Biology:Coiled coil
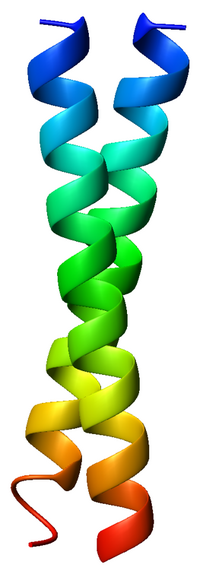
A coiled coil is a structural motif in proteins in which two to seven[1] alpha-helices are coiled together like the strands of a rope. (Dimers and trimers are the most common types.) They have been found in roughly 5-10% of proteins and have a variety of functions.[2] They are one of the most widespread motifs found in protein-protein interactions. To aid protein study, several tools have been developed to predict coiled-coils in protein structures.[3] Many coiled coil-type proteins are involved in important biological functions, such as the regulation of gene expression — e.g., transcription factors. Notable examples are the oncoproteins c-Fos and c-Jun, as well as the muscle protein tropomyosin.
Discovery
The possibility of coiled coils for α-keratin was initially somewhat controversial. Linus Pauling and Francis Crick independently came to the conclusion that this was possible at about the same time. In the summer of 1952, Pauling visited the laboratory in England where Crick worked. Pauling and Crick met and spoke about various topics; at one point, Crick asked whether Pauling had considered coiled coils (a term Crick came up with), to which Pauling said he had. Upon returning to the United States, Pauling resumed research on the topic. He concluded that coiled coils exist, and submitted a lengthy manuscript to the journal Nature in October. Pauling's son Peter Pauling worked at the same lab as Crick, and mentioned the report to him. Crick believed that Pauling had stolen his idea, and submitted a shorter note to Nature a few days after Pauling's manuscript arrived. Eventually, after some controversy and frequent correspondences, Crick's lab declared that the idea had been reached independently by both researchers, and that no intellectual theft had occurred.[4] In his note (which was published first due to its shorter length), Crick proposed the coiled coil and as well as mathematical methods for determining their structure.[5] Remarkably, this was soon after the structure of the alpha helix was suggested in 1951 by Linus Pauling and coworkers.[6] These studies were published in the absence of knowledge of a keratin sequence. The first keratin sequences were determined by Hanukoglu and Fuchs in 1982.[7][8]
Based on sequence and secondary structure prediction analyses identified the coiled-coil domains of keratins.[8] These models have been confirmed by structural analyses of coiled-coil domains of keratins.[9]
Molecular structure
Coiled coils usually contain a repeated pattern, hxxhcxc, of hydrophobic (h) and charged (c) amino-acid residues, referred to as a heptad repeat.[10] The positions in the heptad repeat are usually labeled abcdefg, where a and d are the hydrophobic positions, often being occupied by isoleucine, leucine, or valine. Folding a sequence with this repeating pattern into an alpha-helical secondary structure causes the hydrophobic residues to be presented as a 'stripe' that coils gently around the helix in left-handed fashion, forming an amphipathic structure. The most favorable way for two such helices to arrange themselves in the water-filled environment of the cytoplasm is to wrap the hydrophobic strands against each other sandwiched between the hydrophilic amino acids. Thus, it is the burial of hydrophobic surfaces that provides the thermodynamic driving force for the oligomerization. The packing in a coiled-coil interface is exceptionally tight, with almost complete van der Waals contact between the side-chains of the a and d residues. This tight packing was originally predicted by Francis Crick in 1952[5] and is referred to as knobs into holes packing.
The α-helices may be parallel or anti-parallel, and usually adopt a left-handed super-coil (Figure 1). Although disfavored, a few right-handed coiled coils have also been observed in nature and in designed proteins.[11]
Biological roles
As coiled-coil domains are common among a significant amount of proteins in a wide variety of protein families, they help proteins fulfill various functions in the cell. Their primary feature is to facilitate protein-protein interaction and keep proteins or domains interlocked. This feature corresponds to several subfunctions, including membrane fusion, molecular spacing, oligomerization tags, vesicle movement, aid in movement proteins, cell structure, and more.[12]
Membrane fusion

A coiled coil domain plays a role in human immunodeficiency virus type 1 (HIV-1) infection. Viral entry into CD4-positive cells commences when three subunits of a glycoprotein 120 (gp120) bind to CD4 receptor and a coreceptor.[13] Glycoprotein gp120 is closely associated with a trimer of gp41 via van der Waals interactions. Eventually, the gp41 N-terminal fusion peptide sequence anchors into the host cell. A spring-loaded mechanism is responsible for bringing the viral and cell membranes in close enough proximity that they will fuse. The origin of the spring-loaded mechanism lies within the exposed gp41, which contains two consecutive heptad repeats (HR1 and HR2) following the fusion peptide at the N terminus of the protein. HR1 forms a parallel, trimeric coiled coil onto which HR2 region coils, forming the trimer-of-hairpins (or six-helix bundle) structure, thereby facilitating membrane fusion through bringing the membranes close to each other.[14] The virus then enters the cell and begins its replication. Recently, inhibitors derived from HR2 such as Fuzeon (DP178, T-20) that bind to the HR1 region on gp41 have been developed.[15] However, peptides derived from HR1 have little viral inhibition efficacy due to the propensity for these peptides to aggregate in solution. Chimeras of these HR1-derived peptides with GCN4 leucine zippers have been developed and have shown to be more active than Fuzeon.[16] Human immunodeficiency virus type 2 has a membrane envelope glycoprotein with similar structure to HIV-1 gp41, but containing substitutions for a glycine amino acid residue in the coiled coil domain that may impact trimer stability.[17]
The proteins SNAP-25, synaptobrevin, and syntaxin-1 have alpha-helices which interact with each other to form a coiled-coil SNARE complex. Zippering the domains together provides the necessary energy for vesicle fusion to occur.[18]
Molecular spacers
The coiled-coil motif may also act as a spacer between two objects within a cell. The lengths of these molecular spacer coiled-coil domains are highly conserved. The purpose of these molecular spacers may be to separate protein domains, thus keeping them from interacting, or to separate vesicles within the cell to mediate vesicle transport. An example of this first purpose is Omp‐α found in T. maritima.[19] Other proteins keep vesicles apart, such as p115, giantin, and GM130 which interact with each other via coiled-coil motifs and act as a tether between the Golgi and a nearby vesicle.[20] The family of proteins related to this activity of tethering vesicles to the Golgi are known as golgins.[21] Finally, there are several proteins with coiled-coil domains involved in the kinetochore, which keeps chromosomes separated during cell division. These proteins include Ndc-80, and Nuf2p. Related proteins interact with microtubules during cell division, of which mutation leads to cell death.[22]
As oligomerization tags
Because of their specific interaction coiled coils can be used as "tags" to stabilize or enforce a specific oligomerization state.[23] A coiled coil interaction has been observed to drive the oligomerization of the BBS2 and BBS7 subunits of the BBSome.[24][25] Because coiled-coils generally interact with other coiled coils, they are found in proteins which are required to form dimers or tetramers with more copies of themselves.[26] Because of their ability in driving protein oligomerization, they have also been studied in their use in forming synthetic nanostructures.[27]
Design

The general problem of deciding on the folded structure of a protein when given the amino acid sequence (the so-called protein folding problem) has only been solved partially. However, the coiled coil is one of a relatively small number of folding motifs for which the relationships between the sequence and the final folded structure are comparatively well understood.[29][30] Harbury et al. performed a landmark study using an archetypal coiled coil, GCN4, in which rules that govern the way that peptide sequence affects the oligomeric state (that is, the number of alpha-helices in the final assembly) were established.[31][32] The GCN4 coiled coil is a 31-amino-acid (which equates to just over four heptads) parallel, dimeric (i.e., consisting of two alpha-helices) coiled coil and has a repeated isoleucine (or I, in single-letter code) and leucine (L) at the a and d positions, respectively, and forms a dimeric coiled coil. When the amino acids in the a and d positions were changed from I at a and L at d to I at a and I at d, a trimeric (three alpha-helices) coiled coil was formed. Furthermore, switching the positions of L to a and I to d resulted in the formation of a tetrameric (four alpha-helices) coiled coil. These represent a set of rules for the determination of coiled coil oligomeric states and allows scientists to effectively "dial-in" the oligomerization behavior. Another aspect of coiled coil assembly that is relatively well understood, at least in the case of dimeric coiled coils, is that placing a polar residue (in particular asparagine, N) at opposing a positions forces parallel assembly of the coiled coil. This effect is due to a self-complementary hydrogen bonding between these residues, which would go unsatisfied if an N were paired with, for instance, an L on the opposing helix.[33]
It was recently demonstrated by Peacock, Pikramenou and co-workers that coiled coils may be self-assembled using lanthanide(III) ions as a template, thus producing novel imaging agents.[34]
Biomedical applications

Coiled-coil motifs have been experimented on as possible building block for nanostructures, in part because of their simple design and wide range of function based primarily on facilitating protein-protein interaction. Simple guidelines for de novo synthesis of new proteins containing coiled-coil domains have led to many applications being hypothesized, including drug delivery, regenerating tissue, protein origami, and much more.[35] In regards to drug delivery, coiled-coil domains would help overcome some of the hazards of chemotherapeutic drugs, by keeping them from leaking into healthy tissue as they are transported to their target. Coiled-coil domains can be made to bind to specific proteins or cell surface markers, allowing for more precise targeting in drug delivery.[36] Other functions would be to help store and transport drugs within the body that would otherwise degrade rapidly, by creating nanotubes and other structure svia the interlocking of coiled-coil motifs.[35] By utilizing the function of oligomerization of proteins via coiled-coil domains, antigen display can be amplified in vaccines, increasing their effectiveness.[37]
The oligomerization of coiled-coil motifs allows for the creation of protein origami and protein building blocks. Metal-ligand interactions, covalent bonds, and ionic interactions have been studied to manipulate possible coiled-coil interactions in this field of study.[35] Several different nanostructures can be made by combining coiled-coil motifs such that they are self-assembling building blocks. However, several difficulties remain with stability.[38] Using peptides with coiled-coil motifs for scaffolding has made it easier to create 3D structures for cell culturing. 3D hydrogels can be made with these peptides, and then cells may be loaded into the matrix.[39] This has applications in the study of tissue, tissue engineering, and more.[35]
References
- ↑ "A seven-helix coiled coil". Proceedings of the National Academy of Sciences of the United States of America 103 (42): 15457–62. Oct 2006. doi:10.1073/pnas.0604871103. PMID 17030805. Bibcode: 2006PNAS..10315457L.
- ↑ Szczepaniak, Krzysztof; Bukala, Adriana; da Silva Neto, Antonio Marinho; Ludwiczak, Jan; Dunin-Horkawicz, Stanislaw (2021-04-01). Elofsson, Arne. ed. "A library of coiled-coil domains: from regular bundles to peculiar twists" (in en). Bioinformatics 36 (22–23): 5368–5376. doi:10.1093/bioinformatics/btaa1041. ISSN 1367-4803. PMID 33325494. PMC 8016460. https://academic.oup.com/bioinformatics/article/36/22-23/5368/6039120.
- ↑ Walshaw, John; Woolfson, Derek N (2001-04-13). "SOCKET: a program for identifying and analysing coiled-coil motifs within protein structures11Edited by J. Thornton". Journal of Molecular Biology 307 (5): 1427–1450. doi:10.1006/jmbi.2001.4545. ISSN 0022-2836. PMID 11292353. https://www.sciencedirect.com/science/article/pii/S0022283601945450.
- ↑ "Narrative 43, Coils Upon Coils". Linus Pauling and the Structure of Proteins. Oregon State University Special Collections and Archives Research Center. http://scarc.library.oregonstate.edu/coll/pauling/proteins/narrative/page43.html.
- ↑ 5.0 5.1 "Is alpha-keratin a coiled coil?". Nature 170 (4334): 882–883. November 1952. doi:10.1038/170882b0. PMID 13013241. Bibcode: 1952Natur.170..882C.
- ↑ "The structure of proteins; two hydrogen-bonded helical configurations of the polypeptide chain". Proceedings of the National Academy of Sciences of the United States of America 37 (4): 205–211. April 1951. doi:10.1073/pnas.37.4.205. PMID 14816373. Bibcode: 1951PNAS...37..205P.
- ↑ "The cDNA sequence of a human epidermal keratin: divergence of sequence but conservation of structure among intermediate filament proteins". Cell 31 (1): 243–252. November 1982. doi:10.1016/0092-8674(82)90424-X. PMID 6186381. https://zenodo.org/record/890743.
- ↑ 8.0 8.1 "The cDNA sequence of a Type II cytoskeletal keratin reveals constant and variable structural domains among keratins". Cell 33 (3): 915–924. July 1983. doi:10.1016/0092-8674(83)90034-X. PMID 6191871. https://zenodo.org/record/890739.
- ↑ "Proteopedia entry: coiled-coil structure of keratins". Biochemistry and Molecular Biology Education 42 (1): 93–94. Jan 2014. doi:10.1002/bmb.20746. PMID 24265184.
- ↑ "Coiled coil domains: stability, specificity, and biological implications". ChemBioChem 5 (2): 170–176. February 2004. doi:10.1002/cbic.200300781. PMID 14760737.
- ↑ "High-resolution protein design with backbone freedom". Science 282 (5393): 1462–1467. November 1998. doi:10.1126/science.282.5393.1462. PMID 9822371.
- ↑ Rose, Annkatrin; Schraegle, Shannon J.; Stahlberg, Eric A.; Meier, Iris (2005-11-16). "Coiled-coil protein composition of 22 proteomes – differences and common themes in subcellular infrastructure and traffic control". BMC Evolutionary Biology 5 (1): 66. doi:10.1186/1471-2148-5-66. ISSN 1471-2148. PMID 16288662. Bibcode: 2005BMCEE...5...66R.
- ↑ "Structural basis of coreceptor recognition by HIV-1 envelope spike". Nature 565 (7739): 318–323. January 2019. doi:10.1038/s41586-018-0804-9. PMID 30542158.
- ↑ "HIV: cell binding and entry". Cold Spring Harbor Perspectives in Medicine 2 (8). August 2012. doi:10.1101/cshperspect.a006866. PMID 22908191.
- ↑ "Resistance to enfuvirtide, the first HIV fusion inhibitor". The Journal of Antimicrobial Chemotherapy 54 (2): 333–40. August 2004. doi:10.1093/jac/dkh330. PMID 15231762.
- ↑ "Design of potent inhibitors of HIV-1 entry from the gp41 N-peptide region". Proceedings of the National Academy of Sciences of the United States of America 98 (20): 11187–11192. September 2001. doi:10.1073/pnas.201392898. PMID 11572974. Bibcode: 2001PNAS...9811187E.
- ↑ "Structure of the transmembrane domain of HIV-1 envelope glycoprotein". The FEBS Journal 284: 1171-1177. 2017. doi:10.1111/febs.13954. PMID 27868386.
- ↑ Chen, Yu A.; Scheller, Richard H. (February 2001). "SNARE-mediated membrane fusion" (in en). Nature Reviews Molecular Cell Biology 2 (2): 98–106. doi:10.1038/35052017. ISSN 1471-0072. PMID 11252968. https://www.nature.com/articles/35052017.
- ↑ Truebestein, Linda; Leonard, Thomas A. (September 2016). "Coiled-coils: The long and short of it" (in en). BioEssays 38 (9): 903–916. doi:10.1002/bies.201600062. ISSN 0265-9247. PMID 27492088.
- ↑ Linstedt, Adam D.; Jesch, Stephen A.; Mehta, Amy; Lee, Tina H.; Garcia-Mata, Rafael; Nelson, David S.; Sztul, Elizabeth (April 2000). "Binding Relationships of Membrane Tethering Components". Journal of Biological Chemistry 275 (14): 10196–10201. doi:10.1074/jbc.275.14.10196. ISSN 0021-9258. PMID 10744704.
- ↑ Witkos, Tomasz M.; Lowe, Martin (2016-01-11). "The Golgin Family of Coiled-Coil Tethering Proteins". Frontiers in Cell and Developmental Biology 3: 86. doi:10.3389/fcell.2015.00086. ISSN 2296-634X. PMID 26793708.
- ↑ Jeyaprakash, A. Arockia; Santamaria, Anna; Jayachandran, Uma; Chan, Ying Wai; Benda, Christian; Nigg, Erich A.; Conti, Elena (May 2012). "Structural and Functional Organization of the Ska Complex, a Key Component of the Kinetochore-Microtubule Interface". Molecular Cell 46 (3): 274–286. doi:10.1016/j.molcel.2012.03.005. ISSN 1097-2765. PMID 22483620.
- ↑ "Your personalized protein structure: Andrei N. Lupas fused to GCN4 adaptors". Journal of Structural Biology 186 (3): 380–5. June 2014. doi:10.1016/j.jsb.2014.01.013. PMID 24486584.
- ↑ Chou, Hui-Ting; Apelt, Luise; Farrell, Daniel P.; White, Susan Roehl; Woodsmith, Jonathan; Svetlov, Vladimir; Goldstein, Jaclyn S.; Nager, Andrew R. et al. (3 September 2019). "The Molecular Architecture of Native BBSome Obtained by an Integrated Structural Approach". Structure 27 (9): 1384–1394. doi:10.1016/j.str.2019.06.006. PMID 31303482.
- ↑ Ludlam, WG; Aoba, T; Cuéllar, J; Bueno-Carrasco, MT; Makaju, A; Moody, JD; Franklin, S; Valpuesta, JM et al. (17 September 2019). "Molecular architecture of the Bardet-Biedl syndrome protein 2-7-9 subcomplex.". The Journal of Biological Chemistry 294 (44): 16385–16399. doi:10.1074/jbc.RA119.010150. PMID 31530639.
- ↑ Cabezon, Elena; Butler, P. Jonathan G.; Runswick, Michael J.; Walker, John E. (August 2000). "Modulation of the Oligomerization State of the Bovine F1-ATPase Inhibitor Protein, IF1, by pH" (in en). Journal of Biological Chemistry 275 (33): 25460–25464. doi:10.1074/jbc.M003859200. PMID 10831597.
- ↑ Park, Won Min (2020-05-19). "Coiled-Coils: The Molecular Zippers that Self-Assemble Protein Nanostructures" (in en). International Journal of Molecular Sciences 21 (10): 3584. doi:10.3390/ijms21103584. ISSN 1422-0067. PMID 32438665.
- ↑ 28.0 28.1 Lapenta, Fabio; Aupič, Jana; Strmšek, Žiga; Jerala, Roman (2018). "Coiled coil protein origami: from modular design principles towards biotechnological applications" (in en). Chemical Society Reviews 47 (10): 3530–3542. doi:10.1039/C7CS00822H. ISSN 0306-0012. PMID 29400389. http://xlink.rsc.org/?DOI=C7CS00822H.
- ↑ "Peptide and protein building blocks for synthetic biology: from programming biomolecules to self-organized biomolecular systems". ACS Chemical Biology 3 (1): 38–50. January 2008. doi:10.1021/cb700249v. PMID 18205291.
- ↑ "Complex networks govern coiled-coil oligomerization--predicting and profiling by means of a machine learning approach". Molecular & Cellular Proteomics 10 (5). May 2011. doi:10.1074/mcp.M110.004994. PMID 21311038.
- ↑ "A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants". Science 262 (5138): 1401–1407. November 1993. doi:10.1126/science.8248779. PMID 8248779. Bibcode: 1993Sci...262.1401H.
- ↑ "Crystal structure of an isoleucine-zipper trimer". Nature 371 (6492): 80–83. September 1994. doi:10.1038/371080a0. PMID 8072533. Bibcode: 1994Natur.371...80H.
- ↑ "The design of coiled-coil structures and assemblies". Fibrous Proteins: Coiled-Coils, Collagen and Elastomers. Advances in Protein Chemistry. 70. 2005. pp. 79–112. doi:10.1016/S0065-3233(05)70004-8. ISBN 978-0-12-034270-9.
- ↑ "De novo design of Ln(III) coiled coils for imaging applications". Journal of the American Chemical Society 136 (4): 1166–1169. January 2014. doi:10.1021/ja408741h. PMID 24405157.
- ↑ 35.0 35.1 35.2 35.3 Jorgensen, Michael D.; Chmielewski, Jean (2022). "Recent advances in coiled-coil peptide materials and their biomedical applications". Chemical Communications 58 (83): 11625–11636. doi:10.1039/d2cc04434j. ISSN 1359-7345. PMID 36172799.
- ↑ McFarlane, Ainsley A.; Orriss, George L.; Stetefeld, Jörg (December 2009). "The use of coiled-coil proteins in drug delivery systems" (in en). European Journal of Pharmacology 625 (1–3): 101–107. doi:10.1016/j.ejphar.2009.05.034. PMID 19835864.
- ↑ Schroeder, Ulrich; Graff, Alexandra; Buchmeier, Sabine; Rigler, Per; Silvan, Unai; Tropel, David; Jockusch, Brigitte M.; Aebi, Ueli et al. (March 2009). "Peptide Nanoparticles Serve as a Powerful Platform for the Immunogenic Display of Poorly Antigenic Actin Determinants". Journal of Molecular Biology 386 (5): 1368–1381. doi:10.1016/j.jmb.2008.11.023. ISSN 0022-2836. PMID 19063898.
- ↑ Park, Won Min (January 2020). "Coiled-Coils: The Molecular Zippers that Self-Assemble Protein Nanostructures" (in en). International Journal of Molecular Sciences 21 (10): 3584. doi:10.3390/ijms21103584. ISSN 1422-0067. PMID 32438665.
- ↑ Dexter, A. F.; Fletcher, N. L.; Creasey, R. G.; Filardo, F.; Boehm, M. W.; Jack, K. S. (2017). "Fabrication and characterization of hydrogels formed from designer coiled-coil fibril-forming peptides" (in en). RSC Advances 7 (44): 27260–27271. doi:10.1039/C7RA02811C. ISSN 2046-2069. Bibcode: 2017RSCAd...727260D. http://xlink.rsc.org/?DOI=C7RA02811C.
Further reading
- "The Packing of α-Helices: Simple Coiled-Coils". Acta Crystallogr 6 (8): 689–697. 1953. doi:10.1107/S0365110X53001964. Bibcode: 1953AcCry...6..689C.
- "Geometrical criteria for formation of coiled-coil structures of polypeptide chains". Macromolecules 9 (3): 395–407. 1976. doi:10.1021/ma60051a004. PMID 940353. Bibcode: 1976MaMol...9..395N.
- "A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants". Science 262 (5138): 1401–1407. November 1993. doi:10.1126/science.8248779. PMID 8248779. Bibcode: 1993Sci...262.1401H.
- "An engineered allosteric switch in leucine-zipper oligomerization". Nature Structural Biology 3 (6): 510–515. June 1996. doi:10.1038/nsb0696-510. PMID 8646536.
- "High-resolution protein design with backbone freedom". Science 282 (5393): 1462–1467. November 1998. doi:10.1126/science.282.5393.1462. PMID 9822371.
- "Coiled-coils: stability, specificity, and drug delivery potential". Advanced Drug Delivery Reviews 54 (8): 1113–1129. October 2002. doi:10.1016/S0169-409X(02)00058-3. PMID 12384310.
- "Improving coiled-coil stability by optimizing ionic interactions". Journal of Molecular Biology 318 (3): 901–910. May 2002. doi:10.1016/S0022-2836(02)00114-6. PMID 12054832.
- "Long coiled-coil proteins and membrane traffic". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1641 (2–3): 71–85. August 2003. doi:10.1016/S0167-4889(03)00088-0. PMID 12914949.
- "Coiled coil domains: stability, specificity, and biological implications". ChemBioChem 5 (2): 170–176. February 2004. doi:10.1002/cbic.200300781. PMID 14760737.
External links
Coiled-coil related software
Prediction, detection, and visualization
- Paircoil2 / Paircoil
- bCIPA Estimates Tm values for coiled coil pairs
- bCIPA library screen Screens a library of sequences against a single defined target and estimates Tm values for all coiled coils pairs.
- bCIPA Interactome Screen Screens all interactions between a selection of defined sequences and estimates Tm values for all coiled coil pairs.
- STRAP contains an algorithm to predict coiled-coils from AA-sequences.
- PrOCoil predicts the oligomerization of coiled coil proteins and visualizes the contribution of each individual amino acid to the overall oligomeric tendency.
- DrawCoil creates helical wheel diagrams for coiled coils of any oligomerization state and orientation.
Databases
- Spiricoil uses protein domain annotation to predict coiled coil presence and oligormeric state for all completely sequenced organisms
- CC+ is a relational database of coiled coils found in the PDB
- SUPERFAMILY protein domain annotation for all completely sequenced organisms based on the expertly curated SCOP coiled coil class
 |
