eukaryotes remained in low-oxygen marine waters, although at higher abundances.[234]
Radiation of crown eukaryotes
In absence of a reliable fossil record, hypotheses regarding the early evolution of crown eukaryotes are based on the phylogenomics of modern protists coupled with morphological and ecological comparisons.[228] LECA is widely hypothesized as having a typical excavate morphology, with two flagella and a ventral groove used for feeding on bacteria by phagocytosis.[62][64] In addition, LECA was likely a facultative anaerobe which could live in both aerobic and, more commonly during the Proterozoic, anaerobic environments.[237][238]
Following LECA, a series of ecological and evolutionary innovations took place in the span of 300 million years, between the Paleoproterozoic and Mesoproterozoic, resulting in a rapid radiation that originated all major eukaryotic supergroups.[239][235][228] These events can be outlined in four broad stages.[lower-alpha 7] Firstly, the direct descendants of LECA, excavate-like flagellates feeding on bacteria by altering water currents, diversify and give rise to the most basal groups such as the anaerobic metamonads and the aerobic jakobids. Secondly, surface-associated protists emerge with the development of gliding motility and the cell plasticity to form thin pseudopodia, resulting in lineages such as ancyromonads, apusomonads and CRuMs (i.e., the podiates), which pick bacteria directly off of surfaces. The third stage involves the lineage Diaphoretickes, in which agile swimming motility was prioritized and several adaptations led to an active predatory lifestyle (e.g., provorans, telonemids) hunting both bacteria and, for the first time, other eukaryotes. In response, some prey might have developed stronger pellicles, as seen in rigifilids and apusomonads. Lastly, the fourth stage saw the appearance of both "super-predators" capable of hunting other eukaryovores (as in the SAR lineages), and more specialized bacterivores with amoeboid movement (as in amoebozoans and opisthokonts), resulting in a "mature" biosphere.[228] Early amoebozoans of the Mesoproterozoic adapted to grazing on microbial mats, the dominant food source at the time, resulting in multiple losses of flagella and large amoeboid bodies.[240]
Below is a consensus phylogenetic tree of eukaryotes, including all major supergroups that originated from this time period.[228][61] The position of the root is still debated, hence the polytomy.[lower-alpha 8] Excavate groups are marked *.
| LECA |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Neoproterozoic expansion
The presence of modern eukaryotes in the fossil record remained modest[lower-alpha 10] until the Tonian period (1000–720 million years ago), when biomarker molecules and microfossils became abundant.[232] Fossils at around 1000 million years ago (Mya) include the oldest specimens that are assigned with high confidence to crown eukaryotes.[227][236] Bangiomorpha and Proterocladus are the oldest multicellular red and green algae, respectively.[245] The oldest fossils of opisthokonts are Ourasphaira giraldae, interpreted as the earliest fungus,[234] and Bicellum brasieri, the earliest holozoan, showing traits associated with animal-like multicellularity such as different cell types.[246] Heterotrophic protists appear abundantly throughout the Tonian, as exemplified by the vase-shaped microfossils (780–720 Mya), interpreted as marine testate amoebae[247] and as such composing the oldest representatives of filose (Cercozoa) and lobose (Amoebozoa) amoebae.[248][249] Other microfossils include some poorly preserved tubular shells from 716–635 Mya rocks.[250]
As oxygen levels rose, crown eukaryotes may have outcompeted stem eukaryotes, expanding into oxygen-rich marine environments that supported an aerobic metabolism enabled by their mitochondria. Stem eukaryotes may have gone extinct due to competition and the extreme climatic changes of the Cryogenian glaciations (720–635 Mya) and subsequent global warming, cementing the dominance of crown eukaryotes which began to appear abundantly in this era, fueled by the proliferation of algae.[245][234] After the Gaskiers glaciation of the Late Ediacaran (ca. 579 Mya), fossils of heterotrophic protists diversify further. Some fossils at 548 Mya, similar to vase-shaped microfossils, are interpreted as the oldest traces of foraminifers (e.g., Protolagena),[247] but their foraminiferal affinity is doubtful.[250]
Phanerozoic diversifications and major events
At the very start of the Paleozoic era, the first definitive fossils of radiolarian[251][252][253] and foraminiferal[254][250] shells are found, alongside the first small shelly fauna.[255] Following the Cambrian explosion of animals, the Precambrian microbe-dominated ecosystems were replaced by primarily benthic and nekto-benthic communities, with most marine organisms limited to the depths of shallow water environments.[256] Mirroring the animal radiation, there was a radiation of phytoplanktonic protists (acritarchs)[257] around 520–510 Ma, followed by a decrease in diversity around 500 Ma.[258] The surviving acritarchs expanded in diversity and morphological innovation[257] due to a decrease in predation from benthic animals, which suffered extinction due to various proposed environmental factors such as anoxia.[259] Both phytoplankton and zooplankton (e.g., radiolarians) flourished, as signaled by an increase of organic carbon buried in the sediment known as the SPICE event (~497 Mya).[256][259] This abundant biomass supported a second animal radiation known as the Great Ordovician Biodiversification Event. This period is also known as the 'Ordovician Plankton Revolution' due to the significant diversification of planktonic protists.[256] Starting in the middle Ordovician, the earliest fossils of eugelnids (Moyeria) appear.[260]
In the Devonian period, the first fossils of freshwater arcellinid testate amoebae are found (e.g., Palaeoleptochlamys, Cangweulla),[261] as well as various types of freshwater green algae, including charophytes, volvocaceans and desmids,[262] and some fossils that might represent glaucophytes.[263] Some benthic foraminifera acquired the ability of calcifying,[264] and particularly the giant fusulinids became the dominant fossilizable protists. This period also includes the molecular origin of haptophytes (~310 Mya) and silicoflagellates (397–382 Mya), which did not leave fossil traces until later in the Mesozoic. After the Late Devonian extinction (372 Mya), nassellarian-like radiolarians appeared for the first time, with a unique body plan among marine protists.[264]
During the Carboniferous period, no new fossilizable protists originated despite the major environmental changes. However, radiolarian diversity and productivity increased, causing the accumulation of large amounts of biosiliceous sediment (chert) worldwide until the Early Cretaceous.[265][266][267] Around the Capitanian mass extinction event (262–259 Mya) of the Permian period, coccolithophores genetically diverged from the rest of haptophytes, possibly as a response to a reduction in atmospheric oxygen, and there was a faunal turnover from larger to smaller fusulinids.[264] Spumellarian radiolarians appear in the latest Permian.[265]
The Permian-Triassic extinction event (~251.9 Mya) caused the extinction of many radiolarians, which manifests as a gap in the chert record. The Triassic period saw the acceleration of radiolarian diversity[265] and the appearance of several groups of calcaerous nannofossils, including dinocysts, the oldest identifiable coccolithophore Crucirhabdus minutus, and the oldest fossils of Phaeodaria.[264] There's a variety of protozoa, including soft-bodied ciliates, and filamentous algae found in amber from the Late Triassic (220–230 Ma).[268]
Around the Early–Middle Jurassic, after the global Toarcian Oceanic Anoxic Event there was a diversification of dinoflagellates and coccolithophores, in both species and abundance. This interval also saw the completion of a symbiosis between Acantharia radiolarians and lineages of Phaeocystis haptophytes, as well as the appearance of planktonic foraminifera.[264] The period of low atmospheric oxygen ends in the Aptian-Albian boundary during the Early Cretaceous, and the first fossils of diatoms and silicoflagellates appear.[264] Samples of amber from around 100 Ma contain the oldest fossil records of apicomplexans (particularly malarian agents and gregarines), trypanosomes,[269] and metamonads—particularly mutualistic parabasalids of cockroaches, representing the earliest record of mutualism between protists and animals.[270][271]
Across the Mesozoic era, coccolithophores, dinoflagellates and later diatoms became the dominating eukaryotic producers in oceans until today, as opposed to cyanobacteria and green algae which dominated earlier. Their diversification caused an accelerated transfer of primary production into higher trophic levels, which in turn caused the animal "Mesozoic marine revolution", characterized by the appearance of widespread predation among most invertebrate phyla.[272]
The Cenozoic era began with another extinction event (~66 Ma) that caused the replacement of mesozoic forms of dinoflagellates, foraminifers, coccolithophores, and silicoflagellates with forms that dominate marine habitats today. Right after this event, putative ebridians begin appearing in the fossil record (e.g., Ammodochium), but the oldest reliable ebridian fossils belong to the upper middle Eocene (42–33.7 Ma).[264] Around this time, the oldest fossils of synurids appear (~49–40 Ma).[273] Following the Middle Eocene Climatic Optimum (~40 Ma), diatoms became the dominant agents of marine silicon precipitation as opposed to radiolarians, and the fossil record shows the first raphid diatoms and collodarians.[264]
See also
- Protistology
- Glossary of protistology
- Protist locomotion
Footnotes
- ↑ 1.0 1.1 Eukaryotic flagella are interchangeable with 'cilia' from a biological perspective. The usage of these two names depends on the author: some prefer to reserve cilia for shorter appendages and flagella for longer ones, while others prefer cilia for eukaryotes and flagella for prokaryotes. The term 'undulipodium' was proposed to unify the two concepts, as it refers specifically to the homologous microtubular structure found in both, but not found in prokaryotic flagella.[18][19][20]
- ↑ There was, however, one kingdom-based system that persisted into the 21st century, developed by Thomas Cavalier-Smith. He proposed two non-monophyletic kingdoms of protists, the Protozoa and the Chromista. He argued that protists with red algal-derived plastids and their heterotrophic relatives (i.e., Stramenopiles, Alveolata, Haptista and Cryptista) shared a single common photosynthetic ancestor, and composed the Chromalveolata or, later with the addition of Rhizaria, the Chromista,[10][52] which was polyphyletic.[53] This scheme endured until 2022, the year of his last publication.[54]
- ↑ A 2007 report on protist diversity included a table listing the described number of species for protist and fungal groups. The total sum of the listed species, excluding fungi, is 76,144.[76]
- ↑ The terms "mixotroph" and "mixoplankton" almost exclusively refer to protists that perform photosynthesis and phagocytosis (photo-phagotrophs). Osmotrophy is always present, but not taken into account. As such, "pure" phototrophs (incapable of phagocytosis) and "pure" phagotrophs (incapable of photosynthesis) are technically mixotrophic due to their innate ability for osmotrophy, but are not usually reported in this sense.[131]
- ↑ 5.0 5.1 Every sexual cycle involves syngamy and meiosis, which increase or decrease the ploidy (i.e., number of chromosome sets, represented by the letter n), respectively. Syngamy is the fusion of two haploid (1n) reproductive cells, known as gametes, into a diploid (2n) cell called zygote, which then undergoes meiosis to generate haploid cells.[163]
- ↑ Given the importance of aquatic algae, the equally abundant soil algae may provide a larger contribution to the global carbon cycle than previously thought, but the magnitude of their carbon fixation has yet to be quantified.[189]
- ↑ The exact temporal relationship between these proposed stages is currently impossible to determine. They merely represent the sequence of ecological and evolutionary innovations that took place between LECA and modern eukaryotes. The second and third stages might have overlapped.[228]
- ↑ Some analyses place the root of the eukaryote tree within or next to Metamonada,[238][228] while others split the tree between a branch leading to Diaphoretickes and Discoba ('Diphoda') and a branch leading to Podiata ('Opimoda').[241]
- ↑ Telonemia may be more closely related to SAR, forming the hypothesized TSAR clade,[242] or it may branch with Haptista instead.[243][244]
- ↑ There is a considerable fossil gap between the first stem eukaryotes of the late Paleoproterozoic and early Mesoproterozoic (~1.6 billion years ago) and the first crown eukaryotes of the Neoproterozoic (~1 billion years ago). This gap is known as the 'boring billion'.[236]
References
- ↑ 1.0 1.1 1.2 "The other eukaryotes in light of evolutionary protistology". Biology & Philosophy 28 (2): 299–330. 2012. doi:10.1007/s10539-012-9354-y.
- ↑ Sebé-Pedrós, Arnau; Degnan, Bernard M.; Ruiz-Trillo, Iñaki (8 May 2017). "The origin of Metazoa: a unicellular perspective". Nature Reviews Genetics 18 (8): 498–512. doi:10.1038/nrg.2017.21. PMID 28479598.
- ↑ 3.0 3.1 3.2 3.3 Burki, Fabien; Sandin, Miguel M.; Jamy, Mahwash (2021). "Diversity and ecology of protists revealed by metabarcoding". Current Biology 31 (19): R1267–R1280. doi:10.1016/j.cub.2021.07.066. PMID 34637739. Bibcode: 2021CBio...31R1267B.
- ↑ Harper, David; Benton, Michael (2009). "Protists". Introduction to Paleobiology and the Fossil Record. Wiley-Blackwell. pp. 204–233. ISBN 978-1-4051-4157-4. https://www.a-centauri.com/archivio/geo/Paleontologia/Benton_Introduction%20to%20Paleobiology%20and%20the%20Fossil%20Record.pdf.
- ↑ 5.0 5.1 5.2 5.3 "Not plants or animals: A brief history of the origin of Kingdoms Protozoa, Protista, and Protoctista". International Microbiology 2 (4): 207–221. 1999. PMID 10943416. http://revistes.iec.cat/index.php/IM/article/viewFile/9219/9217.
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedLevine-19802 - ↑ Karpov, Sergey A.; Mamkaeva, Maria A.; Aleoshin, Vladimir V.; Nassonova, Elena; Lilje, Osu; Gleason, Frank H. (2014). "Morphology, phylogeny, and ecology of the aphelids (Aphelidea, Opisthokonta) and proposal for the new superphylum Opisthosporidia". Frontiers in Microbiology 5: 112. doi:10.3389/fmicb.2014.00112. PMID 24734027. Bibcode: 2014FrMic...500112K.
- ↑ Corliss, J. O. (1998). "Haeckel's Kingdom Protista and Current Concepts in Systematic Protistology". Stapfia 56: 85–104. https://www.zobodat.at/pdf/STAPFIA_0056_0085-0104.pdf.
- ↑ Corliss, John O. (19 April 2001). "Protist systematics". Encyclopedia of Life Sciences. John Wiley & Sons. doi:10.1038/npg.els.0003153. ISBN 978-0-470-01590-2.
- ↑ 10.0 10.1 "A revised six-kingdom system of life". Biological Reviews of the Cambridge Philosophical Society 73 (3): 203–266. August 1998. doi:10.1111/j.1469-185X.1998.tb00030.x. PMID 9809012. Bibcode: 1998BioRv..73..203C.
- ↑ 11.00 11.01 11.02 11.03 11.04 11.05 11.06 11.07 11.08 11.09 11.10 11.11 11.12 11.13 11.14 11.15 11.16 11.17 Adl, Sina M.; Bass, David; Lane, Christopher E.; Lukeš, Julius; Schoch, Conrad L. et al. (2019). "Revisions to the Classification, Nomenclature, and Diversity of Eukaryotes". Journal of Eukaryotic Microbiology 66 (1): 4–119. doi:10.1111/JEU.12691. PMID 30257078.
- ↑ Tedersoo, Leho; Sánchez-Ramírez, Santiago; Kõljalg, Urmas; Bahram, Mohammad; Döring, Markus et al. (2018), "High-level classification of the Fungi and a tool for evolutionary ecological analyses", Fungal Diversity 90: 135–159, doi:10.1007/s13225-018-0401-0
- ↑ Wijayawardene, N.N.; Hyde, K.D.; Dai, D.Q.; Sánchez-García, M.; Goto, B.T. et al. (2022). "Outline of Fungi and fungus-like taxa – 2021". Mycosphere 13 (1): 53–453. doi:10.5943/mycosphere/13/1/2. https://www.researchgate.net/publication/358798332.
- ↑ 14.0 14.1 14.2 Hausmann, Klaus; Hülsmann, Norbert; Radek, Renate (1996). "Cellular Organization of Protists". Protistology (3rd ed.). Berlin: E. Schweizerbart'sche Verlagsbuchhandlung. pp. 13–29. ISBN 3-510-65208-8. https://archive.org/details/protistology0000haus.
- ↑ 15.0 15.1 15.2 15.3 Lamża, Łukasz (December 2023). "Diversity of 'simple' multicellular eukaryotes: 45 independent cases and six types of multicellularity". Biological Reviews 98 (6): 2188–2209. doi:10.1111/brv.13001. PMID 37475165.
- ↑ 16.0 16.1 Sleigh, Michael A. (1989). "Protozoa as members of the Protista". Protozoa and other protists. New York: E. Arnold. pp. 1–12. ISBN 0-7131-2943-3. https://archive.org/details/lbg.593.2571.
- ↑ Jeuck, Alexandra; Arndt, Hartmut (2013). "A Short Guide to Common Heterotrophic Flagellates of Freshwater Habitats Based on the Morphology of Living Organisms". Protist 164 (6): 842–860. doi:10.1016/j.protis.2013.08.003.
- ↑ "Lynn Margulis replies". BioScience 36 (5): 293–294. 1986. doi:10.1093/bioscience/36.5.293-a.
- ↑ Margulis, Lynn (1980). "Undulipodia, flagella and cilia". Biosystems 12 (1–2): 105–108. doi:10.1016/0303-2647(80)90041-6. PMID 7378551. Bibcode: 1980BiSys..12..105M.
- ↑ Andersen, R. A.; Barr, D. J. S.; Lynn, D. H.; Melkonian, M.; Moestrup, Ø.; Sleigh, M. A. (1991). "Terminology and nomenclature of the cytoskeletal elements associated with the flagellar/ciliary apparatus in protists". Protoplasma 164 (1–3): 1–8. doi:10.1007/BF01320809. Bibcode: 1991Prpls.164....1A.
- ↑ Taylor, F. J. R. (1990). "Phylum Dinoflagellata". in Margulis, Lynn. Handbook of Protoctista. Boston: Jones and Bartlett Publishers. pp. 419–437. ISBN 0-86720-052-9. https://archive.org/details/handbookofprotoc0000unse/page/418.
- ↑ Esteban, Genoveva F.; Fenchel, Tom M. (2020). "What is a protozoon?". Ecology of Protozoa: The Biology of Free-living Phagotrophic Protists. Cham: Springer. pp. 1–14. doi:10.1007/978-3-030-59979-9_1.
- ↑ Gooday, A.J; Aranda da Silva, A.; Pawlowski, J. (2011). "Xenophyophores (Rhizaria, Foraminifera) from the Nazare Canyon (Portuguese margin, NE Atlantic)". Deep-Sea Research Part II 58 (23–24): 2401–2419. doi:10.1016/j.dsr2.2011.04.005. Bibcode: 2011DSRII..58.2401G.
- ↑ 24.0 24.1 Pánek, Tomáš; Simpson, Alastair G. B.; Brown, Matthew W.; Dyer, Betsey Dexter (2017). "Heterolobosea". Handbook of the Protists. 2 (2nd ed.). Springer. pp. 1005–1046. doi:10.1007/978-3-319-28149-0_10. ISBN 978-3-319-28147-6.
- ↑ H. Lynn, Denis (2017). "Ciliophora". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 679–730. doi:10.1007/978-3-319-28149-0_23. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Kostka, Martin (2017). "Opalinata". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 543–565. doi:10.1007/978-3-319-28149-0_4. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Margulis, Lynn (1990). "Introduction". in Margulis, Lynn. Handbook of Protoctista. Boston: Jones and Bartlett Publishers. pp. xi–xiii. ISBN 0-86720-052-9. https://archive.org/details/handbookofprotoc0000unse.
- ↑ "A newly revised classification of the Protozoa". Journal of Protozoology 27 (1): 37–58. 1980. doi:10.1111/j.1550-7408.1980.tb04228.x. PMID 6989987.
- ↑ Price, Dana C.; Steiner, Jürgen M.; Yoon, Hwan Su; Bhattacharya, Debashish; Löffelhardt, Wolfgang (2017). "Glaucophyta". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 23–87. doi:10.1007/978-3-319-28149-0_42. ISBN 978-3-319-28149-0. https://tuaulavirtual.educatic.unam.mx/pluginfile.php/1747217/mod_resource/content/1/Price2016_ReferenceWorkEntry_Glaucophyta.pdf. Retrieved 9 June 2025.
- ↑ Keeling, Patrick J. (2017). "Chlorarachniophytes". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 765–781. doi:10.1007/978-3-319-28149-0_34. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ 31.0 31.1 Graham, Linda E.; Graham, James M.; Wilcox, Lee W.; Cook, Martha E. (2022). "Chapter 1. Introduction to the Algae". Algae (4th ed.). LJLM Press. pp. 1-1–1-21. ISBN 978-0-9863935-4-9.
- ↑ Spiegel, F.W. (2016). "Unikonts, Evolution and Diversification of (with Emphasis on Fungal-Like Forms)". Encyclopedia of Evolutionary Biology. Elsevier. p. 325–332. doi:10.1016/b978-0-12-800049-6.00248-1. ISBN 978-0-12-800426-5. https://linkinghub.elsevier.com/retrieve/pii/B9780128000496002481. Retrieved 12 May 2025.
- ↑ Gleason, Frank H.; Lilje, Osu; Lange, Lene (2018). "What has happened to the "aquatic phycomycetes" (sensu Sparrow)? Part II: Shared properties of zoosporic true fungi and fungus-like microorganisms". Fungal Biology Reviews 32 (2): 52–61. doi:10.1016/j.fbr.2017.09.003. Bibcode: 2018FunBR..32...52G.
- ↑ Neuhauser, Sigrid; Glockling, Sally L.; Leaño, Eduardo M.; Lilje, Osu; Marano, Agostina V.; Gleason, Frank H. (2012). "An introduction to fungus-like microorganisms". Marine fungi. Marine and Freshwater Botany. De Gruyter. pp. 137–152. doi:10.1515/9783110264067.137. ISBN 978-3-11-026406-7.
- ↑ Michael D. Guiry; Øjvind Moestrup; Robert A. Andersen (11 October 2023). "Validation of the phylum name Heterokontophyta". Notulae Algarum 2023 (297). https://www.notulaealgarum.com/2023/documents/Notulae%20Algarum%20No.%20297.pdf.
- ↑ Umen, James; Herron, Matthew D. (23 November 2021). "Green Algal Models for Multicellularity". Annual Review of Genetics 55 (1): 603–632. doi:10.1146/annurev-genet-032321-091533. ISSN 0066-4197. https://www.annualreviews.org/docserver/fulltext/genet/55/1/annurev-genet-032321-091533.pdf. Retrieved 27 February 2026.
- ↑ Laundon, Davis; Larson, Ben T.; McDonald, Kent; King, Nicole; Burkhardt, Pawel (12 April 2019). "The architecture of cell differentiation in choanoflagellates and sponge choanocytes". PLOS Biology 17 (4). doi:10.1371/journal.pbio.3000226. ISSN 1545-7885. PMID 30978201.
- ↑ 38.0 38.1 38.2 38.3 38.4 38.5 38.6 Yazaki, Euki; Shiratori, Takashi; Inagaki, Yuji (18 August 2025). "Protists with Uncertain Phylogenetic Affiliations for Resolving the Deep Tree of Eukaryotes". Microorganisms 13 (8). doi:10.3390/microorganisms13081926. ISSN 2076-2607. PMID 40871430.
- ↑ Barry S. C. Leadbeater; Sharon M. M. McReady (2000). "Chapter 1. The flagellates: historical perspectives". The Flagellates. Unity, diversity and evolution. London: Taylor & Francis. pp. 1–26. doi:10.1201/9781482268225. ISBN 978-0-429-18213-6.
- ↑ 40.0 40.1 Marc J. Ratcliff (2009). "The Emergence of the Systematics of Infusoria". The Quest for the Invisible: Microscopy in the Enlightenment. Ashgate. pp. 177–216. ISBN 978-1-4094-8026-6. https://books.google.com/books?id=GTZt1c5IYioC.
- ↑ John Hogg (1860). "On the distinctions of a Plant and an Animal, and on a Fourth Kingdom of Nature". Edinburgh New Philosophical Journal. 2nd series 12: 216–225. https://babel.hathitrust.org/cgi/pt?id=hvd.32044089575245;view=1up;seq=232. "... I here suggest a fourth or an additional kingdom, under the title of the Primigenal kingdom, ... This Primigenal kingdom would comprise all the lower creatures, or the primary organic beings, – 'Protoctista,' – from πρώτος, first, and χτιστά, created beings; ...".
- ↑ Haeckel, Ernst (1866) (in de). Generelle Morphologie der Organismen. 1. Berlin, (Germany): G. Reimer. pp. 215ff. https://archive.org/details/generellemorphol01haec/page/214. From p. 215: "VII. Character des Protistenreiches." (VII. Character of the kingdom of Protists.)
- ↑ "The Kingdoms of Organisms". Quarterly Review of Biology 13 (4): 383–420. 1938. doi:10.1086/394568.
- ↑ Margulis, Lynn; Chapman, Michael J. (2009). Kingdoms & Domains: An Illustrated Guide to the Phyla of Life on Earth (4th ed.). AP. ISBN 978-0-12-373621-5.
- ↑ "New concepts of kingdoms or organisms: Evolutionary relations are better represented by new classifications than by the traditional two kingdoms". Science 163 (3863): 150–160. January 1969. doi:10.1126/science.163.3863.150. PMID 5762760. Bibcode: 1969Sci...163..150W.
- ↑ Hagen, Joel B. (2012). "Five Kingdoms, More or Less: Robert Whittaker and the Broad Classification of Organisms". BioScience 62: 67–74. doi:10.1525/bio.2012.62.1.11.
- ↑ 47.0 47.1 Patterson, David J. (1994). "Protozoa: Evolution and Systematics". Progress in Protozoology: Proceedings of the IX International Congress of Protozoology, Berlin 25–31 July 1993. Gustav Fischer. pp. 1–14. https://archive.org/details/progressinprotoz0000klau/page/1.
- ↑ Patterson, David J. (October 1999). "The Diversity of Eukaryotes". The American Naturalist 154 (S4): S96–S124. doi:10.1086/303287. PMID 10527921. https://www.journals.uchicago.edu/doi/epdf/10.1086/303287.
- ↑ Schlegel, M.; Hulsmann, N. (2007). "Protists – A textbook example for a paraphyletic taxon☆". Organisms Diversity & Evolution 7 (2): 166–172. doi:10.1016/j.ode.2006.11.001. Bibcode: 2007ODivE...7..166S.
- ↑ Adl, Sina M.; Simpson, Alastair G. B.; Farmer, Mark A.; Andersen, Robert A.; Anderson, O. Roger et al. (19 October 2005). "The New Higher Level Classification of Eukaryotes with Emphasis on the Taxonomy of Protists". The Journal of Eukaryotic Microbiology 52 (2): 399–451. doi:10.1111/j.1550-7408.2005.00053.x. PMID 16248873.
- ↑ Adl, Sina M.; Simpson, Alastair G. B.; Lane, Christopher E.; Lukeš, Julius; Bass, David et al. (28 September 2012). "The Revised Classification of Eukaryotes". The Journal of Eukaryotic Microbiology 59 (2): 429–514. doi:10.1111/j.1550-7408.2012.00644.x. PMID 23020233.
- ↑ Ruggiero, Michael A.; Gordon, Dennis P.; Orrell, Thomas M.; Bailly, Nicolas; Bourgoin, Thierry; Brusca, Richard C.; Cavalier-Smith, Thomas; Guiry, Michael D. et al. (2015). "A higher level classification of all living organisms". PLOS ONE 10 (4). doi:10.1371/journal.pone.0119248. PMID 25923521. Bibcode: 2015PLoSO..1019248R.
- ↑ Burki, Fabien; Kaplan, Maia; Tikhonenkov, Denis V.; Zlatogursky, Vasily; Minh, Bui Quang et al. (27 January 2016). "Untangling the early diversification of eukaryotes: a phylogenomic study of the evolutionary origins of Centrohelida, Haptophyta and Cryptista". Proceedings of the Royal Society B: Biological Sciences 283 (1823). doi:10.1098/rspb.2015.2802. ISSN 0962-8452. PMID 26817772.
- ↑ "Ciliary transition zone evolution and the root of the eukaryote tree: implications for opisthokont origin and classification of kingdoms Protozoa, Plantae, and Fungi". Protoplasma 259 (3): 487–593. May 2022. doi:10.1007/s00709-021-01665-7. PMID 34940909. Bibcode: 2022Prpls.259..487C.
- ↑ 55.0 55.1 Lax, Gordon; Eglit, Yana; Eme, Laura; Bertrand, Erin M.; Roger, Andrew J.; Simpson, Alastair G. B. (2018-11-14). "Hemimastigophora is a novel supra-kingdom-level lineage of eukaryotes" (in En). Nature 564 (7736): 410–414. doi:10.1038/s41586-018-0708-8. ISSN 0028-0836. PMID 30429611. Bibcode: 2018Natur.564..410L.
- ↑ Burki, Fabien; Roger, Andrew J.; Brown, Matthew W.; Simpson, Alastair G.B. (9 October 2019). "The New Tree of Eukaryotes". Trends in Ecology & Evolution 35 (1): 43–55. doi:10.1016/j.tree.2019.08.008. PMID 31606140. Bibcode: 2020TEcoE..35...43B. https://www.cell.com/article/S0169534719302575/pdf. Retrieved 2025-06-05.
- ↑ "Microbial predators form a new supergroup of eukaryotes". Nature 612 (7941): 714–719. 2022. doi:10.1038/s41586-022-05511-5. PMID 36477531. Bibcode: 2022Natur.612..714T.
- ↑ Eglit, Yana; Shiratori, Takashi; Jerlström-Hultqvist, Jon; Williamson, Kelsey; Roger, Andrew J.; Ishida, Ken-Ichiro; Simpson, Alastair G.B. (22 January 2024). "Meteora sporadica, a protist with incredible cell architecture, is related to Hemimastigophora". Current Biology 34 (2): 451–459. doi:10.1016/j.cub.2023.12.032. PMID 38262350. Bibcode: 2024CBio...34..451E.
- ↑ "Phylogenomics reshuffles the eukaryotic supergroups". PLOS ONE 2 (8). August 2007. doi:10.1371/journal.pone.0000790. PMID 17726520. Bibcode: 2007PLoSO...2..790B.
- ↑ Simpson, Alastair G. B.; Slamovits, Claudio H.; Archibald, John M. (2017). "Protist Diversity and Eukaryote Phylogeny". Handbook of the Protists. 1 (2nd ed.). Springer. pp. 1–22. doi:10.1007/978-3-319-28149-0_45. ISBN 978-3-319-28147-6.
- ↑ 61.0 61.1 61.2 Valt, Marek; Pánek, Tomáš; Mirzoyan, Seda; Tice, Alexander K.; Jones, Robert E. et al. (19 November 2025). "Rare microbial relict sheds light on an ancient eukaryotic supergroup" (in en). Nature. doi:10.1038/s41586-025-09750-0. ISSN 0028-0836. https://www.researchsquare.com/article/rs-5245440/v1.pdf. Retrieved 25 November 2025.
- ↑ 62.0 62.1 Simpson, Alastair G. B. (2003). "Cytoskeletal organization, phylogenetic affinities and systematics in the contentious taxon Excavata (Eukaryota)". International Journal of Systematic and Evolutionary Microbiology 53 (6): 1759–1777. doi:10.1099/ijs.0.02578-0. PMID 14657103. Bibcode: 2003IJSEM..53.1759S.
- ↑ 63.0 63.1 Madigan, Michael T.; Bender, Kelly S.; Buckley, Daniel H.; Sattley, W. Matthew; Stahl, David A. (2019). "Diversity of Microbial Eukarya". Brock Biology of Microorganisms (15th, Global ed.). Pearson. pp. 593–618. ISBN 978-1-292-23510-3.
- ↑ 64.0 64.1 Suzuki-Tellier, Sei; Kiørboe, Thomas; Simpson, Alastair G. B. (2023). "The function of the feeding groove of 'typical excavate' flagellates". Journal of Eukaryotic Microbiology 71 (2). doi:10.1111/jeu.13016. PMID 38108228.
- ↑ Jirsová, Dagmar; Wideman, Jeremy G (8 January 2024). "Integrated overview of stramenopile ecology, taxonomy, and heterotrophic origin". The ISME Journal 18 (1). doi:10.1093/ismejo/wrae150. ISSN 1751-7362. PMID 39077993.
- ↑ H.S. Yoon; R.A. Andersen; S.M. Boo; D. Bhattacharya (17 February 2009). "Stramenopiles". Encyclopedia of Microbiology (Third ed.). pp. 721–731. doi:10.1016/B978-012373944-5.00253-4. ISBN 978-0-12-373944-5. https://www.sciencedirect.com/science/article/abs/pii/B9780123739445002534. Retrieved 2 March 2024.
- ↑ 67.0 67.1 Tikhonenkov, Denis V.; Strassert, Jürgen F.H.; Janouškovec, Jan; Mylnikov, Alexander P.; Aleoshin, Vladimir V.; Burki, Fabien; Keeling, Patrick J. (2020). "Predatory colponemids are the sister group to all other alveolates". Molecular Phylogenetics and Evolution 149. doi:10.1016/j.ympev.2020.106839. https://linkinghub.elsevier.com/retrieve/pii/S1055790320301111. Retrieved 19 February 2026.
- ↑ H. Lynn, Denis (2017). "Ciliophora". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 679–730. doi:10.1007/978-3-319-28149-0_23. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Janouškovec, Jan; Tikhonenkov, Denis V.; Burki, Fabien; Howe, Alexis T.; Kolísko, Martin; Mylnikov, Alexander P.; Keeling, Patrick J. (2015). "Factors mediating plastid dependency and the origins of parasitism in apicomplexans and their close relatives". PNAS 112 (33): 10200–10207. doi:10.1073/pnas.1423790112. PMID 25717057. Bibcode: 2015PNAS..11210200J.
- ↑ Mathur, Varsha; Kolísko, Martin; Hehenberger, Elisabeth; Irwin, Nicholas A.T.; Leander, Brian S. et al. (2019). "Multiple Independent Origins of Apicomplexan-Like Parasites". Current Biology 29 (17): 2936–2941. doi:10.1016/j.cub.2019.07.019. PMID 31422883.
- ↑ Holt, Corey C.; Hehenberger, Elisabeth; Tikhonenkov, Denis V.; Jacko-Reynolds, Victoria K. L.; Okamoto, Noriko; Cooney, Elizabeth C.; Irwin, Nicholas A. T.; Keeling, Patrick J. (3 November 2023). "Multiple parallel origins of parasitic Marine Alveolates". Nature Communications 14 (1). doi:10.1038/s41467-023-42807-0. ISSN 2041-1723. PMID 37923716.
- ↑ Cavalier-Smith, Thomas; Chao, Ema E.; Lewis, Rhodri (2018). "Multigene phylogeny and cell evolution of chromist infrakingdom Rhizaria: contrasting cell organisation of sister phyla Cercozoa and Retaria". Protoplasma 255 (5): 1517–1574. doi:10.1007/s00709-018-1241-1. PMID 29666938. Bibcode: 2018Prpls.255.1517C.
- ↑ Brown, Matthew W.; Kolisko, Martin; Silberman, Jeffrey D.; Rogers, Andrew J. (2012). "Aggregative multicellularity evolved independently in the eukaryotic supergroup Rhizaria". Current Biology 22 (12): 1123–1127. doi:10.1016/j.cub.2012.04.021. PMID 22608512. Bibcode: 2012CBio...22.1123B.
- ↑ Lax, Gordon; Cooney, Elizabeth C.; Zlatogursky, Vasily; Mtawali, Mahara; Okamoto, Noriko et al. (30 January 2026). "Phylogenomic tree of Cercozoa based on single-cell transcriptomes from 100 uncultured cells". BMC Biology 24 (1). doi:10.1186/s12915-026-02536-4. ISSN 1741-7007. PMID 41618358.
- ↑ 75.0 75.1 Burki, Fabien; Keeling, Patrick J. (2014). "Rhizaria". Current Biology 24 (3): R103–R107. doi:10.1016/j.cub.2013.12.025.
- ↑ 76.0 76.1 76.2 76.3 Adl, Sina M.; Leander, Brian S.; Simpson, Alastair G. B.; Archibald, John M.; Anderson, O. Roger. et al. (1 August 2007). "Diversity, Nomenclature, and Taxonomy of Protists". Systematic Biology 56 (4): 684–689. doi:10.1080/10635150701494127. ISSN 1076-836X. PMID 17661235. https://academic.oup.com/sysbio/article-pdf/56/4/684/24203799/56-4-684.pdf. Retrieved 15 July 2025.
- ↑ Tikhonenkov, Denis V.; Jamy, Mahwash; Borodina, Anastasia S.; Belyaev, Artem O.; Zagumyonnyi, Dmitry G.; Prokina, Kristina I.; Mylnikov, Alexander P.; Burki, Fabien et al. (2022). "On the origin of TSAR: morphology, diversity and phylogeny of Telonemia". Open Biology (The Royal Society) 12 (3). doi:10.1098/rsob.210325. ISSN 2046-2441. PMID 35291881.
- ↑ Zlatogursky, Vasily; Boscaro, Vittorio; Lax, Gordon; Wanntorp, Matias; Pohl, Nina; Burki, Fabien; Keeling, Patrick J. (15 August 2025). "Phylogenetic position and mitochondrial genome evolution of "orphan" eukaryotic lineages". iScience 28 (8). doi:10.1016/j.isci.2025.113184. PMID 40948565.
- ↑ Mostazo‐Zapata, Helena; Gàlvez‐Morante, Alex; Berney, Cédric; Maya‐Figuerola, Xènia; Sigona, Cristiana; López‐Escardó, David; Sà, Elisabet L.; Vaqué, Dolors et al. (2025). "Description of a New Telonemia Genus and Species With Novel Observations Providing Insights Into Its Hidden Diversity". Journal of Eukaryotic Microbiology 72 (6). doi:10.1111/jeu.70050. ISSN 1066-5234. PMID 41131848.
- ↑ Zagumyonnyi, Dmitry G.; Radaykina, Liudmila V.; Tikhonenkov, Denis V. (11 December 2021). "Triangulopteris lacunata gen. et sp. nov. (Centroplasthelida), a New Centrohelid Heliozoan from Soil". Diversity 13 (12). doi:10.3390/d13120658. ISSN 1424-2818.
- ↑ Eikrem, Wenche; Medlin, Linda K.; Henderiks, Jorijntje; Rokitta, Sebastian; Rost, Björn et al. (2017). "Haptophyta". Handbook of the Protists. 2 (2nd ed.). Cham: Springer International Publishing. pp. 893–953. doi:10.1007/978-3-319-28149-0_38. ISBN 978-3-319-28149-0. https://tuaulavirtual.educatic.unam.mx/pluginfile.php/1747249/mod_resource/content/1/Eikrem2017_ReferenceWorkEntry_Haptophyta.pdf. Retrieved 9 June 2025.
- ↑ Yazaki, Euki; Yabuki, Akinori; Imaizumi, Ayaka; Kume, Keitaro; Hashimoto, Tetsuo; Inagaki, Yuji (2022-04-13). "The closest lineage of Archaeplastida is revealed by phylogenomics analyses that include Microheliella maris". Open Biology 12 (4). doi:10.1098/rsob.210376. PMID 35414259. PMC 9006020. https://royalsocietypublishing.org/doi/10.1098/rsob.210376.
- ↑ Hoef-Emden, Kerstin; Archibald, John M. (2017). "Cryptophyta (Cryptomonads)". Handbook of the Protists. 2 (2nd ed.). Cham: Springer International Publishing. pp. 851–891. doi:10.1007/978-3-319-28149-0_35. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Prokina, Kristina I.; Tikhonenkov, Denis V.; López-García, Purificación; Moreira, David (2023). "Morphological and molecular characterization of a new member of the phylum Rhodelphidia". Journal of Eukaryotic Microbiology 71 (2). doi:10.1111/jeu.12995. PMID 37548159.
- ↑ Schön, Max E.; Zlatogursky, Vasily V.; Singh, Rohan P.; Poirier, Camille; Wilken, Susanne; Mathur, Varsha; Strassert, Jürgen F. H.; Pinhassi, Jarone et al. (2021). "Single cell genomics reveals plastid-lacking Picozoa are close relatives of red algae". Nature Communications 12 (1): 6651. doi:10.1038/s41467-021-26918-0. PMID 34789758. Bibcode: 2021NatCo..12.6651S.
- ↑ Price, Dana C.; Steiner, Jürgen M.; Yoon, Hwan Su; Bhattacharya, Debashish; Löffelhardt, Wolfgang (2017). "Glaucophyta". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 23–87. doi:10.1007/978-3-319-28149-0_42. ISBN 978-3-319-28149-0. https://tuaulavirtual.educatic.unam.mx/pluginfile.php/1747217/mod_resource/content/1/Price2016_ReferenceWorkEntry_Glaucophyta.pdf. Retrieved 9 June 2025.
- ↑ Bowles, Alexander M.C.; Williamson, Christopher J.; Williams, Tom A.; Lenton, Timothy M.; Donoghue, Philip C.J. (2023). "The origin and early evolution of plants". Trends in Plant Science 28 (3): 312–329. doi:10.1016/j.tplants.2022.09.009.
- ↑ Tikhonenkov, Denis V.; Mikhailov, Kirill V.; Gawryluk, Ryan M. R.; Belyaev, Artem O.; Mathur, Varsha et al. (22 December 2022). "Microbial predators form a new supergroup of eukaryotes". Nature 612 (7941): 714–719. doi:10.1038/s41586-022-05511-5. PMID 36477531.
- ↑ Belyaev, Artem O.; Karpov, Sergey A.; Keeling, Patrick J.; Tikhonenkov, Denis V. (18 December 2024). "The nature of 'jaws': a new predatory representative of Provora and the ultrastructure of nibbling protists". Open Biology 14 (12). doi:10.1098/rsob.240158. PMID 39689855.
- ↑ Smirnov, Alexey; Nassonova, Elena; Berney, Cédric; Fahrni, José; Bolivar, Ignacio; Pawlowski, Jan (2005). "Molecular Phylogeny and Classification of the Lobose Amoebae". Protist 156 (2): 129–142. doi:10.1016/j.protis.2005.06.002. PMID 16171181.
- ↑ Kang, Seungho; Tice, Alexander K; Spiegel, Frederick W; Silberman, Jeffrey D; Pánek, Tomáš et al. (September 2017). "Between a Pod and a Hard Test: The Deep Evolution of Amoebae". Molecular Biology and Evolution 34 (9): 2258–2270. doi:10.1093/molbev/msx162. PMID 28505375.
- ↑ Tekle, Yonas I.; Wang, Fang; Wood, Fiona C.; Anderson, O. Roger; Smirnov, Alexey (1 July 2022). "New insights on the evolutionary relationships between the major lineages of Amoebozoa". Scientific Reports 12 (1): 11173. doi:10.1038/s41598-022-15372-7. PMID 35778543. Bibcode: 2022NatSR..1211173T.
- ↑ González-Miguéns, Rubén; Todorov, Milcho; Blandenier, Quentin; Duckert, Clément; Porfirio-Sousa, Alfredo L.; Ribeiro, Giulia M.; Ramos, Diana; Lahr, Daniel J.G. et al. (2022). "Deconstructing Difflugia: The tangled evolution of lobose testate amoebae shells (Amoebozoa: Arcellinida) illustrates the importance of convergent evolution in protist phylogeny". Molecular Phylogenetics and Evolution 175. doi:10.1016/j.ympev.2022.107557. PMID 35777650. Bibcode: 2022MolPE.17507557G.
- ↑ Brown, Matthew W.; Silberman, Jeffrey D.; Spiegel, Frederick W. (26 November 2010). ""Slime Molds" among the Tubulinea (Amoebozoa): Molecular Systematics and Taxonomy of Copromyxa". Protist 162 (2): 277–287. doi:10.1016/j.protis.2010.09.003. PMID 21112814.
- ↑ 95.0 95.1 Lamża, Łukasz (2023). "Diversity of 'simple' multicellular eukaryotes: 45 independent cases and six types of multicellularity". Biological Reviews 98 (6): 2188–2209. doi:10.1111/brv.13001. ISSN 1464-7931.
- ↑ Martinez, Augusto Julio; Visvesvara, Govinda S. (1997). "Free‐living, Amphizoic and Opportunistic Amebas". Brain Pathology 7 (1): 583–598. doi:10.1111/j.1750-3639.1997.tb01076.x. ISSN 1015-6305. PMID 9034567.
- ↑ 97.0 97.1 97.2 Pawlowski, Jan; Audic, Stéphane; Adl, Sina; Bass, David; Belbahri, Lassaâd et al. (6 November 2012). "CBOL Protist Working Group: Barcoding Eukaryotic Richness beyond the Animal, Plant, and Fungal Kingdoms". PLoS Biology 10 (11). doi:10.1371/journal.pbio.1001419. ISSN 1545-7885. PMID 23139639.
- ↑ Aguilera-Campos, Karla Iveth; Boisard, Julie; Törnblom, Viktor; Jerlström-Hultqvist, Jon; Behncké-Serra, Ada; Cotillas, Elena Aramendia; Stairs, Courtney Weir (2 January 2025). "Anaerobic breviate protist survival in microcosms depends on microbiome metabolic function". The ISME Journal 19 (1). doi:10.1093/ismejo/wraf171. ISSN 1751-7362. PMID 40795332.
- ↑ 99.0 99.1 99.2 Heiss, Aaron A.; Brown, Matthew W.; Simpson, Alastair G. B. (2017). "Apusomonadida". Handbook of the Protists. 2 (2nd ed.). Cham: Springer International Publishing. pp. 1619–1645. doi:10.1007/978-3-319-28149-0_12. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ 100.0 100.1 Torruella, Guifré; Galindo, Luis Javier; Moreira, David; Ciobanu, Maria; Heiss, Aaron A. et al. (2023). "Expanding the molecular and morphological diversity of Apusomonadida, a deep‐branching group of gliding bacterivorous protists". Journal of Eukaryotic Microbiology 70 (2). doi:10.1111/jeu.12956. PMID 36453005. https://hal.science/hal-04384861.
- ↑ Torruella, Guifré; de Mendoza, Alex; Grau-Bové, Xavier; Antó, Meritxell; Chaplin, Mark A. et al. (2015). "Phylogenomics Reveals Convergent Evolution of Lifestyles in Close Relatives of Animals and Fungi". Current Biology 25 (18): 2404–2410. doi:10.1016/j.cub.2015.07.053. PMID 26365255.
- ↑ 102.0 102.1 Gabaldón, Toni; Völcker, Eckhard; Torruella, Guifré (2022). "On the Biology, Diversity and Evolution of Nucleariid Amoebae (Amorphea, Obazoa, Opisthokonta)". Protist 173 (4). doi:10.1016/j.protis.2022.125895. PMID 35841659.
- ↑ Brunet, Thibaut; Albert, Marvin; Roman, William; Coyle, Maxwell C; Spitzer, Danielle C; King, Nicole (15 January 2021). "A flagellate-to-amoeboid switch in the closest living relatives of animals". eLife 10. doi:10.7554/eLife.61037. ISSN 2050-084X.
- ↑ Richter, Daniel J.; Nitsche, Frank (2017). "Choanoflagellatea". Handbook of the Protists. 2 (2nd ed.). Springer. pp. 1479–1496. doi:10.1007/978-3-319-28149-0_5. ISBN 978-3-319-28147-6.
- ↑ Hehenberger, Elisabeth; Tikhonenkov, Denis V.; Kolisko, Martin; del Campo, Javier; Esaulov, Anton S.; Mylnikov, Alexander P.; Keeling, Patrick J. (10 July 2017). "Novel Predators Reshape Holozoan Phylogeny and Reveal the Presence of a Two-Component Signaling System in the Ancestor of Animals". Current Biology 27 (13): 2043–2050. doi:10.1016/j.cub.2017.06.006. PMID 28648822. Bibcode: 2017CBio...27E2043H.
- ↑ Tikhonenkov, Denis V.; Mikhailov, Kirill V.; Hehenberger, Elisabeth; Karpov, Sergei A.; Prokina, Kristina I. et al. (16 November 2020). "New Lineage of Microbial Predators Adds Complexity to Reconstructing the Evolutionary Origin of Animals". Current Biology 30 (22): 4500–4509.e5. doi:10.1016/J.CUB.2020.08.061. PMID 32976804. Bibcode: 2020CBio...30E4500T.
- ↑ Glockling, Sally L.; Marshall, Wyth L.; Gleason, Frank H. (25 April 2013). "Phylogenetic interpretations and ecological potentials of the Mesomycetozoea (Ichthyosporea)". Fungal Ecology 6 (4): 237–247. doi:10.1016/j.funeco.2013.03.005.
- ↑ Urrutia, Ander; Mitsi, Konstantina; Foster, Rachel; Ross, Stuart; Carr, Martin et al. (2022). "Txikispora philomaios n. sp., n. g., a micro-eukaryotic pathogen of amphipods, reveals parasitism and hidden diversity in Class Filasterea". Journal of Eukaryotic Microbiology 69 (2). doi:10.1111/jeu.12875. PMID 34726818. https://pure.hud.ac.uk/en/publications/1dd4e1c5-f638-48cf-ab99-95c9c90636a1.
- ↑ Kostygov, Alexei Y.; Karnkowska, Anna; Votýpka, Jan; Tashyreva, Daria; Maciszewski, Kacper; Yurchenko, Vyacheslav; Lukeš, Julius (2021). "Euglenozoa: taxonomy, diversity and ecology, symbioses and viruses". Open Biology 11 (3). doi:10.1098/rsob.200407. PMID 33715388.
- ↑ Simpson, Alastair G. B. (2017). "Jakobida". Handbook of the Protists. 2 (2nd ed.). Springer. pp. 973–1004. doi:10.1007/978-3-319-28149-0_6. ISBN 978-3-319-28147-6.
- ↑ 111.0 111.1 Boscaro, Vittorio; James, Erick R.; Fiorito, Rebecca; del Campo, Javier; Scheffrahn, Rudolf H.; Keeling, Patrick J. (2024). "Updated classification of the phylum Parabasalia". Journal of Eukaryotic Microbiology 71 (4). doi:10.1111/jeu.13035. ISSN 1066-5234.
- ↑ Karnkowska, Anna; Vacek, Vojtěch; Zubáčová, Zuzana; Treitli, Sebastian C.; Petrželková, Romana et al. (2016). "A Eukaryote without a Mitochondrial Organelle". Current Biology 26 (10): 1274–1284. doi:10.1016/j.cub.2016.03.053.
- ↑ Yubuki, Naoji; Huang, Sam S.C.; Leander, Brian S. (2016). "Comparative Ultrastructure of Fornicate Excavates, Including a Novel Free-living Relative of Diplomonads: Aduncisulcus paluster gen. et sp. nov.". Protist 167 (6): 584–596. doi:10.1016/j.protis.2016.10.001.
- ↑ 114.0 114.1 Heiss, Aaron A.; Warring, Sally D.; Lukacs, Kaleigh; Favate, John; Yang, Ashley et al. (2021). "Description of Imasa heleensis , gen. nov., sp. nov. (Imasidae, fam. nov.), a Deep‐Branching Marine Malawimonad and Possible Key Taxon in Understanding Early Eukaryotic Evolution". Journal of Eukaryotic Microbiology 68 (2). doi:10.1111/jeu.12837.
- ↑ 115.0 115.1 Yubuki, Naoji; Torruella, Guifré; Galindo, Luis Javier; Heiss, Aaron A.; Ciobanu, Maria Cristina et al. (22 August 2023). "Molecular and morphological characterization of four new ancyromonad genera and proposal for an updated taxonomy of the Ancyromonadida". Journal of Eukaryotic Microbiology 70 (6). doi:10.1111/jeu.12997. ISSN 1550-7408. PMID 37606230.
- ↑ "How Many Species Are There on Earth and in the Ocean?". PLOS Biology 9 (8). 2011. doi:10.1371/journal.pbio.1001127. PMID 21886479.
- ↑ "Evolutionary cell biology of proteins from protists to humans and plants". J. Eukaryot. Microbiol. 65 (2): 255–289. 2018. doi:10.1111/jeu.12449. PMID 28719054. http://nbn-resolving.de/urn:nbn:de:bsz:352-2-1kezn7g0ruv3m7.
- ↑ 118.0 118.1 118.2 118.3 118.4 118.5 118.6 118.7 Levandowsky, Michael (2012). "Physiological Adaptations of Protists". Cell Physiology Sourcebook: Essentials of Membrane Biophysics (Fourth ed.). Amsterdam; Boston: Elsevier/AP. pp. 874–890. doi:10.1016/B978-012656976-6/50153-0. ISBN 978-0-12-387738-3.
- ↑ 119.0 119.1 119.2 119.3 Wiser, Mark F. (1 July 2024). "Feeding Mechanisms of Pathogenic Protozoa with a Focus on Endocytosis and the Digestive Vacuole". Parasitologia 4 (3): 222–237. doi:10.3390/parasitologia4030019.
- ↑ "Nomenclature of nutritional types of microorganisms.". Cold Spring Harbor Symposia on Quantitative Biology 11: 302–3. 1946. http://symposium.cshlp.org/content/11/local/back-matter.pdf.
- ↑ 121.0 121.1 Hickman, Cleveland P. Jr.; Keen, Susan L.; Eisenhour, David J.; Larson, Allan; l'Anson, Helen (2017). "Unicellular Eukaryotes: Protozoan Groups". Integrated Principles of Zoology (7th ed.). New York: McGraw Hill. pp. 216–245. ISBN 978-1-259-56231-0.
- ↑ 122.0 122.1 122.2 122.3 Leander, Brian S. (2020). "Predatory protists". Current Biology 30 (10): R510–R516. doi:10.1016/j.cub.2020.03.052.
- ↑ 123.0 123.1 123.2 123.3 123.4 Ruppert, Edward E.; Fox, Richard S.; Barnes, Robert D. (2004). "Protozoa". Invertebrate Zoology: A Functional Evolutionary Approach (7th ed.). Thomson Brooks/Cole. pp. 22–57. ISBN 0-03-025982-7.
- ↑ 124.0 124.1 124.2 124.3 Esteban, Genoveva F.; Fenchel, Tom M. (2020). "Feeding". Ecology of Protozoa: The Biology of Free-living Phagotrophic Protists (2nd ed.). Cham: Springer Nature Switzerland AG. pp. 33–54. doi:10.1007/978-3-030-59979-9_4. ISBN 978-3-030-59979-9.
- ↑ 125.0 125.1 125.2 125.3 Geisen, Stefan; Koller, Robert; Hünninghaus, Maike; Dumack, Kenneth; Urich, Tim; Bonkowski, Michael (2016). "The soil food web revisited: Diverse and widespread mycophagous soil protists". Soil Biology and Biochemistry 94: 10–18. doi:10.1016/j.soilbio.2015.11.010. Bibcode: 2016SBiBi..94...10G.
- ↑ Geisen, Stefan; Rosengarten, Jamila; Koller, Robert; Mulder, Christian; Urich, Tim; Bonkowski, Michael (16 June 2015). "Pack-hunting protists attacking nematodes". Environmental Microbiology 17 (11): 4538–4546. doi:10.1111/1462-2920.12949. PMID 26079718.
- ↑ 127.0 127.1 127.2 "Euglenozoan kleptoplasty illuminates the early evolution of photoendosymbiosis". Proceedings of the National Academy of Sciences of the United States of America 120 (12). 21 March 2023. doi:10.1073/pnas.2220100120. PMID 36927158. Bibcode: 2023PNAS..12020100K.
- ↑ 128.0 128.1 Richards, Thomas A.; Talbot, Nicholas J. (10 September 2013). "Horizontal gene transfer in osmotrophs: playing with public goods". Nature Reviews Microbiology 11 (10): 720–727. doi:10.1038/nrmicro3108. PMID 24018383.
- ↑ Harmer, Jane; Yurchenko, Vyacheslav; Nenarokova, Anna; Lukeš, Julius; Ginger, Michael L. (13 June 2018). "Farming, slaving and enslavement: histories of endosymbioses during kinetoplastid evolution". Parasitology 145 (10): 1311–1323. doi:10.1017/S0031182018000781. PMID 29895336. https://pure.hud.ac.uk/ws/files/13639420/Harmer_et_al_2018a_Parasitology_in_press.pdf.
- ↑ "Alternative Nutritional Strategies in Protists: Symposium Introduction and a Review of Freshwater Protists that Combine Photosynthesis and Heterotrophy". Journal of Eukaryotic Microbiology 58 (3): 181–184. 2011. doi:10.1111/j.1550-7408.2011.00543.x. PMID 21477096.
- ↑ Flynn, Kevin J.; Mitra, Aditee; Anestis, Konstantinos; Anschütz, Anna A.; Calbet, Albert; Ferreira, Guilherme Duarte; Gypens, Nathalie; Hansen, Per J. et al. (15 July 2019). "Mixotrophic protists and a new paradigm for marine ecology: where does plankton research go now?". Journal of Plankton Research 41 (4): 375–391. doi:10.1093/plankt/fbz026.
- ↑ 132.0 132.1 Mitra, Aditee; Flynn, Kevin J.; Tillmann, Urban; Raven, John A.; Caron, David et al. (2016). "Defining Planktonic Protist Functional Groups on Mechanisms for Energy and Nutrient Acquisition: Incorporation of Diverse Mixotrophic Strategies". Protist 167 (2): 106–120. doi:10.1016/j.protis.2016.01.003. PMID 26927496.
- ↑ 133.0 133.1 Faure, Emile; Not, Fabrice; Benoiston, Anne-Sophie; Labadie, Karine; Bittner, Lucie; Ayata, Sakina-Dorothée (April 2019). "Mixotrophic protists display contrasted biogeographies in the global ocean". The ISME Journal 13 (4): 1072–1083. doi:10.1038/s41396-018-0340-5. PMID 30643201. Bibcode: 2019ISMEJ..13.1072F.
- ↑ Prokopchuk, Galina; Korytář, Tomáš; Juricová, Valéria; Majstorović, Jovana; Horák, Aleš; Šimek, Karel; Lukeš, Julius (18 January 2022). "Trophic flexibility of marine diplonemids - switching from osmotrophy to bacterivory". The ISME Journal 16 (5): 1409–1419. doi:10.1038/s41396-022-01192-0. PMID 35042972. Bibcode: 2022ISMEJ..16.1409P.
- ↑ 135.0 135.1 Patterson, D. J. (February 1980). "Contractile vacuoles and associated structures: their organization and function". Biological Reviews 55 (1): 1–46. doi:10.1111/j.1469-185x.1980.tb00686.x. Bibcode: 1980BioRv..55....1P.
- ↑ Moreno, Silvia N. J.; Docampo, Roberto (2009). "The Role of Acidocalcisomes in Parasitic Protists 1". Journal of Eukaryotic Microbiology 56 (3): 208–213. doi:10.1111/j.1550-7408.2009.00404.x. ISSN 1066-5234. PMID 19527347.
- ↑ Gawryluk, Ryan M.R.; Kamikawa, Ryoma; Stairs, Courtney W.; Silberman, Jeffrey D.; Brown, Matthew W.; Roger, Andrew J. (2016). "The Earliest Stages of Mitochondrial Adaptation to Low Oxygen Revealed in a Novel Rhizarian". Current Biology 26 (20): 2729–2738. doi:10.1016/j.cub.2016.08.025.
- ↑ Gawryluk, Ryan M.R.; Stairs, Courtney W. (2021). "Diversity of electron transport chains in anaerobic protists". Biochimica et Biophysica Acta (BBA) - Bioenergetics 1862 (1). doi:10.1016/j.bbabio.2020.148334.
- ↑ Voleman, Luboš; Doležal, Pavel (21 November 2019). "Mitochondrial dynamics in parasitic protists". PLOS Pathogens 15 (11). doi:10.1371/journal.ppat.1008008. ISSN 1553-7374.
- ↑ Burger, Gertraud; Gray, Michael W; Franz Lang, B (2003). "Mitochondrial genomes: anything goes". Trends in Genetics 19 (12): 709–716. doi:10.1016/j.tig.2003.10.012.
- ↑ Zíková, Alena; Hampl, Vladimír; Paris, Zdeněk; Týč, Jiří; Lukeš, Julius (2016). "Aerobic mitochondria of parasitic protists: Diverse genomes and complex functions". Molecular and Biochemical Parasitology 209 (1-2): 46–57. doi:10.1016/j.molbiopara.2016.02.007.
- ↑ Pánek, Tomáš; Eliáš, Marek; Vancová, Marie; Lukeš, Julius; Hashimi, Hassan (2020). "Returning to the Fold for Lessons in Mitochondrial Crista Diversity and Evolution". Current Biology 30 (10): R575–R588. doi:10.1016/j.cub.2020.02.053.
- ↑ Benchimol, Marlene (2010). "The Mastigont System in Trichomonads". Structures and Organelles in Pathogenic Protists. Microbiology Monographs. 17. Berlin: Springer-Verlag. pp. 1–26. doi:10.1007/978-3-642-12863-9_1.
- ↑ 144.0 144.1 Yubuki, Naoji; Čepička, Ivan; Leander, Brian S. (2016). "Evolution of the microtubular cytoskeleton (flagellar apparatus) in parasitic protists". Molecular and Biochemical Parasitology 209 (1-2): 26–34. doi:10.1016/j.molbiopara.2016.02.002.
- ↑ 145.0 145.1 145.2 145.3 145.4 Yubuki, Naoji; Leander, Brian S. (2013). "Evolution of microtubule organizing centers across the tree of eukaryotes". The Plant Journal 75 (2): 230–244. doi:10.1111/tpj.12145. ISSN 0960-7412.
- ↑ Moestrup, Øjvind (2000). "The flagellate cytoskeleton: Introduction of a general terminology for microtubular flagellar roots in protists". The flagellates: Unity, diversity and evolution. The Systematics Association Special Volume Series. 59. London: Taylor & Francis. pp. 69–94. https://archive.org/details/flagellatesunity0000unse/page/69/mode/1up.
- ↑ Karpov, Sergey A. (2016). "Flagellar apparatus structure of choanoflagellates". Cilia 5 (1). doi:10.1186/s13630-016-0033-5. ISSN 2046-2530. PMID 27148446.
- ↑ Chapman, Michael; Alliegro, Mark C. (2012). "The Karyomastigont as an Evolutionary Seme". The Quarterly Review of Biology 87 (4): 315–324. doi:10.1086/668165. ISSN 0033-5770.
- ↑ Moestrup, Øjvind (1982). "Phycological Reviews 7. Flagellar structure in algae: a review, with new observations particularly on the Chrysophyceae, Phaeophyceae (Fucophyceae), Euglenophyceae and Reckertia". Phycologia 21 (4): 427–528. doi:10.2216/i0031-8884-21-4-427.1.
- ↑ Hoppenrath, M; Bachvaroff, TR; Handy, SM; Delwiche, CF; Leander, BS (25 May 2009). "Molecular phylogeny of ocelloid-bearing dinoflagellates (Warnowiaceae) as inferred from SSU and LSU rDNA sequences.". BMC Evolutionary Biology 9 (1): 116. doi:10.1186/1471-2148-9-116. PMID 19467154. Bibcode: 2009BMCEE...9..116H.
- ↑ Decelle, Johan; Colin, Sébastien; Foster, Rachel A. (2015). "Photosymbiosis in Marine Planktonic Protists". Marine Protists: Diversity and Dynamics. Tokyo: Springer. pp. 465–500. doi:10.1007/978-4-431-55130-0_19.
- ↑ Foster, Rachel A.; Zehr, Jonathan P. (8 September 2019). "Diversity, Genomics, and Distribution of Phytoplankton-Cyanobacterium Single-Cell Symbiotic Associations". Annual Review of Microbiology 73 (1): 435–456. doi:10.1146/annurev-micro-090817-062650. ISSN 0066-4227.
- ↑ Fenchel, Tom; Bernard, Catherine (1993). "Endosymbiotic purple non-sulphur bacteria in an anaerobic ciliated protozoon". FEMS Microbiology Letters 110 (1): 21–25. doi:10.1111/j.1574-6968.1993.tb06289.x.
- ↑ Muñoz-Gómez, Sergio A.; Kreutz, Martin; Hess, Sebastian (11 June 2021). "A microbial eukaryote with a unique combination of purple bacteria and green algae as endosymbionts". Science Advances 7 (24). doi:10.1126/sciadv.abg4102. ISSN 2375-2548. PMID 34117067. PMC 8195481. https://www.science.org/doi/10.1126/sciadv.abg4102. Retrieved 26 March 2026.
- ↑ 155.0 155.1 Yubuki, Naoji; Leander, Brian S. (18 July 2018). "Diversity and Evolutionary History of the Symbiontida (Euglenozoa)". Frontiers in Ecology and Evolution 6. doi:10.3389/fevo.2018.00100. ISSN 2296-701X.
- ↑ Seah, Brandon K. B.; Schwaha, Thomas; Volland, Jean-Marie; Huettel, Bruno; Dubilier, Nicole; Gruber-Vodicka, Harald R. (12 July 2017). "Specificity in diversity: single origin of a widespread ciliate-bacteria symbiosis". Proceedings of the Royal Society B: Biological Sciences 284 (1858). doi:10.1098/rspb.2017.0764. ISSN 0962-8452. PMID 28701560.
- ↑ Bright, Monika; Espada-Hinojosa, Salvador; Lagkouvardos, Ilias; Volland, Jean-Marie (7 April 2014). "The giant ciliate Zoothamnium niveum and its thiotrophic epibiont Candidatus Thiobios zoothamnicoli: a model system to study interspecies cooperation". Frontiers in Microbiology 5. doi:10.3389/fmicb.2014.00145. ISSN 1664-302X. PMID 24778630.
- ↑ Hamann, Emmo; Gruber-Vodicka, Harald; Kleiner, Manuel; Tegetmeyer, Halina E.; Riedel, Dietmar et al. (9 June 2016). "Environmental Breviatea harbour mutualistic Arcobacter epibionts". Nature 534 (7606): 254–258. doi:10.1038/nature18297. ISSN 0028-0836. PMID 27279223.
- ↑ Husnik, Filip; Tashyreva, Daria; Boscaro, Vittorio; George, Emma E.; Lukeš, Julius; Keeling, Patrick J. (2021). "Bacterial and archaeal symbioses with protists". Current Biology 31 (13): R862–R877. doi:10.1016/j.cub.2021.05.049. https://linkinghub.elsevier.com/retrieve/pii/S0960982221007478. Retrieved 26 March 2026.
- ↑ Petroni, Giulio; Spring, Stefan; Schleifer, Karl-Heinz; Verni, Franco; Rosati, Giovanna (15 February 2000). "Defensive extrusive ectosymbionts of Euplotidium (Ciliophora) that contain microtubule-like structures are bacteria related to Verrucomicrobia". Proceedings of the National Academy of Sciences 97 (4): 1813–1817. doi:10.1073/pnas.030438197. ISSN 0027-8424. PMID 10660683.
- ↑ Dyer, Betsey Dexter; Khalsa, Ongkar (1993). "Surface bacteria of Streblomastix strix are sensory symbionts". Biosystems 31 (2-3): 169–180. doi:10.1016/0303-2647(93)90046-F.
- ↑ Young, Neil D.; Dyková, Iva; Crosbie, Philip B.B.; Wolf, Matthias; Morrison, Richard N. et al. (2014). "Support for the coevolution of Neoparamoeba and their endosymbionts, Perkinsela amoebae-like organisms". European Journal of Protistology 50 (5): 509–523. doi:10.1016/j.ejop.2014.07.004.
- ↑ 163.0 163.1 163.2 163.3 163.4 163.5 163.6 163.7 163.8 Rizos, Iris; Frada, Miguel J.; Bittner, Lucie; Not, Fabrice (31 July 2024). "Life cycle strategies in free- living unicellular eukaryotes: Diversity, evolution, and current molecular tools to unravel the private life of microorganisms". Journal of Eukaryotic Microbiology 71 (6). doi:10.1111/jeu.13052. PMID 39085163.
- ↑ 164.0 164.1 "Chapter 1. DNA repair as the primary adaptive function of sex in bacteria and eukaryotes". DNA Repair: New Research. Hauppauge, N.Y.: Nova Sci. Publ.. 2012. pp. 1–49. ISBN 978-1-62100-756-2. https://www.researchgate.net/publication/285932986.
- ↑ 165.0 165.1 Milgroom, Michael G. (26 November 2023). "Protozoa". Biology of Infectious Disease. Cham: Springer. pp. 71–87. doi:10.1007/978-3-031-38941-2_6. ISBN 978-3-031-38940-5.
- ↑ Cecere, Ester; Petrocelli, Antonella; Verlaque, Marc (2011). "Vegetative reproduction by multicellular propagules in Rhodophyta: an overview". Marine Ecology 32 (4): 419–437. doi:10.1111/j.1439-0485.2011.00448.x. Bibcode: 2011MarEc..32..419C.
- ↑ Hörandl, Elvira; Bast, Jens; Brandt, Alexander; Scheu, Stefan; Bleidorn, Christoph; Cordellier, Mathilde; Nowrousian, Minou; Begerow, Dominik et al. (2020). "Chapter 7. Genome Evolution of Asexual Organisms and the Paradox of Sex in Eukaryotes". Evolutionary Biology—A Transdisciplinary Approach. Cham: Springer Nature Switzerland AG. doi:10.1007/978-3-030-57246-4_7. ISBN 978-3-030-57246-4. https://serval.unil.ch/notice/serval:BIB_E19BFD9D1BE0.
- ↑ 168.0 168.1 Gibson, Wendy (16 April 2021). "The sexual side of parasitic protists". Molecular and Biochemical Parasitology 243. doi:10.1016/j.molbiopara.2021.111371. PMID 33872659.
- ↑ Hahn, Matthew W, ed (August 2007). "An expanded inventory of conserved meiotic genes provides evidence for sex in Trichomonas vaginalis". PLOS ONE 3 (8). doi:10.1371/journal.pone.0002879. PMID 18663385. Bibcode: 2008PLoSO...3.2879M.
- ↑ "The first sexual lineage and the relevance of facultative sex". Journal of Molecular Evolution 48 (6): 779–783. June 1999. doi:10.1007/PL00013156. PMID 10229582. Bibcode: 1999JMolE..48..779D.
- ↑ "A phylogenomic inventory of meiotic genes; evidence for sex in Giardia and an early eukaryotic origin of meiosis". Current Biology 15 (2): 185–191. January 2005. doi:10.1016/j.cub.2005.01.003. PMID 15668177. Bibcode: 2005CBio...15..185R.
- ↑ "Population genetics provides evidence for recombination in Giardia". Current Biology 17 (22): 1984–1988. November 2007. doi:10.1016/j.cub.2007.10.020. PMID 17980591. Bibcode: 2007CBio...17.1984C.
- ↑ "The chastity of amoebae: re-evaluating evidence for sex in amoeboid organisms". Proceedings of the Royal Society B: Biological Sciences 278 (1715): 2081–2090. July 2011. doi:10.1098/rspb.2011.0289. PMID 21429931. Bibcode: 2011PBioS.278.2081L.
- ↑ Dobell, C. (1909). "Chromidia and the binuclearity hypotheses: A review and a criticism". Quarterly Journal of Microscopical Science 53: 279–326. http://jcs.biologists.org/content/s2-53/210/279.full.pdf.
- ↑ 175.0 175.1 Mulisch, Maria (1996). "Nuclei and Sexual Reproduction". Protistology (3rd ed.). Berlin: E. Schweizerbart'sche Verlagsbuchhandlung. pp. 241–255. ISBN 3-510-65208-8. https://archive.org/details/protistology0000haus.
- ↑ Hall, John D.; McCourt, Richard M. (2017). "Zygnematophyta". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 135–330. doi:10.1007/978-3-319-28149-0_33. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Lynn, Denis H. (2017). "Ciliophora". Handbook of the Protists. 1 (2nd ed.). Springer. pp. 679–730. doi:10.1007/978-3-319-28149-0_23. ISBN 978-3-319-28147-6.
- ↑ Heesch, Svenja; Serrano-Serrano, Martha; Barrera-Redondo, Josué; Luthringer, Rémy; Peters, Akira F.; Destombe, Christophe; Cock, J. Mark; Valero, Myriam et al. (1 July 2021). "Evolution of life cycles and reproductive traits: Insights from the brown algae". Journal of Evolutionary Biology 34 (7): 992–1009. doi:10.1111/jeb.13880. PMID 34096650. Bibcode: 2021JEBio..34..992H.
- ↑ Yoon, Hwan Su; Nelson, Wendy; Lindstrom, Sandra C.; Boo, Sung Min; Pueschel, Curt; Qiu, Huan; Bhattacharya, Debashish (2017). "Rhodophyta". Handbook of the Protists. 1 (2nd ed.). Cham: Springer International Publishing. pp. 89–135. doi:10.1007/978-3-319-28149-0_33. ISBN 978-3-319-28149-0. https://ficoherb.fciencias.unam.mx/Recursos/Bibl/Handbookoftheprotists.pdf.
- ↑ Timmons, Caitlin; Le, Kristine; Rappaport, H. B.; Sterner, Elinor G.; Maurer-Alcalá, Xyrus X. et al. (13 March 2024). "Foraminifera as a model of eukaryotic genome dynamism". mBio 15 (3). doi:10.1128/mbio.03379-23. ISSN 2150-7511. PMID 38329358.
- ↑ "Gametocytogenesis: the puberty of Plasmodium falciparum". Malaria Journal 3: 24. July 2004. doi:10.1186/1475-2875-3-24. PMID 15253774.
- ↑ Votýpka, Jan; Modrý, David; Oborník, Miroslav; Šlapeta, Jan; Lukeš, Julius (2017). "Apicomplexa". Handbook of the Protists. 1 (2nd ed.). Springer. pp. 567–624. doi:10.1007/978-3-319-28149-0_20. ISBN 978-3-319-28147-6.
- ↑ Bulman, Simon; Neuhauser, Sigrid (2017). "Phytomyxea". Handbook of the Protists. 1 (2nd ed.). Springer. pp. 783–803. doi:10.1007/978-3-319-28149-0_24. ISBN 978-3-319-28147-6.
- ↑ Dubey, Jitender P. (2014). "The History and Life Cycle of Toxoplasma gondii". Toxoplasma gondii: The Model Apicomplexan - Perspectives and Methods (2nd ed.). Academic Press. pp. 1–17. doi:10.1016/B978-0-12-396481-6.00001-5. ISBN 978-0-12-396481-6.
- ↑ "Demonstration of genetic exchange during cyclical development of Leishmania in the sand fly vector". Science 324 (5924): 265–268. April 2009. doi:10.1126/science.1169464. PMID 19359589. Bibcode: 2009Sci...324..265A.
- ↑ da Silva, Verônica Santana; Machado, Carlos Renato (2022). "Sex in protists: A new perspective on the reproduction mechanisms of trypanosomatids". Genetics and Molecular Biology 45 (3): e20220065. doi:10.1590/1678-4685-GMB-2022-0065. PMID 36218381.
- ↑ "Are eukaryotic microorganisms clonal or sexual? A population genetics vantage". Proceedings of the National Academy of Sciences of the United States of America 88 (12): 5129–33. June 1991. doi:10.1073/pnas.88.12.5129. PMID 1675793. Bibcode: 1991PNAS...88.5129T.
- ↑ Epstein, Slava; López-García, Purificación (2007). ""Missing" protists: a molecular perspective". Biodiversity and Conservation 17 (2): 261–276. doi:10.1007/s10531-007-9250-y.
- ↑ 189.0 189.1 189.2 189.3 189.4 189.5 189.6 189.7 189.8 Singer, David; Seppey, Christophe V.W.; Lentendu, Guillaume; Dunthorn, Micah; Bass, David et al. (January 2021). "Protist taxonomic and functional diversity in soil, freshwater and marine ecosystems". Environment International 146 (106262). doi:10.1016/j.envint.2020.106262. PMID 33221595. Bibcode: 2021EnInt.14606262S.
- ↑ 190.0 190.1 190.2 Geisen, Stefan; Mitchell, Edward A. D.; Adl, Sina; Bonkowski, Michael; Dunthorn, Micah; Ekelund, Flemming; Fernández, Leonardo D.; Jousset, Alexandre et al. (May 2018). "Soil protists: a fertile frontier in soil biology research". FEMS Microbiology Reviews 42 (3): 293–323. doi:10.1093/femsre/fuy006. PMID 29447350.
- ↑ Mahé, Frédéric; de Vargas, Colomban; Bass, David; Czech, Lucas; Stamatakis, Alexandros et al. (2017). "Parasites dominate hyperdiverse soil protist communities in Neotropical rainforests". Nature Ecology and Evolution 1 (4): 0091. doi:10.1038/s41559-017-0091. PMID 28812652. https://hal.science/hal-03971032.
- ↑ "Oceanic protists with different forms of acquired phototrophy display contrasting biogeographies and abundance". Proceedings of the Royal Society B: Biological Sciences 284 (1860). August 2017. doi:10.1098/rspb.2017.0664. PMID 28768886. Bibcode: 2017PBioS.28470664L.
- ↑ 193.0 193.1 Thoré, Eli S.J.; Muylaert, Koenraad; Bertram, Michael G.; Brodin, Thomas (6 February 2023). "Microalgae". Current Biology 33 (3): R91–R95. doi:10.1016/j.cub.2022.12.032. PMID 36750029. Bibcode: 2023CBio...33R..91T.
- ↑ 194.0 194.1 Pereira, Leonel (7 February 2021). "Macroalgae". Encyclopedia 1 (1): 177–188. doi:10.3390/encyclopedia1010017.
- ↑ 195.0 195.1 Rothäusler, Eva; Gutow, Lars; Thiel, Martin (1 January 2012). "Floating Seaweeds and Their Communities". Seaweed Biology: Novel Insights into Ecophysiology, Ecology and Utilization. Ecological Studies. 219. Springer. pp. 359–380. doi:10.1007/978-3-642-28451-9_17. ISBN 978-3-642-28451-9.
- ↑ "Freshwater protists: unveiling the unexplored in a large floodplain system". Environmental Microbiology 24 (4): 1731–1745. 2022. doi:10.1111/1462-2920.15838. PMID 34783136. Bibcode: 2022EnvMi..24.1731M.
- ↑ 197.0 197.1 197.2 197.3 Rappaport, Hannah B.; Oliverio, Angela M. (16 August 2023). "Extreme environments offer an unprecedented opportunity to understand microbial eukaryotic ecology, evolution, and genome biology". Nature Communications 14 (1). doi:10.1038/s41467-023-40657-4. ISSN 2041-1723. PMID 37587119. Bibcode: 2023NatCo..14.4959R.
- ↑ Rappaport, Hannah B.; Oliverio, Angela M. (16 August 2023). "Extreme environments offer an unprecedented opportunity to understand microbial eukaryotic ecology, evolution, and genome biology". Nature Communications 14 (1). doi:10.1038/s41467-023-40657-4. ISSN 2041-1723. PMID 37587119. Bibcode: 2023NatCo..14.4959R.
- ↑ Bar-On, Yinon M.; Phillips, Rob; Milo, Ron (2018-05-17). "The biomass distribution on Earth" (in en). Proceedings of the National Academy of Sciences 115 (25): 6506–6511. doi:10.1073/pnas.1711842115. ISSN 0027-8424. PMID 29784790. Bibcode: 2018PNAS..115.6506B.
- ↑ Caron, David A.; Worden, Alexandra Z.; Countway, Peter D.; Demir, Elif; Heidelberg, Karla B. (January 2009). "Protists are microbes too: a perspective". The ISME Journal 3 (1): 4–12. doi:10.1038/ismej.2008.101. PMID 19005497. Bibcode: 2009ISMEJ...3....4C.
- ↑ "Local diversity of heathland Cercozoa explored by in-depth sequencing". The ISME Journal 10 (10): 2488–2497. 8 March 2016. doi:10.1038/ismej.2016.31. PMID 26953604. Bibcode: 2016ISMEJ..10.2488H.
- ↑ Santoyo, Gustavo; Orozco-Mosqueda, Ma del Carmen; Babalola, Olubukola Oluranti (2025). "How protists contribute to plant growth and health: Exploring new interactions with the plant microbiome". The Microbe 7. doi:10.1016/j.microb.2025.100361.
- ↑ 203.0 203.1 Moye, Jannika; Hess, Sebastian (3 November 2024). "Broad- range necrophytophagy in the flagellate Orciraptor agilis (Viridiraptoridae, Cercozoa) and the underappreciated role of scavenging among protists". Journal of Eukaryotic Microbiology 72 (2). doi:10.1111/jeu.13065. PMID 39489698.
- ↑ Boddy, Lynne (2016). "Interactions Between Fungi and Other Microbes". in Watkinson, Sarah C.. The Fungi (Third ed.). Academic Press. pp. 337–360. doi:10.1016/B978-0-12-382034-1.00010-4.
- ↑ Lauckner, G. (1980). "Diseases of Protozoa". in Kinne, O.. Diseases of Marine Animals. Volume I. General Aspects. Protozoa to Gastropoda. Chichester: John Wiley & Sons. pp. 75–. https://www.int-res.com/archive/doma_books/DOMA_Vol_I_(general_aspects,_protozoa_to%20gastropoda).pdf.
- ↑ Kamoun, Sophien; Furzer, Oliver; Jones, Jonathan D. G.; Judelson, Howard S.; Ali, Gul Shad et al. (2015). "The Top 10 oomycete pathogens in molecular plant pathology". Molecular Plant Pathology 16 (4): 413–434. doi:10.1111/mpp.12190. ISSN 1464-6722. PMID 25178392.
- ↑ Schwelm, Arne; Badstöber, Julia; Bulman, Simon; Desoignies, Nicolas; Etemadi, Mohammad et al. (2018). "Not in your usual Top 10: protists that infect plants and algae". Molecular Plant Pathology 19 (4): 1029–1044. doi:10.1111/mpp.12580. ISSN 1464-6722. PMID 29024322.
- ↑ Kolářová, Iva; Florent, Isabelle; Valigurová, Andrea (3 August 2022). "Parasitic Protists: Diversity of Adaptations to a Parasitic Lifestyle". Microorganisms 10 (8): 1560. doi:10.3390/microorganisms10081560. ISSN 2076-2607. PMID 36013978.
- ↑ Martinez, Augusto Julio; Visvesvara, Govinda S. (1997). "Free‐living, Amphizoic and Opportunistic Amebas". Brain Pathology 7 (1): 583–598. doi:10.1111/j.1750-3639.1997.tb01076.x. ISSN 1015-6305. PMID 9034567.
- ↑ Baker, J. R. (1977). "Systematics of parasitic protozoa". in Kreier, Julius P.. Parasitic Protozoa. Volume I. Taxonomy, Kinetoplastids, and Flagellates of Fish (1st ed.). New York: Academic Press. pp. 35–56. ISBN 0-12-426001-2. https://archive.org/details/parasiticprotozo0001unse.
- ↑ Brooks, Fred; Rindi, Fabio; Suto, Yasuo; Ohtani, Shuji; Green, Mark (2015). "The Trentepohliales (Ulvophyceae, Chlorophyta): An Unusual Algal Order and its Novel Plant Pathogen— Cephaleuros". Plant Disease 99 (6): 740–753. doi:10.1094/PDIS-01-15-0029-FE. ISSN 0191-2917.
- ↑ Goff, Lynda J.; Ashen, Jon; Moon, Debra (1997). "The evolution of parasites from their hosts: a case study in the parasitic red algae". Evolution 51 (4): 1068–1078. doi:10.1111/j.1558-5646.1997.tb03954.x. ISSN 0014-3820.
- ↑ Haigh, Nicola; Brown, Tamara; Johnson, Devan (2019). "Ichthyotoxic skeleton-forming silicoflagellates in British Columbia, Canada; results from the Harmful Algae Monitoring Program, 1999–2017". in Hess, Philipp. Harmful Algae 2018 – From Ecosystems to Socio-ecosystems. Proceedings of the 18th International Conference on Harmful Algae. Nantes, France: International Society for the Study of Harmful Algae. pp. 167–170. ISBN 978-87-990827-7-3. https://archimer.ifremer.fr/doc/00651/76285/77274.pdf#page=169.
- ↑ Hallegraeff, Gustaaf M. (2003). "Harmful algal blooms: a global overview". in Hallegraeff, G.M.. Manual on harmful marine microalgae (Second ed.). Landais, France: UNESCO Publishing. pp. 25–50. ISBN 92-3-103948-2. https://www.researchgate.net/profile/Donald-Anderson-3/publication/290125074_IOC_Manuals_and_Guides_No33/links/56994aff08aea14769433603/IOC-Manuals-and-Guides-No33.pdf#page=22.
- ↑ Esteban, Genoveva F.; Fenchel, Tom M. (2020). "Symbiosis". Ecology of Protozoa: The Biology of Free-living Phagotrophic Protists (2nd ed.). Cham: Springer. pp. 87–105. doi:10.1007/978-3-030-59979-9_8. ISBN 978-3-030-59979-9.
- ↑ Stanley, George D. (12 May 2006). "Photosymbiosis and the Evolution of Modern Coral Reefs". Science 312 (5775): 857–858. doi:10.1126/science.1123701. ISSN 0036-8075.
- ↑ Decelle, Johan (30 July 2013). "New perspectives on the functioning and evolution of photosymbiosis in plankton: Mutualism or parasitism?". Communicative & Integrative Biology 6 (4). doi:10.4161/cib.24560. ISSN 1942-0889. PMID 23986805.
- ↑ LaJeunesse, Todd C. (2020). "Zooxanthellae". Current Biology 30 (19): R1110–R1113. doi:10.1016/j.cub.2020.03.058.
- ↑ Davy, Simon K.; Allemand, Denis; Weis, Virginia M. (2012). "Cell Biology of Cnidarian-Dinoflagellate Symbiosis". Microbiology and Molecular Biology Reviews 76 (2): 229–261. doi:10.1128/MMBR.05014-11. ISSN 1092-2172.
- ↑ Hernawan, Udhi Eko (6 December 2008). "REVIEW: Symbiosis between the Giant Clams (Bivalvia: Cardiidae) and Zooxanthellae (Dinophyceae)". Biodiversitas Journal of Biological Diversity 9 (1): 53–58. doi:10.13057/biodiv/d090112. ISSN 2085-4722. https://smujo.id/biodiv/article/view/414. Retrieved 27 March 2026.
- ↑ Kaup, Maya; Trull, Sam; Hom, Erik F. Y. (2021). "On the move: sloths and their epibionts as model mobile ecosystems". Biological Reviews 96 (6): 2638–2660. doi:10.1111/brv.12773. ISSN 1464-7931. PMID 34309191.
- ↑ Holt, Corey C.; Boscaro, Vittorio; Van Steenkiste, Niels W. L.; Herranz, Maria; Mathur, Varsha et al. (30 September 2022). "Microscopic marine invertebrates are reservoirs for cryptic and diverse protists and fungi". Microbiome 10 (1). doi:10.1186/s40168-022-01363-3. ISSN 2049-2618. PMID 36180959.
- ↑ Gile, Gillian H. (2024). "Protist symbionts of termites: diversity, distribution, and coevolution". Biological Reviews 99 (2): 622–652. doi:10.1111/brv.13038. ISSN 1464-7931.
- ↑ Lind, Abigail (2025). "Evolution and ecology of commensal gut protists: recent advances". Current Opinion in Genetics & Development 94. doi:10.1016/j.gde.2025.102382. PMID 40683031.
- ↑ "Marine Protists Are Not Just Big Bacteria". Current Biology 27 (11): R541–R549. June 2017. doi:10.1016/j.cub.2017.03.075. PMID 28586691. Bibcode: 2017CBio...27.R541K.
- ↑ Graham, Linda E.; Graham, James M.; Wilcox, Lee W.; Cook, Martha E. (2022). "Chapter 12. The Role of Algae in Biogeochemistry". Algae (4th ed.). LJLM Press. ISBN 978-0-9863935-4-9.
- ↑ 227.0 227.1 227.2 Simpson, A.G.B.; Eglit, Y. (2016). "Protist Diversification". Encyclopedia of Evolutionary Biology. Elsevier. doi:10.1016/b978-0-12-800049-6.00247-x. ISBN 978-0-12-800426-5.
- ↑ 228.0 228.1 228.2 228.3 228.4 228.5 228.6 Lamża, Łukasz (9 November 2025). "Deep‐branching eukaryotes and early events in protist evolution". Biological Reviews 101 (2): 735-750. doi:10.1111/brv.70101. https://www.researchgate.net/profile/Lukasz-Lamza/publication/397445301_Deep-branching_eukaryotes_and_early_events_in_protist_evolution/links/691b6f3cde8143098271e2ac/Deep-branching-eukaryotes-and-early-events-in-protist-evolution.pdf.
- ↑ Archibald, John M. (5 February 2026). "Genomic clues to the origin of eukaryotic cells". Nature 650 (8100): 42–44. doi:10.1038/d41586-025-04094-1. ISSN 0028-0836.
- ↑ Tobiasson, Victor; Luo, Jacob; Wolf, Yuri I.; Koonin, Eugene V. (5 February 2026). "Dominant contribution of Asgard archaea to eukaryogenesis". Nature 650 (8100): 141–149. doi:10.1038/s41586-025-09960-6. ISSN 0028-0836. PMID 41535464.
- ↑ 231.0 231.1 Betts, Holly C.; Puttick, Mark N.; Clark, James W.; Williams, Tom A.; Donoghue, Philip C. J.; Pisani, Davide (20 August 2018). "Integrated genomic and fossil evidence illuminates life's early evolution and eukaryote origin". Nature Ecology & Evolution 2 (10): 1556–1562. doi:10.1038/s41559-018-0644-x. ISSN 2397-334X. PMID 30127539.
- ↑ 232.0 232.1 232.2 232.3 Butterfield, Nicholas J. (2015). "Early evolution of the Eukaryota". Palaeontology 58 (1): 5–17. doi:10.1111/pala.12139. ISSN 0031-0239.
- ↑ 233.0 233.1 Cite error: Invalid
<ref>tag; no text was provided for refs namedStrassert-2021 - ↑ 234.0 234.1 234.2 234.3 234.4 234.5 Brocks, Jochen J.; Nettersheim, Benjamin J.; Adam, Pierre; Schaeffer, Philippe; Jarrett, Amber J. M. et al. (2023). "Lost world of complex life and the late rise of the eukaryotic crown". Nature 618 (7966): 767–773. doi:10.1038/s41586-023-06170-w. PMID 37286610. Bibcode: 2023Natur.618..767B. https://hal.science/hal-04273175/file/Brocks%20et%20al.%20Text%202023%20%2B%20Supplementary.pdf.
- ↑ 235.0 235.1 Porter, Susannah M.; Riedman, Leigh Anne (15 September 2023). "Frameworks for Interpreting the Early Fossil Record of Eukaryotes". Annual Review of Microbiology 77 (1): 173–191. doi:10.1146/annurev-micro-032421-113254. ISSN 0066-4227.
- ↑ 236.0 236.1 236.2 Xiao, Shuhai; Tang, Qing (28 September 2018). "After the boring billion and before the freezing millions: evolutionary patterns and innovations in the Tonian Period". Emerging Topics in Life Sciences 2 (2): 161–171. doi:10.1042/ETLS20170165. ISSN 2397-8554.
- ↑ Müller, Miklós; Mentel, Marek; van Hellemond, Jaap J.; Henze, Katrin; Woehle, Christian et al. (2012). "Biochemistry and Evolution of Anaerobic Energy Metabolism in Eukaryotes". Microbiology and Molecular Biology Reviews 76 (2): 444–495. doi:10.1128/MMBR.05024-11. ISSN 1092-2172.
- ↑ 238.0 238.1 Al Jewari, Caesar; Baldauf, Sandra L. (28 April 2023). "An excavate root for the eukaryote tree of life". Science Advances 9 (17). doi:10.1126/sciadv.ade4973. ISSN 2375-2548. PMID 37115919.
- ↑ Eme, L.; Sharpe, S. C.; Brown, M. W.; Roger, A. J. (1 August 2014). "On the Age of Eukaryotes: Evaluating Evidence from Fossils and Molecular Clocks". Cold Spring Harbor Perspectives in Biology 6 (8): a016139–a016139. doi:10.1101/cshperspect.a016139. ISSN 1943-0264. PMID 25085908.
- ↑ Tekle, Yonas I.; Wang, Fang; Wood, Fiona C.; Anderson, O. Roger; Smirnov, Alexey (1 July 2022). "New insights on the evolutionary relationships between the major lineages of Amoebozoa". Scientific Reports 12 (11173). doi:10.1038/s41598-022-15372-7. PMID 35778543. Bibcode: 2022NatSR..1211173T.
- ↑ Williamson, Kelsey; Eme, Laura; Baños, Hector; McCarthy, Charley G. P.; Susko, Edward et al. (24 April 2025). "A robustly rooted tree of eukaryotes reveals their excavate ancestry". Nature 640 (8060): 974–981. doi:10.1038/s41586-025-08709-5. ISSN 0028-0836.
- ↑ Tikhonenkov, Denis V.; Jamy, Mahwash; Borodina, Anastasia S.; Belyaev, Artem O.; Zagumyonnyi, Dmitry G.; Prokina, Kristina I.; Mylnikov, Alexander P.; Burki, Fabien et al. (2022). "On the origin of TSAR: morphology, diversity and phylogeny of Telonemia". Open Biology (The Royal Society) 12 (3). doi:10.1098/rsob.210325. ISSN 2046-2441. PMID 35291881.
- ↑ Yazaki, Euki; Yabuki, Akinori; Imaizumi, Ayaka; Kume, Keitaro; Hashimoto, Tetsuo; Inagaki, Yuji (2022). "The closest lineage of Archaeplastida is revealed by phylogenomics analyses that include Microheliella maris". Open Biol. 12 (4). doi:10.1098/rsob.210376. PMID 35414259.
- ↑ Torruella, Guifré; Galindo, Luis Javier; Moreira, David; López-García, Purificación (2025). "Phylogenomics of neglected flagellated protists supports a revised eukaryotic tree of life". Current Biology 35 (1): 198–207.e4. doi:10.1016/j.cub.2024.10.075. PMID 39642877. Bibcode: 2025CBio...35..198T.
- ↑ 245.0 245.1 Javaux, Emmanuelle J. (2007). "The Early Eukaryotic Fossil Record". Eukaryotic Membranes and Cytoskeleton. Advances in Experimental Medicine and Biology. 607. New York: Springer. pp. 1–19. doi:10.1007/978-0-387-74021-8_1. ISBN 978-0-387-74020-1.
- ↑ Strother, Paul K.; Brasier, Martin D.; Wacey, David; Timpe, Leslie; Saunders, Martin; Wellman, Charles H. (April 2021). "A possible billion-year-old holozoan with differentiated multicellularity". Current Biology 31 (12): 2658–2665.e2. doi:10.1016/j.cub.2021.03.051. PMID 33852871. Bibcode: 2021CBio...31E2658S.
- ↑ 247.0 247.1 Chai, Shu; Hua, Hong; Ren, Jinjie; Dai, Qiaokun; Cui, Zaihang (January 2021). "Vase-shaped microfossils from the late Ediacaran Dengying Formation of Ningqiang, South China: taxonomy, preservation and biological affinity". Precambrian Research 352. doi:10.1016/j.precamres.2020.105968. Bibcode: 2021PreR..35205968C.
- ↑ "Vase-shaped microfossils from the Neoproterozoic Chuar Group, Grand Canyon: a classification guided by modern testate amoebae". Journal of Paleontology 77 (3): 409–429. May 2003. doi:10.1666/0022-3360(2003)077<0409:VMFTNC>2.0.CO;2. Bibcode: 2003JPal...77..409P. http://www.geol.ucsb.edu/faculty/porter/Papers/Porteretal2003_VSMs.pdf.
- ↑ "Estimating the timing of early eukaryotic diversification with multigene molecular clocks". Proceedings of the National Academy of Sciences of the United States of America 108 (33): 13624–9. August 2011. doi:10.1073/pnas.1110633108. PMID 21810989. Bibcode: 2011PNAS..10813624P.
- ↑ 250.0 250.1 250.2 Min, Xiao; Hua, Hong; Sun, Bo; Dai, Qiaokun; Luo, Jinzhou; Pan, Xiaoqiang; Liu, Ziwei (August 2021). "Diversification of heterotrophic protists at the eve of Cambrian explosion". Global and Planetary Change 203. doi:10.1016/j.gloplacha.2021.103545. Bibcode: 2021GPC...20303545M.
- ↑ Chang, Shan; Feng, Qinglai; Zhang, Lei (14 August 2018). "New Siliceous Microfossils from the Terreneuvian Yanjiahe Formation, South China: The Possible Earliest Radiolarian Fossil Record". Journal of Earth Science 29 (4): 912–919. doi:10.1007/s12583-017-0960-0. Bibcode: 2018JEaSc..29..912C.
- ↑ Zhang, Ke; Feng, Qing-Lai (September 2019). "Early Cambrian radiolarians and sponge spicules from the Niujiaohe Formation in South China". Palaeoworld 28 (3): 234–242. doi:10.1016/j.palwor.2019.04.001.
- ↑ Maletz, Jörg (June 2017). "The identification of putative Lower Cambrian Radiolaria". Revue de Micropaléontologie 60 (2): 233–240. doi:10.1016/j.revmic.2017.04.001. Bibcode: 2017RvMic..60..233M.
- ↑ Pawlowski, Jan; Holzmann, Maria; Berney, Cédric; Fahrni, José; Gooday, Andrew J.; Cedhagen, Thomas; Habura, Andrea; Bowser, Samuel S. (2003). "The evolution of early Foraminifera". Proceedings of the National Academy of Sciences 100 (20): 11494–11498. doi:10.1073/pnas.2035132100. PMID 14504394. Bibcode: 2003PNAS..10011494P.
- ↑ {{{1}}} (2007), "{{{2}}}", in Vickers-Rich, Patricia; Komarower, Patricia, The Rise and Fall of the Ediacaran Biota, Special publications, 286, London: Geological Society, pp. {{{3}}}–{{{4}}}, doi:10.1144/SP286.{{{5}}}, ISBN 9781862392335, OCLC 156823511
- ↑ 256.0 256.1 256.2 Servais, Thomas; Perrier, Vincent; Danelian, Taniel; Klug, Christian; Martin, Ronald; Munnecke, Axel; Nowak, Hendrik; Nützel, Alexander et al. (15 September 2016). "The onset of the 'Ordovician Plankton Revolution' in the late Cambrian". Palaeogeography, Palaeoclimatology, Palaeoecology 458: 12–28. doi:10.1016/j.palaeo.2015.11.003. Bibcode: 2016PPP...458...12S.
- ↑ 257.0 257.1 Vecoli, Marco; Le Hérissé, Alain (October 2004). "Biostratigraphy, taxonomic diversity and patterns of morphological evolution of Ordovician acritarchs (organic-walled microphytoplankton) from the northern Gondwana margin in relation to palaeoclimatic and palaeogeographic changes". Earth-Science Reviews 67 (3–4): 267–311. doi:10.1016/j.earscirev.2004.03.002. Bibcode: 2004ESRv...67..267V.
- ↑ Nowak, Hendrik; Servais, Thomas; Monnet, Claude; Molyneux, Stewart G.; Vandenbroucke, Thijs R.A. (December 2015). "Phytoplankton dynamics from the Cambrian Explosion to the onset of the Great Ordovician Biodiversification Event: A review of Cambrian acritarch diversity". Earth-Science Reviews 151: 117–131. doi:10.1016/j.earscirev.2015.09.005. Bibcode: 2015ESRv..151..117N. http://nora.nerc.ac.uk/id/eprint/514043/1/Nowak%20et%20al%20-%20Cambrian%20Phytoplankton_FINAL%20TEXT.pdf.
- ↑ 259.0 259.1 Zhang, Lei; Algeo, Thomas J.; Zhao, Laishi; Dahl, Tais W.; Chen, Zhong-Qiang; Zhang, Zihu; Poulton, Simon W.; Hughes, Nigel C. et al. (2023-05-12). "Environmental and trilobite diversity changes during the middle-late Cambrian SPICE event". Geological Society of America Bulletin. doi:10.1130/b36421.1. ISSN 0016-7606.
- ↑ Strother, Paul K.; Taylor, Wilson A.; van de Schootbrugge, Bas; Leander, Brian S.; Wellman, Charles H. (2020). "Pellicle ultrastructure demonstrates that Moyeria is a fossil euglenid". Palynology 44 (3): 461–471. doi:10.1080/01916122.2019.1625457. Bibcode: 2020Paly...44..461S.
- ↑ Wang, Kai; Xu, Hong-He; Liu, Bing-Cai; Bai, Jiao; Wang, Yao; Tang, Peng; Lu, Jian-Feng; Wang, Yi (19 May 2023). "Shallow-marine testate amoebae with internal structures from the Lower Devonian of China". iScience 26 (5). doi:10.1016/j.isci.2023.106678. PMID 37182111. Bibcode: 2023iSci...26j6678W.
- ↑ Krings, Michael (September 2022). "Algae from the Lower Devonian Rhynie chert: Populations of a probable saccoderm desmid (Mesotaeniaceae, Zygnematales) preserved in a microbial mat". Review of Palaeobotany and Palynology 304. doi:10.1016/j.revpalbo.2022.104697. Bibcode: 2022RPaPa.30404697K.
- ↑ Krings, Michael; Kerp, Hans (December 2019). "A tiny parasite of unicellular microorganisms from the Lower Devonian Rhynie and Windyfield cherts, Scotland". Review of Palaeobotany and Palynology 271. doi:10.1016/j.revpalbo.2019.104106. Bibcode: 2019RPaPa.27104106K.
- ↑ 264.0 264.1 264.2 264.3 264.4 264.5 264.6 264.7 Suzuki, Noritoshi; Oba, Masahiro (2015). "Oldest Fossil Records of Marine Protists and the Geologic History Toward the Establishment of the Modern-Type Marine Protist World". Marine Protists: Diversity and Dynamics. Springer Japan. pp. 359–394. doi:10.1007/978-4-431-55130-0_15. ISBN 978-4-431-55130-0.
- ↑ 265.0 265.1 265.2 Wever, Patrick; O'Dogherty, Luis; Gorican, Spela (2007). "The plankton turnover at the Permo-Triassic boundary, emphasis on radiolarians". 10th International Meeting of Radiolarian Palaeontologists. 2. Basel: Birkhäuser Verlag. pp. 49–62. doi:10.1007/978-3-7643-8344-2_4.
- ↑ Suzuki, Noritoshi; Aita, Yoshiaki; Yamakita, Satoshi; Kamata, Yoshihito; Takemura, Atsushi; Fujiki, Toru; Ogane, Kaoru; Sakai, Toyosaburo et al. (2007). "The depositional environment of the Induan (Early Triassic) biosiliceous sequence (units 2B and 3 of the Oruatemanu Formation), Arrow Rocks, New Zealand". The oceanic Permian/Triassic boundary sequence at Arrow Rocks (Oruatemanu), Northland, New Zealand. GNS Science Monograph. 24. Lower Hutt, New Zealand: GNS Science. pp. 45–67. ISBN 978-0-478-09919-5. https://www.researchgate.net/publication/280321398.
- ↑ Saesaengseerung, Doungrutai; Agematsu, Sachiko; Sashida, Katsuo; Sardsud, Apsorn (30 June 2009). "Discovery of Lower Permian radiolarian and conodont faunas from the bedded chert of the Chanthaburi area along the Sra Kaeo suture zone, eastern Thailand". Paleontological Research 13 (2): 119–138. doi:10.2517/1342-8144-13.2.119. Bibcode: 2009PalRe..13..119S.
- ↑ Poinar, George O.; Waggoner, Benjamin M.; Bauer, Ulf-Christian (1993). "Terrestrial Soft-Bodied Protists and Other Microorganisms in Triassic Amber". Science 259 (5092): 222–224. doi:10.1126/science.259.5092.222. PMID 17790989. Bibcode: 1993Sci...259..222P.
- ↑ Leung, Tommy L. F. (2017). "Fossils of parasites: what can the fossil record tell us about the evolution of parasitism?". Biological Reviews 92 (1): 410–430. doi:10.1111/brv.12238. PMID 26538112.
- ↑ Poinar, George O. (18 February 2009). "Description of an early Cretaceous termite (Isoptera: Kalotermitidae) and its associated intestinal protozoa, with comments on their co-evolution". Parasites & Vectors 2 (12). doi:10.1186/1756-3305-2-12. PMID 19226475. Bibcode: 2009PVec....2...12P.
- ↑ Poinar, George (October 2009). "Early Cretaceous protist flagellates (Parabasalia: Hypermastigia: Oxymonada) of cockroaches (Insecta: Blattaria) in Burmese amber". Cretaceous Research 30 (5): 1066–1072. doi:10.1016/j.cretres.2009.03.008. Bibcode: 2009CrRes..30.1066P.
- ↑ Knoll, Andrew H.; Follows, Michael J. (26 October 2016). "A bottom-up perspective on ecosystem change in Mesozoic oceans". Proceedings of the Royal Society B: Biological Sciences 283 (1841). doi:10.1098/rspb.2016.1755. PMID 27798303. Bibcode: 2016PBioS.28361755K.
- ↑ "A Molecular Genetic Timescale for the Diversification of Autotrophic Stramenopiles (Ochrophyta): Substantive Underestimation of Putative Fossil Ages". PLOS ONE 5 (9). 2010. doi:10.1371/journal.pone.0012759. PMID 20862282. Bibcode: 2010PLoSO...512759B.
External links
| Wikispecies has information related to Protista |
| Wikispecies has information related to Protoctista |
- UniEuk Taxonomy App
- Tree of Life: Eukaryotes
- Tsukii, Y. (1996). Protist Information Server (database of protist images). Laboratory of Biology, Hosei University. Protist Information Server. Updated: March 22, 2016.
Wikidata ☰ Q10892 entry
 |















