Chemistry:Targeted protein degradation
Targeted protein degradation (TPD) is a drug design strategy that uses small molecules such as PROteolysis TArgeting Chimeras (PROTACs), molecular glues, or related approaches to induce the selective ubiquitination and subsequent proteasomal degradation of target proteins via the ubiquitin–proteasome system or other cellular clearance pathways.[1] Unlike traditional occupancy-driven inhibition, TPD agents catalytically trigger irreversible loss of disease-modifying proteins, enabling the removal of providing sustained duration of action after transient engagement of the target protein.[2] This event-driven approach is being investigated for potential therapeutic applications in oncology,[3] neurodegeneration,[4] and other areas.[5]
Mechanism
Small molecule drugs, compounds typically <1 kD in mass, comprise a large portion of the therapeutic market.[6] These drugs usually operate by agonizing or antagonizing the active site on a disease-linked protein of interest, though allosteric regulation is possible.[7] With an estimated 93% of the human proteome lacking druggable binding sites,[8] methods have been developed to modulate protein activity through binding of any available site rather than only the active site. These drugs contain a target protein binding warhead in addition to a linker-separated active domain. This domain may recruit a second protein to the proximity, induce protease-mediated degradation, or recruit a kinase for directed phosphorylation, among other functions.[9] These drugs expand both the mechanism of action for small molecule therapeutics and the pool of potential protein targets.[9]
Event-driven pharmacology
In the field of targeted protein degraders (TPDs), event-driven pharmacology describes a mechanism of action by which a drug exerts its biological effect not by maintaining continuous occupancy of its target, but rather by initiating an irreversible downstream event, such as the proteolytic degradation of the target protein. This contrasts with traditional occupancy-driven pharmacology, in which drug efficacy depends on sustained binding.[10][2]
TPDs act in a catalytic manner. Transient binding brings the target protein into proximity with an E3 ubiquitin ligase, resulting in its ubiquitination and subsequent degradation by the proteasome. Because the target protein is removed rather than merely inhibited, the pharmacological effect can persist after the TPD dissociates from the target or is cleared from the body, resulting prolonged duration of action at lower drug exposures.[10][2]
Proteolysis-targeting chimeras
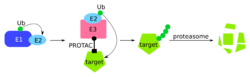
Proteolysis-targeting chimeras (PROTACs) were first reported by Kathleen Sakamoto, Craig Crews, and Raymond Deshaies in 2001. A chimeric molecule consisting of ovalicin (a MetAP-2 small molecule inhibitor) and IκBα phosphopeptide (a recruiter of the SCFβ-TRCP E3 ligase complex) separated by a linker was constructed and shown to induce MetAP-2 degradation in in vitro cell models. Further study confirmed that E3 ligase-mediated ubiquitination and subsequent proteasome degradation was responsible for reduced MetAP-2 levels.[11] Continued work on this system by Craig Crews and others has expanded the potential pool of E3 ligases and degradation targets with Arvinas Inc. founded in 2013 to bring PROTAC drugs to market.[12] As of April 2023, Arvinas has one drug in Stage 3 clinical trials (ARV-471, an estrogen receptor degrader), and two drugs in Stage 2 clinical trials (androgen receptor degraders ARV-110 and ARV-766) for treatment of breast and prostate cancer, respectively.[13] Arvinas released Phase 2 clinical trial results for ARV-471 in December, 2022.[14]
As of May 2025[update], PROTACs in active development that have reached at least Phase II clinical trials:[15]
| INN | Research code | Target | Indication | Developer | NCT number | Phase |
|---|---|---|---|---|---|---|
| Bavdegalutamide | ARV-110 | Androgen receptor | mCRPC (prostate cancer) | Arvinas | NCT03888612 [16] | I |
| Vepdegestrant | ARV-471 | Estrogen receptor | ER+ advanced/metastatic breast cancer | Arvinas / Pfizer | NCT05654623 [17] | III |
| Luxdegalutamide | ARV-766 | Androgen receptor | mCRPC (prostate cancer) | Arvinas | NCT05067140 [18] | II |
| BGB-16673 | BTK | B-cell malignancies (CLL, NHL, MCL) | BeiGene | NCT05006716 [19] | I/II | |
| Zelebrudomide | NX-2127 | BTK, IKZF1/IKZF3 | Relapsed/refractory B-cell malignancies | Nurix Therapeutics | NCT04830137 [20] | I/II |
| Bexobrutideg | NX-5948 | BTK | B-cell malignancies | Nurix Therapeutics | NCT05131022 [21] | I/II |
SNIPERs
SNIPERs (Specific and Non-genetic IAP-dependent Protein Erasers) are chimeric small molecules that hijack the E3 ligase activity of inhibitor of apoptosis proteins (IAPs), such as cIAP1 and XIAP, to selectively induce the ubiquitin-dependent proteasomal degradation of target proteins. SNIPERs are a subclass of PROTAC degraders that specifically use IAP family ligases. Notably, SNIPERs often also degrade the IAP ligases themselves along with the intended targets, which maybe beneficial in cancer cells that overexpress IAPs.[22][23]
Molecular glues
Molecular glues are small molecules that promote targeted protein degradation by stabilizing interactions between E3 ubiquitin ligases and target proteins, enabling their ubiquitin-proteasome mediated breakdown. Unlike bifunctional PROTACs, they lack a linker and directly enhance weak protein–protein interactions, leading to degradation of the target protein. Their small size enables high cell permeability and oral bioavailability. While early examples, such as thalidomide analogs recruiting cereblon, were discovered by accident, mechanistic and structural insights are now starting to drive their rational design.[24]
| Compound name | Target / Mechanism | Primary indication(s) | Most advanced phase | NCT number |
|---|---|---|---|---|
| Iberdomide (CC-220) | CRBN modulator, IKZF1/IKZF3 degrader | Multiple myeloma, lupus | Phase III | NCT04975997 [25] |
| Mezigdomide (CC-92480) | CRBN modulator, IKZF1/3 degrader | Relapsed / refractory multiple myeloma | Phase III | NCT05552976 [26] |
| Avadomide (CC-122) | CRBN modulator, IKZF1/3 degrader | NHL, DLBCL, solid tumors | Phase II | NCT03834623 [27] |
Hydrophobic tag degradation
Hydrophobic tag degraders contain a binding domain in addition to a linker-separated hydrophobic moiety, such as adamantyl, to induce protein degradation. An early example of a hydrophobically tagged degrader is fulvestrant, an estrogen receptor antagonist that contains a long hydrophobic side chain that induces the degradation of the estrogen receptor.[28][29][10] Fulvestrant has inspired the development of additional selective estrogen receptor degraders (SERDs).[30]
As exposed hydrophobicity is characteristic of protein misfolding,[31] the native cell proteasome may recognize and degrade proteins tagged with the hydrophobic moiety. Taavi Neklesa and Craig Crews first reported hydrophobic tag degradation in 2011 as a tool to probe protein function in conjunction with cognate HaloTag fusion proteins.[32] This principle has also been further used to effectively degrade transcription factors[33] (a traditionally difficult class to drug[34]) and cancer-linked EZH2 in in vitro models.[35] As of yet, no drug candidates have been publicly identified making use of this technology.
Alternative strategies
Lysosome-targeting chimeras (LYTACs) have been developed, combining target-binding compounds or antibodies and glycopeptide ligands to stimulate the lysosomal degradation pathway. Unlike the proteasome pathway, this enables the targeted degradation of extracellular and membrane-bound proteins in addition to cytoplasmic ones.[36] Autophagy-targeting chimeras (AUTACs) can be employed to degrade proteins as well as protein aggregates and organelles.[37] AUTAC degradation tags are typically derived from guanine though the particular mechanism of action is still unclear.[38] Autophagosome-tethering compounds (ATTECs) mimic this strategy, directly appending a target protein to the autophagosome membrane for degradation absent the use of a linker.[39] Phosphorylation-inducing chimeric small molecules (PHICS) employ the warhead-linker-recruiter structure to direct phosphorylation of a given target by proximity to a desired kinase. This technique does not necessarily involve protein degradation and may instead be used to modulate protein function to direct or inhibit certain pathways.[40] Further work in the Crews Lab has used chimeric oligonucleotides, the dCas9 protein, and chimeric small molecules to create the TRAFTAC system for generalizable transcription factor degradation.[41]
Advantages
The ability to inhibit or modify enzyme function absent a catalytic pocket binding site target greatly expands the potentially druggable portion of the proteome.[42] Furthermore, most classes of chimeric small molecules can act on many targets over their life cycle,[43] lowering the effective dose compared to traditional inhibitors that act only on one protein at a time.[44] These therapeutics provide an alternative mechanism of action that may be useful as a combination therapy in diseases where drug resistance is a concern.[45] Chimeric drug activity is also highly dependent on distance between targeted proteins[46] allowing effect to be effectively tuned through optimization of the linker structure.
Challenges
The existence of two or more binding domains increases the difficulty of synthesis for chimeric molecules. Each component must be discovered, optimized, and synthesized in such a way that they can be linked together, driving up cost relative to single-domain inhibitors. The large size of chimeric molecules (typically 700-1100 Da) makes effective delivery difficult and increases complexity in pharmacokinetic design.[47][48] Care must be taken to ensure that the molecule is capable of passing through the cell membrane[49] and subsisting long enough to have therapeutic effect. Additionally, protein-protein ternary complexes are generally unstable, adding to the difficulty of chimeric drug design[50]
References
- ↑ "Molecular glues and PROTACs in targeted protein degradation: mechanisms, advances, and therapeutic potential". Biochemical Pharmacology 242 (Pt 3). September 2025. doi:10.1016/j.bcp.2025.117297. PMID 40907798.
- ↑ 2.0 2.1 2.2 "Generating Surprisingly Powerful Pharmacology from Chemically Induced Protein Interactions". Accounts of Chemical Research 58 (15): 2394–2401. August 2025. doi:10.1021/acs.accounts.5c00225. PMID 40705033.
- ↑ "Targeted protein degradation with small molecules for cancer immunotherapy". Asian Journal of Pharmaceutical Sciences 20 (4). August 2025. doi:10.1016/j.ajps.2025.101058. PMID 40791660.
- ↑ "Contemporary trends in targeted protein degradation for neurodegenerative diseases". European Journal of Medicinal Chemistry 300. August 2025. doi:10.1016/j.ejmech.2025.118110. PMID 40914014.
- ↑ "Targeted protein degradation: mechanisms, strategies and application". Signal Transduction and Targeted Therapy 7 (1). April 2022. doi:10.1038/s41392-022-00966-4. PMID 35379777.
- ↑ "Molecular properties that influence the oral bioavailability of drug candidates". Journal of Medicinal Chemistry 45 (12): 2615–2623. June 2002. doi:10.1021/jm020017n. PMID 12036371.
- ↑ "Mechanisms of Action for Small Molecules Revealed by Structural Biology in Drug Discovery". International Journal of Molecular Sciences 21 (15): 5262. July 2020. doi:10.3390/ijms21155262. PMID 32722222.
- ↑ "Elucidating the druggability of the human proteome with eFindSite". Journal of Computer-Aided Molecular Design 33 (5): 509–519. May 2019. doi:10.1007/s10822-019-00197-w. PMID 30888556. Bibcode: 2019JCAMD..33..509K.
- ↑ 9.0 9.1 "Protein degradation technology: a strategic paradigm shift in drug discovery". Journal of Hematology & Oncology 14 (1): 138. September 2021. doi:10.1186/s13045-021-01146-7. PMID 34488823.
- ↑ 10.0 10.1 10.2 "Induced protein degradation: an emerging drug discovery paradigm". Nature Reviews. Drug Discovery 16 (2): 101–114. February 2017. doi:10.1038/nrd.2016.211. PMID 27885283.
- ↑ "Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation". Proceedings of the National Academy of Sciences of the United States of America 98 (15): 8554–8559. July 2001. doi:10.1073/pnas.141230798. PMID 11438690. Bibcode: 2001PNAS...98.8554S.
- ↑ "Nature Biotechnology's academic spinouts of 2013". Nature Biotechnology 32 (3): 229–238. March 2014. doi:10.1038/nbt.2846. PMID 24727773.
- ↑ "PROTAC Protein Degrader Pipeline". Arvinas. https://www.arvinas.com/research-and-development/pipeline/.
- ↑ "Pfizer Strengthens Cancer Standing with Protein Degrader Collaboration". BioSpace. 22 July 2021. https://www.biospace.com/article/pfizer-strikes-another-deal-with-protein-degrader-arvinas-/.
- ↑ "PROTAC Degraders in Clinical Trails: 2025 Update". 2025-05-13. https://www.biochempeg.com/article/434.html.
- ↑ Clinical trial number NCT03888612 for "Trial of ARV-110 in Patients With Metastatic Castration Resistant Prostate Cancer (mCRPC) " at ClinicalTrials.gov
- ↑ Clinical trial number NCT05654623 for "A Study to Learn About a New Medicine Called Vepdegestrant (ARV-471, PF-07850327) in People Who Have Advanced Metastatic Breast Cancer (VERITAC-2) " at ClinicalTrials.gov
- ↑ Clinical trial number NCT05067140 for "A Study of ARV-766 Given by Mouth in Men With Metastatic Prostate Cancer" at ClinicalTrials.gov
- ↑ Clinical trial number NCT05006716 for "A Dose-Escalation and Expansion Study of BGB-16673 in Participants With B-Cell Malignancies (CaDAnCe-101) " at ClinicalTrials.gov
- ↑ Clinical trial number NCT04830137 for "A Study of NX-2127 in Adults With Relapsed/Refractory B-cell Malignancies " at ClinicalTrials.gov
- ↑ Clinical trial number NCT05131022 for "A Study of NX-5948 in Adults With Relapsed/Refractory B-cell Malignancies " at ClinicalTrials.gov
- ↑ "SNIPERs-Hijacking IAP activity to induce protein degradation". Drug Discovery Today. Technologies 31: 35–42. April 2019. doi:10.1016/j.ddtec.2018.12.002. PMID 31200857.
- ↑ "Recent advances in IAP-based PROTACs (SNIPERs) as potential therapeutic agents". Journal of Enzyme Inhibition and Medicinal Chemistry 37 (1): 1437–1453. December 2022. doi:10.1080/14756366.2022.2074414. PMID 35589670.
- ↑ "From serendipity to strategy: molecular glue degraders in cancer therapeutics". Critical Reviews in Biochemistry and Molecular Biology: 1–31. October 2025. doi:10.1080/10409238.2025.2564068. PMID 41032400.
- ↑ Clinical trial number NCT04975997 for "Open-label Study Comparing Iberdomide, Daratumumab and Dexamethasone (IberDd) Versus Daratumumab, Bortezomib, and Dexamethasone (DVd) in Participants With Relapsed or Refractory Multiple Myeloma (RRMM) (EXCALIBER-RRMM)" at ClinicalTrials.gov
- ↑ Clinical trial number NCT05552976 for "A Study to Evaluate Mezigdomide in Combination With Carfilzomib and Dexamethasone (MeziKD) Versus Carfilzomib and Dexamethasone (Kd) in Participants With Relapsed or Refractory Multiple Myeloma (SUCCESSOR-2) (SUCCESSOR-2)" at ClinicalTrials.gov
- ↑ Clinical trial number NCT03834623 for "Avadomide (CC-122) in Combination With Nivolumab in Advanced Melanoma" at ClinicalTrials.gov
- ↑ "Hydrophobic tag-based protein degradation: Development, opportunity and challenge". European Journal of Medicinal Chemistry 260. November 2023. doi:10.1016/j.ejmech.2023.115741. PMID 37607438.
- ↑ "Induced protein degradation: an emerging drug discovery paradigm". Nature Reviews. Drug Discovery 16 (2): 101–114. February 2017. doi:10.1038/nrd.2016.211. PMID 27885283.
- ↑ "Oral SERDs changing the scenery in hormone receptor positive breast cancer, a comprehensive review". Cancer Treatment Reviews 130. November 2024. doi:10.1016/j.ctrv.2024.102825. PMID 39293125.
- ↑ "Exposed hydrophobicity is a key determinant of nuclear quality control degradation". Molecular Biology of the Cell 22 (13): 2384–2395. July 2011. doi:10.1091/mbc.E11-03-0256. PMID 21551067.
- ↑ "Small-molecule hydrophobic tagging-induced degradation of HaloTag fusion proteins". Nature Chemical Biology 7 (8): 538–543. July 2011. doi:10.1038/nchembio.597. PMID 21725302.
- ↑ "Hydrophobic Tagging-Mediated Degradation of Transcription Coactivator SRC-1". International Journal of Molecular Sciences 22 (12): 6407. June 2021. doi:10.3390/ijms22126407. PMID 34203850.
- ↑ "The Human Transcription Factors". Cell 172 (4): 650–665. February 2018. doi:10.1016/j.cell.2018.01.029. PMID 29425488.
- ↑ "Discovery of a first-in-class EZH2 selective degrader". Nature Chemical Biology 16 (2): 214–222. February 2020. doi:10.1038/s41589-019-0421-4. PMID 31819273.
- ↑ "Lysosome Targeting Chimeras (LYTACs) for the Degradation of Secreted and Membrane Proteins | Biological and Medicinal Chemistry". ChemRxiv (Cambridge Open Engage). 2019. doi:10.26434/chemrxiv.7927061.v1. https://chemrxiv.org/engage/chemrxiv/article-details/60c745e6337d6ca40ee27078. Retrieved 2023-04-14.
- ↑ "AUTACs: Cargo-Specific Degraders Using Selective Autophagy". Molecular Cell 76 (5): 797–810.e10. December 2019. doi:10.1016/j.molcel.2019.09.009. PMID 31606272.
- ↑ "Emerging New Concepts of Degrader Technologies". Trends in Pharmacological Sciences 41 (7): 464–474. July 2020. doi:10.1016/j.tips.2020.04.005. PMID 32416934.
- ↑ "ATTEC: a potential new approach to target proteinopathies". Autophagy 16 (1): 185–187. January 2020. doi:10.1080/15548627.2019.1688556. PMID 31690177.
- ↑ "Phosphorylation-Inducing Chimeric Small Molecules". Journal of the American Chemical Society 142 (33): 14052–14057. August 2020. doi:10.1021/jacs.0c05537. PMID 32787262.
- ↑ "Targeted degradation of transcription factors by TRAFTACs: TRAnscription Factor TArgeting Chimeras". Cell Chemical Biology 28 (5): 648–661.e5. May 2021. doi:10.1016/j.chembiol.2021.03.011. PMID 33836141.
- ↑ "Commentary: PROTACs make undruggable targets druggable: Challenge and opportunity". Acta Pharmaceutica Sinica. B 11 (10): 3335–3336. October 2021. doi:10.1016/j.apsb.2021.07.017. PMID 34729320.
- ↑ "PROTAC targeted protein degraders: the past is prologue". Nature Reviews. Drug Discovery 21 (3): 181–200. March 2022. doi:10.1038/s41573-021-00371-6. PMID 35042991.
- ↑ "Recent Advances in PROTACs for Drug Targeted Protein Research". International Journal of Molecular Sciences 23 (18). September 2022. doi:10.3390/ijms231810328. PMID 36142231.
- ↑ "Overcoming Cancer Drug Resistance Utilizing PROTAC Technology". Frontiers in Cell and Developmental Biology 10. 2022. doi:10.3389/fcell.2022.872729. PMID 35547806.
- ↑ "Impact of linker length on the activity of PROTACs". Molecular BioSystems 7 (2): 359–364. February 2011. doi:10.1039/c0mb00074d. PMID 20922213.
- ↑ "Fundamental aspects of DMPK optimization of targeted protein degraders". Drug Discovery Today 25 (6): 969–982. June 2020. doi:10.1016/j.drudis.2020.03.012. PMID 32298797.
- ↑ "Optimising proteolysis-targeting chimeras (PROTACs) for oral drug delivery: a drug metabolism and pharmacokinetics perspective". Drug Discovery Today 25 (10): 1793–1800. October 2020. doi:10.1016/j.drudis.2020.07.013. PMID 32693163.
- ↑ "Understanding and Improving the Membrane Permeability of VH032-Based PROTACs". ACS Medicinal Chemistry Letters 11 (9): 1732–1738. September 2020. doi:10.1021/acsmedchemlett.0c00265. PMID 32939229.
- ↑ "On Ternary Complex Stability in Protein Degradation: In Silico Molecular Glue Binding Affinity Calculations". Journal of Chemical Information and Modeling 63 (8): 2382–2392. April 2023. doi:10.1021/acs.jcim.2c01386. PMID 37037192.
 |
