Naive Bayes classifier
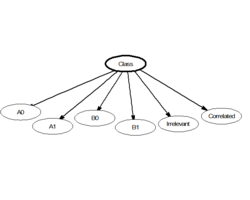
In statistics, naive (sometimes simple or idiot's) Bayes classifiers are a family of "probabilistic classifiers" which assumes that the features are conditionally independent, given the target class.[1] In other words, a naive Bayes model assumes the information about the class provided by each variable is unrelated to the information from the others, with no information shared between the predictors. The highly unrealistic nature of this assumption, called the naive independence assumption, is what gives the classifier its name. These classifiers are some of the simplest Bayesian network models.[2]
Naive Bayes classifiers generally perform worse than more advanced models like logistic regressions, especially at quantifying uncertainty (with naive Bayes models often producing wildly overconfident probabilities). However, they are highly scalable, requiring only one parameter for each feature or predictor in a learning problem. Maximum-likelihood training can be done by evaluating a closed-form expression (simply by counting observations in each group),[3]: 718 rather than the expensive iterative approximation algorithms required by most other models.
Despite the use of Bayes' theorem in the classifier's decision rule, naive Bayes is not (necessarily) a Bayesian method, and naive Bayes models can be fit to data using either Bayesian or frequentist methods.[1][3]
Introduction
Naive Bayes is a simple technique for constructing classifiers: models that assign class labels to problem instances, represented as vectors of feature values, where the class labels are drawn from some finite set. There is not a single algorithm for training such classifiers, but a family of algorithms based on a common principle: all naive Bayes classifiers assume that the value of a particular feature is independent of the value of any other feature, given the class variable. For example, a fruit may be considered to be an apple if it is red, round, and about 10 cm in diameter. A naive Bayes classifier considers each of these features to contribute independently to the probability that this fruit is an apple, regardless of any possible correlations between the color, roundness, and diameter features.
In many practical applications, parameter estimation for naive Bayes models uses the method of maximum likelihood; in other words, one can work with the naive Bayes model without accepting Bayesian probability or using any Bayesian methods.
Despite their naive design and apparently oversimplified assumptions, naive Bayes classifiers have worked quite well in many complex real-world situations. In 2004, an analysis of the Bayesian classification problem showed that there are sound theoretical reasons for the apparently implausible efficacy of naive Bayes classifiers.[4] Still, a comprehensive comparison with other classification algorithms in 2006 showed that Bayes classification is outperformed by other approaches, such as boosted trees or random forests.[5]
An advantage of naive Bayes is that it only requires a small amount of training data to estimate the parameters necessary for classification.[6]
Probabilistic model
Abstractly, naive Bayes is a conditional probability model: it assigns probabilities for each of the K possible outcomes or classes given a problem instance to be classified, represented by a vector encoding some n features (independent variables).[7]
The problem with the above formulation is that if the number of features n is large or if a feature can take on a large number of values, then basing such a model on probability tables is infeasible. The model must therefore be reformulated to make it more tractable. Using Bayes' theorem, the conditional probability can be decomposed as:
In plain English, using Bayesian probability terminology, the above equation can be written as
In practice, there is interest only in the numerator of that fraction, because the denominator does not depend on and the values of the features are given, so that the denominator is effectively constant. The numerator is equivalent to the joint probability model which can be rewritten as follows, using the chain rule for repeated applications of the definition of conditional probability:
Now the "naive" conditional independence assumptions come into play: assume that all features in are mutually independent, conditional on the category . Under this assumption,
Thus, the joint model can be expressed as where denotes proportionality since the denominator is omitted.
This means that under the above independence assumptions, the conditional distribution over the class variable is: where the evidence is a scaling factor dependent only on , that is, a constant if the values of the feature variables are known.
Often, it is only necessary to discriminate between classes. In that case, the scaling factor is irrelevant, and it is sufficient to calculate the log-probability up to a factor:The scaling factor is irrelevant, since discrimination subtracts it away:There are two benefits of using log-probability. One is that it allows an interpretation in information theory, where log-probabilities are units of information in nats. Another is that it avoids arithmetic underflow.
Constructing a classifier from the probability model
The discussion so far has derived the independent feature model, that is, the naive Bayes probability model. The naive Bayes classifier combines this model with a decision rule. One common rule is to pick the hypothesis that is most probable so as to minimize the probability of misclassification; this is known as the maximum a posteriori' or MAP decision rule. The corresponding classifier, a Bayes classifier, is the function that assigns a class label for some k as follows:
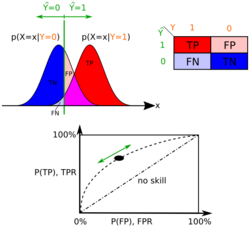
Parameter estimation and event models
A class's prior may be calculated by assuming equiprobable classes, i.e., , or by calculating an estimate for the class probability from the training set: To estimate the parameters for a feature's distribution, one must assume a distribution or generate nonparametric models for the features from the training set.[8]
The assumptions on distributions of features are called the "event model" of the naive Bayes classifier. For discrete features like the ones encountered in document classification (include spam filtering), multinomial and Bernoulli distributions are popular. These assumptions lead to two distinct models, which are often confused.[9][10]
Gaussian naive Bayes
When dealing with continuous data, a typical assumption is that the continuous values associated with each class are distributed according to a normal (or Gaussian) distribution. For example, suppose the training data contains a continuous attribute, . The data is first segmented by the class, and then the mean and variance of is computed in each class. Let be the mean of the values in associated with class , and let be the Bessel corrected variance of the values in associated with class . Suppose one has collected some observation value . Then, the probability density of given a class , i.e., , can be computed by plugging into the equation for a normal distribution parameterized by and . Formally,
Another common technique for handling continuous values is to use binning to discretize the feature values and obtain a new set of Bernoulli-distributed features. Some literature suggests that this is required in order to use naive Bayes, but it is not true, as the discretization may throw away discriminative information.[1]
Sometimes the distribution of class-conditional marginal densities is far from normal. In these cases, kernel density estimation can be used for a more realistic estimate of the marginal densities of each class. This method, which was introduced by John and Langley,[8] can boost the accuracy of the classifier considerably.[11][12]
Multinomial naive Bayes
With a multinomial event model, samples (feature vectors) represent the frequencies with which certain events have been generated by a multinomial where is the probability that event i occurs (or K such multinomials in the multiclass case). A feature vector is then a histogram, with counting the number of times event i was observed in a particular instance. This is the event model typically used for document classification, with events representing the occurrence of a word in a single document (see bag of words assumption).[13] The likelihood of observing a histogram x is given by: where .
The multinomial naive Bayes classifier becomes a linear classifier when expressed in log-space:[14] where and . Estimating the parameters in log space is advantageous since multiplying a large number of small values can lead to significant rounding error. Applying a log transform reduces the effect of this rounding error.
If a given class and feature value never occur together in the training data, then the frequency-based probability estimate will be zero, because the probability estimate is directly proportional to the number of occurrences of a feature's value. This is problematic because it will wipe out all information in the other probabilities when they are multiplied. Therefore, it is often desirable to incorporate a small-sample correction, called pseudocount, in all probability estimates such that no probability is ever set to be exactly zero. This way of regularizing naive Bayes is called Laplace smoothing when the pseudocount is one, and Lidstone smoothing in the general case.
Rennie et al. discuss problems with the multinomial assumption in the context of document classification and possible ways to alleviate those problems, including the use of tf–idf weights instead of raw term frequencies and document length normalization, to produce a naive Bayes classifier that is competitive with support vector machines.[14]
Bernoulli naive Bayes
In the multivariate Bernoulli event model, features are independent Boolean variables (binary variables) describing inputs. Like the multinomial model, this model is popular for document classification tasks,[9] where binary term occurrence features are used rather than term frequencies. If is a Boolean expressing the occurrence or absence of the i'th term from the vocabulary, then the likelihood of a document given a class is given by:[9] where is the probability of class generating the term . This event model is especially popular for classifying short texts. It has the benefit of explicitly modelling the absence of terms. Note that a naive Bayes classifier with a Bernoulli event model is not the same as a multinomial NB classifier with frequency counts truncated to one.
Semi-supervised parameter estimation
Given a way to train a naive Bayes classifier from labeled data, it's possible to construct a semi-supervised training algorithm that can learn from a combination of labeled and unlabeled data by running the supervised learning algorithm in a loop:[15]
- Given a collection of labeled samples L and unlabeled samples U, start by training a naive Bayes classifier on L.
- Until convergence, do:
- Predict class probabilities for all examples x in .
- Re-train the model based on the probabilities (not the labels) predicted in the previous step.
Convergence is determined based on improvement to the model likelihood , where denotes the parameters of the naive Bayes model.
This training algorithm is an instance of the more general expectation–maximization algorithm (EM): the prediction step inside the loop is the E-step of EM, while the re-training of naive Bayes is the M-step. The algorithm is formally justified by the assumption that the data are generated by a mixture model, and the components of this mixture model are exactly the classes of the classification problem.[15]
Discussion
Despite the fact that the far-reaching independence assumptions are often inaccurate, the naive Bayes classifier has several properties that make it surprisingly useful in practice. In particular, the decoupling of the class conditional feature distributions means that each distribution can be independently estimated as a one-dimensional distribution. This helps alleviate problems stemming from the curse of dimensionality, such as the need for data sets that scale exponentially with the number of features. While naive Bayes often fails to produce a good estimate for the correct class probabilities,[16] this may not be a requirement for many applications. For example, the naive Bayes classifier will make the correct MAP decision rule classification so long as the correct class is predicted as more probable than any other class. This is true regardless of whether the probability estimate is slightly, or even grossly inaccurate. In this manner, the overall classifier can be robust enough to ignore serious deficiencies in its underlying naive probability model.[17] Other reasons for the observed success of the naive Bayes classifier are discussed in the literature cited below.
Relation to logistic regression
In the case of discrete inputs (indicator or frequency features for discrete events), naive Bayes classifiers form a generative-discriminative pair with multinomial logistic regression classifiers: each naive Bayes classifier can be considered a way of fitting a probability model that optimizes the joint likelihood , while logistic regression fits the same probability model to optimize the conditional .[18]
More formally, we have the following:
Theorem — Naive Bayes classifiers on binary features are subsumed by logistic regression classifiers.
Consider a generic multiclass classification problem, with possible classes , then the (non-naive) Bayes classifier gives, by Bayes theorem:
The naive Bayes classifier gives where
This is exactly a logistic regression classifier.
The link between the two can be seen by observing that the decision function for naive Bayes (in the binary case) can be rewritten as "predict class if the odds of exceed those of ". Expressing this in log-space gives:
The left-hand side of this equation is the log-odds, or logit, the quantity predicted by the linear model that underlies logistic regression. Since naive Bayes is also a linear model for the two "discrete" event models, it can be reparametrised as a linear function . Obtaining the probabilities is then a matter of applying the logistic function to , or in the multiclass case, the softmax function.
Discriminative classifiers have lower asymptotic error than generative ones; however, research by Ng and Jordan has shown that in some practical cases naive Bayes can outperform logistic regression because it reaches its asymptotic error faster.[18]
Examples
Person classification
Problem: classify whether a given person is a male or a female based on the measured features. The features include height, weight, and foot size. Although with NB classifier we treat them as independent, they are not in reality.
Training
Example training set below.
| Person | height (feet) | weight (lbs) | foot size (inches) |
|---|---|---|---|
| male | 6 | 180 | 12 |
| male | 5.92 (5'11") | 190 | 11 |
| male | 5.58 (5'7") | 170 | 12 |
| male | 5.92 (5'11") | 165 | 10 |
| female | 5 | 100 | 6 |
| female | 5.5 (5'6") | 150 | 8 |
| female | 5.42 (5'5") | 130 | 7 |
| female | 5.75 (5'9") | 150 | 9 |
The classifier created from the training set using a Gaussian distribution assumption would be (given variances are unbiased sample variances):
| Person | mean (height) | variance (height) | mean (weight) | variance (weight) | mean (foot size) | variance (foot size) |
|---|---|---|---|---|---|---|
| male | 5.855 | 3.5033 × 10−2 | 176.25 | 12.292 | 11.25 | 9.1667 × 10−1 |
| female | 5.4175 | 9.7225 × 10−2 | 132.5 | 5.5833 | 7.5 | 1.6667 |
The following example assumes equiprobable classes so that P(male)= P(female) = 0.5. This prior probability distribution might be based on prior knowledge of frequencies in the larger population or in the training set.
Testing
Below is a sample to be classified as male or female.
| Person | height (feet) | weight (lbs) | foot size (inches) |
|---|---|---|---|
| sample | 6 | 130 | 8 |
In order to classify the sample, one has to determine which posterior is greater, male or female. For the classification as male the posterior is given by
For the classification as female the posterior is given by
The evidence (also termed normalizing constant) may be calculated:
However, given the sample, the evidence is a constant and thus scales both posteriors equally. It therefore does not affect classification and can be ignored. The probability distribution for the sex of the sample can now be determined: where and are the parameters of normal distribution which have been previously determined from the training set. Note that a value greater than 1 is OK here – it is a probability density rather than a probability, because height is a continuous variable.
Since posterior numerator is greater in the female case, the prediction is that the sample is female.
Document classification
Here is a worked example of naive Bayesian classification to the document classification problem. Consider the problem of classifying documents by their content, for example into spam and non-spam e-mails. Imagine that documents are drawn from a number of classes of documents which can be modeled as sets of words where the (independent) probability that the i-th word of a given document occurs in a document from class C can be written as
(For this treatment, things are further simplified by assuming that words are randomly distributed in the document - that is, words are not dependent on the length of the document, position within the document with relation to other words, or other document-context.)
Then the probability that a given document D contains all of the words , given a class C, is
The question that has to be answered is: "what is the probability that a given document D belongs to a given class C?" In other words, what is ?
Now by definition and
Bayes' theorem manipulates these into a statement of probability in terms of likelihood.
Assume for the moment that there are only two mutually exclusive classes, S and ¬S (e.g. spam and not spam), such that every element (email) is in either one or the other; and
Using the Bayesian result above, one can write:
Dividing one by the other gives:
Which can be re-factored as:
Thus, the probability ratio p(S | D) / p(¬S | D) can be expressed in terms of a series of likelihood ratios. The actual probability p(S | D) can be easily computed from log (p(S | D) / p(¬S | D)) based on the observation that p(S | D) + p(¬S | D) = 1.
Taking the logarithm of all these ratios, one obtains:
(This technique of "log-likelihood ratios" is a common technique in statistics. In the case of two mutually exclusive alternatives (such as this example), the conversion of a log-likelihood ratio to a probability takes the form of a sigmoid curve: see logit for details.)
Finally, the document can be classified as follows. It is spam if (i. e., ), otherwise it is not spam.
Spam filtering
Naive Bayes classifiers are a popular statistical technique of e-mail filtering. They typically use bag-of-words features to identify email spam, an approach commonly used in text classification. Naive Bayes classifiers work by correlating the use of tokens (typically words, or sometimes other things), with spam and non-spam e-mails and then using Bayes' theorem to calculate a probability that an email is or is not spam.
Naive Bayes spam filtering is a baseline technique for dealing with spam that can tailor itself to the email needs of individual users and give low false positive spam detection rates that are generally acceptable to users. Bayesian algorithms were used for email filtering as early as 1996. Although naive Bayesian filters did not become popular until later, multiple programs were released in 1998 to address the growing problem of unwanted email.[19] The first scholarly publication on Bayesian spam filtering was by Sahami et al. in 1998.[20]
Variants of the basic technique have been implemented in a number of research works and commercial software products.[21] Many modern mail clients implement Bayesian spam filtering. Users can also install separate email filtering programs. Server-side email filters, such as DSPAM, Rspamd,[22] SpamAssassin,[23] SpamBayes,[24] Bogofilter, and ASSP, make use of Bayesian spam filtering techniques, and the functionality is sometimes embedded within mail server software itself. CRM114, oft cited as a Bayesian filter, is not intended to use a Bayes filter in production, but includes the ″unigram″ feature for reference.[25]
Dealing with rare words
In the case a word has never been met during the learning phase, both the numerator and the denominator are equal to zero, both in the general formula and in the spamicity formula. The software can decide to discard such words for which there is no information available.
More generally, the words that were encountered only a few times during the learning phase cause a problem, because it would be an error to trust blindly the information they provide. A simple solution is to simply avoid taking such unreliable words into account as well.
Applying again Bayes' theorem, and assuming the classification between spam and ham of the emails containing a given word ("replica") is a random variable with beta distribution, some programs decide to use a corrected probability:
where:
- is the corrected probability for the message to be spam, knowing that it contains a given word ;
- is the strength we give to background information about incoming spam ;
- is the probability of any incoming message to be spam ;
- is the number of occurrences of this word during the learning phase ;
- is the spamicity of this word.
(Demonstration:[26])
This corrected probability is used instead of the spamicity in the combining formula.
This formula can be extended to the case where n is equal to zero (and where the spamicity is not defined), and evaluates in this case to .
Other heuristics
"Neutral" words like "the", "a", "some", or "is" (in English), or their equivalents in other languages, can be ignored. These are also known as Stop words. More generally, some bayesian filtering filters simply ignore all the words which have a spamicity next to 0.5, as they contribute little to a good decision. The words taken into consideration are those whose spamicity is next to 0.0 (distinctive signs of legitimate messages), or next to 1.0 (distinctive signs of spam). A method can be for example to keep only those ten words, in the examined message, which have the greatest absolute value |0.5 − pI|.
Some software products take into account the fact that a given word appears several times in the examined message,[27] others don't.
Some software products use patterns (sequences of words) instead of isolated natural languages words.[28] For example, with a "context window" of four words, they compute the spamicity of "Viagra is good for", instead of computing the spamicities of "Viagra", "is", "good", and "for". This method gives more sensitivity to context and eliminates the Bayesian noise better, at the expense of a bigger database.
Disadvantages
Depending on the implementation, Bayesian spam filtering may be susceptible to Bayesian poisoning, a technique used by spammers in an attempt to degrade the effectiveness of spam filters that rely on Bayesian filtering. A spammer practicing Bayesian poisoning will send out emails with large amounts of legitimate text (gathered from legitimate news or literary sources). Spammer tactics include insertion of random innocuous words that are not normally associated with spam, thereby decreasing the email's spam score, making it more likely to slip past a Bayesian spam filter. However, with (for example) Paul Graham's scheme only the most significant probabilities are used, so that padding the text out with non-spam-related words does not affect the detection probability significantly.
Words that normally appear in large quantities in spam may also be transformed by spammers. For example, «Viagra» would be replaced with «Viaagra» or «V!agra» in the spam message. The recipient of the message can still read the changed words, but each of these words is met more rarely by the Bayesian filter, which hinders its learning process. As a general rule, this spamming technique does not work very well, because the derived words end up recognized by the filter just like the normal ones.[29]
Another technique used to try to defeat Bayesian spam filters is to replace text with pictures, either directly included or linked. The whole text of the message, or some part of it, is replaced with a picture where the same text is "drawn". The spam filter is usually unable to analyze this picture, which would contain the sensitive words like «Viagra». However, since many mail clients disable the display of linked pictures for security reasons, the spammer sending links to distant pictures might reach fewer targets. Also, a picture's size in bytes is bigger than the equivalent text's size, so the spammer needs more bandwidth to send messages directly including pictures. Some filters are more inclined to decide that a message is spam if it has mostly graphical contents. A solution used by Google in its Gmail email system is to perform an OCR (Optical Character Recognition) on every mid to large size image, analyzing the text inside.[30][31]
See also
- AODE
- Anti-spam techniques
- Bayes classifier
- Bayesian network
- Bayesian poisoning
- Email filtering
- Linear classifier
- Logistic regression
- Markovian discrimination
- Mozilla Thunderbird mail client with native implementation of Bayes filters[32][33]
- Perceptron
- Random naive Bayes
- Take-the-best heuristic
References
- ↑ 1.0 1.1 1.2 Hand, D. J.; Yu, K. (2001). "Idiot's Bayes — not so stupid after all?". International Statistical Review 69 (3): 385–399. doi:10.2307/1403452. ISSN 0306-7734.
- ↑ McCallum, Andrew. "Graphical Models, Lecture2: Bayesian Network Representation". https://people.cs.umass.edu/~mccallum/courses/gm2011/02-bn-rep.pdf.
- ↑ 3.0 3.1 Russell, Stuart; Norvig, Peter (2003) [1995]. A Modern Approach (2nd ed.). Prentice Hall. ISBN 978-0137903955.
- ↑ Zhang, Harry. "The Optimality of Naive Bayes". FLAIRS2004 conference. http://www.cs.unb.ca/profs/hzhang/publications/FLAIRS04ZhangH.pdf.
- ↑ Caruana, R.; Niculescu-Mizil, A. (2006). "An empirical comparison of supervised learning algorithms". Proc. 23rd International Conference on Machine Learning.
- ↑ "Why does Naive Bayes work better when the number of features >> sample size compared to more sophisticated ML algorithms?". https://stats.stackexchange.com/q/379383.
- ↑ Narasimha Murty, M.; Susheela Devi, V. (2011). Pattern Recognition: An Algorithmic Approach. Springer. ISBN 978-0857294944.
- ↑ 8.0 8.1 John, George H.; Langley, Pat (1995). "Estimating Continuous Distributions in Bayesian Classifiers". Proc. Eleventh Conf. on Uncertainty in Artificial Intelligence. Morgan Kaufmann. pp. 338–345. https://dl.acm.org/doi/10.5555/2074158.2074196.
- ↑ 9.0 9.1 9.2 McCallum, Andrew; Nigam, Kamal (1998). "A comparison of event models for Naive Bayes text classification". AAAI-98 workshop on learning for text categorization. 752. http://www.kamalnigam.com/papers/multinomial-aaaiws98.pdf.
- ↑ Metsis, Vangelis; Androutsopoulos, Ion; Paliouras, Georgios (2006). "Spam filtering with Naive Bayes—which Naive Bayes?". Third conference on email and anti-spam (CEAS). 17. https://www.researchgate.net/publication/221650814.
- ↑ Piryonesi, S. Madeh; El-Diraby, Tamer E. (2020-06-01). "Role of Data Analytics in Infrastructure Asset Management: Overcoming Data Size and Quality Problems". Journal of Transportation Engineering, Part B: Pavements 146 (2): 04020022. doi:10.1061/JPEODX.0000175.
- ↑ Hastie, Trevor. (2001). The elements of statistical learning : data mining, inference, and prediction : with 200 full-color illustrations. Tibshirani, Robert., Friedman, J. H. (Jerome H.). New York: Springer. ISBN 0-387-95284-5. OCLC 46809224.
- ↑ James, Gareth; Witten, Daniela; Hastie, Trevor; Tibshirani, Robert (2021). An introduction to statistical learning: with applications in R (Second ed.). New York, NY: Springer. p. 157. doi:10.1007/978-1-0716-1418-1. ISBN 978-1-0716-1418-1. https://link.springer.com/book/10.1007/978-1-0716-1418-1. Retrieved 10 November 2024.
- ↑ 14.0 14.1 Rennie, J.; Shih, L.; Teevan, J.; Karger, D. (2003). "Tackling the poor assumptions of naive Bayes classifiers". ICML. http://people.csail.mit.edu/~jrennie/papers/icml03-nb.pdf.
- ↑ 15.0 15.1 Nigam, Kamal; McCallum, Andrew; Thrun, Sebastian; Mitchell, Tom (2000). "Learning to classify text from labeled and unlabeled documents using EM". Machine Learning 39 (2/3): 103–134. doi:10.1023/A:1007692713085. http://www.kamalnigam.com/papers/emcat-aaai98.pdf.
- ↑ Niculescu-Mizil, Alexandru; Caruana, Rich (2005). "Predicting good probabilities with supervised learning". ICML. doi:10.1145/1102351.1102430. http://machinelearning.wustl.edu/mlpapers/paper_files/icml2005_Niculescu-MizilC05.pdf. Retrieved 2016-04-24.
- ↑ Rish, Irina (2001). "An empirical study of the naive Bayes classifier". IJCAI Workshop on Empirical Methods in AI. http://www.research.ibm.com/people/r/rish/papers/RC22230.pdf.
- ↑ 18.0 18.1 Ng, Andrew Y.; Jordan, Michael I. (2002). "On discriminative vs. generative classifiers: A comparison of logistic regression and naive Bayes". NIPS. 14. http://papers.nips.cc/paper/2020-on-discriminative-vs-generative-classifiers-a-comparison-of-logistic-regression-and-naive-bayes.
- ↑ Brunton, Finn (2013). Spam: A Shadow History of the Internet. MIT Press. p. 136. ISBN 9780262018876. https://books.google.com/books?id=QF7EjCRg5CIC&pg=PA136. Retrieved 2017-09-13.
- ↑ "A Bayesian approach to filtering junk e-mail". AAAI'98 Workshop on Learning for Text Categorization. 1998. http://robotics.stanford.edu/users/sahami/papers-dir/spam.pdf.
- ↑ "Junk Mail Controls". MozillaZine. November 2009. http://kb.mozillazine.org/Junk_Mail_Controls.
- ↑ "Rspamd statistic settings". docs.rspamd.com. https://docs.rspamd.com/configuration/statistic.
- ↑ "Installation". Ubuntu manuals. 2010-09-18. http://manpages.ubuntu.com/manpages/gutsy/man1/sa-learn.1p.html. "Gary Robinson’s f(x) and combining algorithms, as used in SpamAssassin"
- ↑ "Background Reading". SpamBayes project. 2010-09-18. https://spambayes.sourceforge.net/background.html. "Sharpen your pencils, this is the mathematical background (such as it is).* The paper that started the ball rolling: Paul Graham's A Plan for Spam.* Gary Robinson has an interesting essay suggesting some improvements to Graham's original approach.* Gary Robinson's Linux Journal article discussed using the chi squared distribution."
- ↑ "Archived copy". https://crm114.sourceforge.net/docs/classify_details.txt.
- ↑ Gary Robinson (2003). "A statistical approach to the spam problem". Linux Journal. http://www.linuxjournal.com/article/6467. Retrieved 2007-07-19.
- ↑ Brian Burton (2003). "SpamProbe - Bayesian Spam Filtering Tweaks". https://spamprobe.sourceforge.net/paper.html.
- ↑ Jonathan A. Zdziarski (2004). "Bayesian Noise Reduction: Contextual Symmetry Logic Utilizing Pattern Consistency Analysis". http://bnr.nuclearelephant.com/l.
- ↑ Paul Graham (2002), A Plan for Spam
- ↑ "Gmail uses Google's innovative technology to keep spam out of your inbox". http://www.google.com/mail/help/intl/en_GB/fightspam/spamexplained.html.
- ↑ Zhu, Z.; Jia, Z; Xiao, H; Zhang, G; Liang, H.; Wang, P. (2014). "A Modified Minimum Risk Bayes and It's Application in Spam Filtering". in Li, S; Jin, Q; Jiang, X et al. (in en). Frontier and Future Development of Information Technology in Medicine and Education. Lecture Notes in Electrical Engineering. 269. Dordrecht: Springer. pp. 2155–2159. doi:10.1007/978-94-007-7618-0_261. ISBN 978-94-007-7617-3.
- ↑ Hristea, Florentina T. (2013) (in en). The Naïve Bayes Model for Unsupervised Word Sense Disambiguation. London; Berlin: Springer- Verlag Heidelberg Berlin. pp. 70. ISBN 978-3-642-33692-8.
- ↑ Zheng, J.; Tang, Yongchuan (2005). "One Generalization of the Naive Bayes to Fuzzy Sets and the Design of the Fuzzy Naive Bayes Classifier". in Mira, Jose; Álvarez, Jose R (in en). Artificial Intelligence and Knowledge Engineering Applications: A Bioinspired Approach. Lecture Notes in Computer Science. 3562. Berlin: Springer, Berlin, Heidelberg. p. 281. doi:10.1007/11499305_29. ISBN 978-3-540-26319-7.
Further reading
- Domingos, Pedro; Pazzani, Michael (1997). "On the optimality of the simple Bayesian classifier under zero-one loss". Machine Learning 29 (2/3): 103–137. doi:10.1023/A:1007413511361. http://citeseer.ist.psu.edu/domingos97optimality.html.
- Webb, G. I.; Boughton, J.; Wang, Z. (2005). "Not So Naive Bayes: Aggregating One-Dependence Estimators". Machine Learning 58 (1): 5–24. doi:10.1007/s10994-005-4258-6.
- Mozina, M.; Demsar, J.; Kattan, M.; Zupan, B. (2004). "Nomograms for Visualization of Naive Bayesian Classifier". Proc. PKDD-2004. pp. 337–348. http://eprints.fri.uni-lj.si/154/01/PKDD_camera_mozina.pdf. Retrieved 2014-04-01.
- Maron, M. E. (1961). "Automatic Indexing: An Experimental Inquiry". Journal of the ACM 8 (3): 404–417. doi:10.1145/321075.321084.
- Minsky, M. (1961). "Steps toward Artificial Intelligence". Proc. IRE. 49. pp. 8–30.
External links
- Book Chapter: Naive Bayes text classification, Introduction to Information Retrieval
- Naive Bayes for Text Classification with Unbalanced Classes
 |
